Abstract
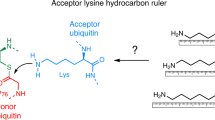
Ubiquitin is a versatile cellular signaling molecule that can form polymers of eight different linkages, and individual linkage types have been associated with distinct cellular functions. Though little is currently known about Lys11-linked ubiquitin chains, recent data indicate that they may be as abundant