Abstract
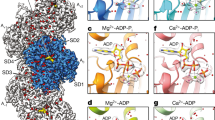
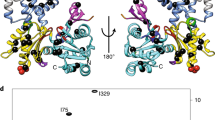
Actin has maintained an exquisite degree of sequence conservation over large evolutionary distances for reasons that are not understood. The desire to explain phenomena from muscle contraction to cytokinesis in mechanistic detail has driven the generation of an atomic model of the actin filament (F-actin). Here we use electron cryomicroscopy to show that frozen-hydrated actin filaments contain a multiplicity of different structural states. We show (at ∼10 Å resolution) that subdomain 2 can be disordered and can make multiple contacts with the C terminus of a subunit above it. We link a number of disease-causing mutations in the human ACTA1 gene to the most structurally dynamic elements of actin. Because F-actin is structurally polymorphic, it cannot be described using only one atomic model and must be understood as an ensemble of different states.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Feuer, G., Molnar, F., Pettko, E. & Straub, F.B. Studies on the composition and polymerization of actin. Hung. Acta Physiol. 1, 150–163 (1948).
Pollard, T.D. & Borisy, G.G. Cellular motility driven by assembly and disassembly of actin filaments. Cell 112, 453–465 (2003).
Cooke, R. The mechanism of muscle contraction. CRC Crit. Rev. Biochem. 21, 53–118 (1986).
Salwinski, L. et al. The Database of Interacting Proteins: 2004 update. Nucleic Acids Res. 32, D449–D451 (2004).
Kabsch, W., Mannherz, H.G., Suck, D., Pai, E.F. & Holmes, K.C. Atomic structure of the actin:DNase I complex. Nature 347, 37–44 (1990).
Otterbein, L.R., Graceffa, P. & Dominguez, R. The crystal structure of uncomplexed actin in the ADP state. Science 293, 708–711 (2001).
Rould, M.A., Wan, Q., Joel, P.B., Lowey, S. & Trybus, K.M. Crystal structures of expressed non-polymerizable monomeric actin in the ADP and ATP states. J. Biol. Chem. 281, 31909–31919 (2006).
Holmes, K.C., Popp, D., Gebhard, W. & Kabsch, W. Atomic model of the actin filament. Nature 347, 44–49 (1990).
Lorenz, M., Popp, D. & Holmes, K.C. Refinement of the F-actin model against x-ray fiber diffraction data by the use of a directed mutation algorithm. J. Mol. Biol. 234, 826–836 (1993).
Tirion, M.M., ben-Avraham, D., Lorenz, M. & Holmes, K.C. Normal modes as refinement parameters for the F-actin model. Biophys. J. 68, 5–12 (1995).
Oda, T., Iwasa, M., Aihara, T., Maeda, Y. & Narita, A. The nature of the globular- to fibrous-actin transition. Nature 457, 441–445 (2009).
Egelman, E.H., Francis, N. & DeRosier, D.J. F-actin is a helix with a random variable twist. Nature 298, 131–135 (1982).
Galkin, V.E., VanLoock, M.S., Orlova, A. & Egelman, E.H. A new internal mode in F-actin helps explain the remarkable evolutionary conservation of actin's sequence and structure. Curr. Biol. 12, 570–575 (2002).
Schmid, M.F., Sherman, M.B., Matsudaira, P. & Chiu, W. Structure of the acrosomal bundle. Nature 431, 104–107 (2004).
Kim, E. & Reisler, E. Intermolecular coupling between loop 38–52 and the c-terminus in actin filaments. Biophys. J. 71, 1914–1919 (1996).
Kim, E. et al. Cross-linking constraints on F-actin structure. J. Mol. Biol. 299, 421–429 (2000).
Heintz, D., Kany, H. & Kalbitzer, H.R. Mobility of the N-terminal segment of rabbit skeletal muscle F-actin detected by 1H and 19F nuclear magnetic resonance spectroscopy. Biochemistry 35, 12686–12693 (1996).
Orlova, A., Yu, X. & Egelman, E.H. Three-dimensional reconstruction of a co-complex of F-actin with antibody Fab fragments to actin's amino-terminus. Biophys. J. 66, 276–285 (1994).
Oztug Durer, Z.A., Diraviyam, K., Sept, D., Kudryashov, D.S. & Reisler, E. F-actin structure destabilization and DNase I binding loop: fluctuations mutational cross-linking and electron microscopy analysis of loop states and effects on F-actin. J. Mol. Biol. 395, 544–557 (2010).
Doolittle, R.F. The origins and evolution of eukaryotic proteins. Phil. Trans. R. Soc. Lond. B 349, 235–240 (1995).
Sheterline, P., Clayton, J. & Sparrow, J. Actin. Protein Profile 2, 1–103 (1995).
Derman, A.I. et al. Phylogenetic analysis identifies many uncharacterized actin-like proteins (Alps) in bacteria: regulated polymerization, dynamic instability and treadmilling in Alp7A. Mol. Microbiol. 73, 534–552 (2009).
Egelman, E.H. Actin allostery again? Nat. Struct. Biol. 8, 735–736 (2001).
McCormack, E.A., Llorca, O., Carrascosa, J.L., Valpuesta, J.M. & Willison, K.R. Point mutations in a hinge linking the small and large domains of beta-actin result in trapped folding intermediates bound to cytosolic chaperonin CCT. J. Struct. Biol. 135, 198–204 (2001).
Drummond, D.A., Bloom, J.D., Adami, C., Wilke, C.O. & Arnold, F.H. Why highly expressed proteins evolve slowly. Proc. Natl. Acad. Sci. USA 102, 14338–14343 (2005).
Strzelecka-Gołaszewska, H., Mossakowska, M., Wozniak, A., Moraczewska, J. & Nakayama, H. Long-range conformational effects of proteolytic removal of the last three residues of actin. Biochem. J. 307, 527–534 (1995).
Kim, E., Motoki, M., Seguro, K., Muhlrad, A. & Reisler, E. Conformational changes in subdomain 2 of G-actin: fluorescence probing by dansyl ethylenediamine attached to Gln-41. Biophys. J. 69, 2024–2032 (1995).
Süel, G.M., Lockless, S.W., Wall, M.A. & Ranganathan, R. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nat. Struct. Biol. 10, 59–69 (2003).
Drummond, D.R., Peckham, M., Sparrow, J.C. & White, D.C. Alteration in crossbridge kinetics caused by mutations in actin. Nature 348, 440–442 (1990).
Prochniewicz, E. & Yanagida, T. Inhibition of sliding movement of F-actin by crosslinking emphasizes the role of actin structure in the mechanism of motility. J. Mol. Biol. 216, 761–772 (1990).
Kim, E. et al. Intrastrand cross-linked actin between Gln-41 and Cys-374. III. Inhibition of motion and force generation with myosin. Biochemistry 37, 17801–17809 (1998).
Schwyter, D.H., Kron, S.J., Toyoshima, Y.Y., Spudich, J.A. & Reisler, E. Subtilisin cleavage of actin inhibits in vitro sliding movement of actin filaments over myosin. J. Cell Biol. 111, 465–470 (1990).
Egelman, E.H. A robust algorithm for the reconstruction of helical filaments using single-particle methods. Ultramicroscopy 85, 225–234 (2000).
Kudryashov, D.S. et al. The crystal structure of a cross-linked actin dimer suggests a detailed molecular interface in F-actin. Proc. Natl. Acad. Sci. USA 102, 13105–13110 (2005).
Stokasimov, E., McKane, M. & Rubenstein, P.A. Role of intermonomer ionic bridges in the stabilization of the actin filament. J. Biol. Chem. 283, 34844–34854 (2008).
Dominguez, R. Actin-binding proteins—a unifying hypothesis. Trends Biochem. Sci. 29, 572–578 (2004).
Kim, E., Miller, C.J. & Reisler, E. Polymerization and in vitro motility properties of yeast actin: a comparison with rabbit skeletal α-actin. Biochemistry 35, 16566–16572 (1996).
McKane, M., Wen, K.K., Meyer, A. & Rubenstein, P.A. Effect of the substitution of muscle actin-specific subdomain 1 and 2 residues in yeast actin on actin function. J. Biol. Chem. 281, 29916–29928 (2006).
Orlova, A. & Egelman, E.H. A conformational change in the actin subunit can change the flexibility of the actin filament. J. Mol. Biol. 232, 334–341 (1993).
Prochniewicz, E., Katayama, E., Yanagida, T. & Thomas, D.D. Cooperativity in F-actin: chemical modifications of actin monomers affect the functional interactions of myosin with unmodified monomers in the same actin filament. Biophys. J. 65, 113–123 (1993).
Drewes, G. & Faulstich, H. Cooperative effects on filament stability in actin modified at the C-terminus by substitution or truncation. Eur. J. Biochem. 212, 247–253 (1993).
Orlova, A., Prochniewicz, E. & Egelman, E.H. Structural dynamics of F-actin. II. Co-operativity in structural transitions. J. Mol. Biol. 245, 598–607 (1995).
Miki, M., Wahl, P. & Auchet, J.-C. Fluorescence anisotropy of labeled F-actin: influence of divalent cations on the interaction between F-actin and myosin heads. Biochemistry 21, 3661–3665 (1982).
Muhlrad, A., Cheung, P., Phan, B., Miller, C. & Reisler, E. Dynamic properties of actin. Structural changes induced by beryllium fluoride. J. Biol. Chem. 269, 11852–11858 (1994).
Oosawa, F., Fujime, S., Ishiwata, S. & Mihashi, K. Dynamic property of F-actin and thin filament. Cold Spring Harb. Symp. Quant. Biol. 37, 277–285 (1973).
Laing, N.G. et al. Mutations and polymorphisms of the skeletal muscle alpha-actin gene (ACTA1). Hum. Mutat. 30, 1267–1277 (2009).
Nowak, K.J. et al. Nemaline myopathy caused by absence of alpha-skeletal muscle actin. Ann. Neurol. 61, 175–184 (2007).
Sparrow, J.C. et al. Muscle disease caused by mutations in the skeletal muscle alpha-actin gene (ACTA1). Neuromuscul. Disord. 13, 519–531 (2003).
Hennessey, E.S., Drummond, D.R. & Sparrow, J.C. Molecular genetics of actin function. Biochem. J. 291, 657–671 (1993).
Feng, J.J. & Marston, S. Genotype-phenotype correlations in ACTA1 mutations that cause congenital myopathies. Neuromuscul. Disord. 19, 6–16 (2009).
Morín, M. et al. In vivo and in vitro effects of two novel gamma-actin (ACTG1) mutations that cause DFNA20/26 hearing impairment. Hum. Mol. Genet. 18, 3075–3089 (2009).
Ilkovski, B. et al. Evidence for a dominant-negative effect in ACTA1 nemaline myopathy caused by abnormal folding, aggregation and altered polymerization of mutant actin isoforms. Hum. Mol. Genet. 13, 1727–1743 (2004).
Chik, J.K., Lindberg, U. & Schutt, C.E. The structure of an open state of β-actin at 2.65 Ångstrom resolution. J. Mol. Biol. 263, 607–623 (1996).
Khaitlina, S.Y. & Strzelecka-Golaszewska, H. Role of the DNase-I-binding loop in dynamic properties of actin filament. Biophys. J. 82, 321–334 (2002).
Kuznetsova, I., Antropova, O., Turoverov, K. & Khaitlina, S. Conformational changes in subdomain I of actin induced by proteolytic cleavage within the DNase I-binding loop: energy transfer from tryptophan to AEDANS. FEBS Lett. 383, 105–108 (1996).
Kuang, B. & Rubenstein, P.A. The effects of severely decreased hydrophobicity in a subdomain 3/4 loop on the dynamics and stability of yeast G-actin. J. Biol. Chem. 272, 4412–4418 (1997).
McKane, M. et al. A mammalian actin substitution in yeast actin (H372R) causes a suppressible mitochondria/vacuole phenotype. J. Biol. Chem. 280, 36494–36501 (2005).
Frank, J. et al. SPIDER and WEB: Processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996).
Ludtke, S.J., Baldwin, P.R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Galkin, V.E., Orlova, A., Cherepanova, O., Lebart, M.C. & Egelman, E.H. High-resolution cryo-EM structure of the F-actin-fimbrin/plastin ABD2 complex. Proc. Natl. Acad. Sci. USA 105, 1494–1498 (2008).
Schröder, G.F., Brunger, A.T. & Levitt, M. Combining efficient conformational sampling with a deformable elastic network model facilitates structure refinement at low resolution. Structure 15, 1630–1641 (2007).
Acknowledgements
This work was supported by a grant from the US National Institutes of Health (NIH GM081303 to E.H.E.).
Author information
Authors and Affiliations
Contributions
A.O. prepared specimens and EM; V.E.G. analyzed images, made classifications, created three-dimensional reconstructions, and built preliminary models; R.S. built and refined models; V.E.G., A.O. and E.H.E. discussed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Methods (PDF 834 kb)
Supplementary Video 1
When the power spectra derived from the raw images of mode 1 and mode 5 classes are superimposed, significant changes in the position and relative intensities of layer line maxima are evident. (GIF 89 kb)
Rights and permissions
About this article
Cite this article
Galkin, V., Orlova, A., Schröder, G. et al. Structural polymorphism in F-actin. Nat Struct Mol Biol 17, 1318–1323 (2010). https://doi.org/10.1038/nsmb.1930
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1930
This article is cited by
-
Magic angle spinning NMR structure of human cofilin-2 assembled on actin filaments reveals isoform-specific conformation and binding mode
Nature Communications (2022)
-
Bending forces and nucleotide state jointly regulate F-actin structure
Nature (2022)
-
Flagella-like beating of actin bundles driven by self-organized myosin waves
Nature Physics (2022)
-
Structural and functional analysis of actin point mutations leading to nemaline myopathy to elucidate their role in actin function
Biophysical Reviews (2022)
-
Structural analysis of cross α-helical nanotubes provides insight into the designability of filamentous peptide nanomaterials
Nature Communications (2021)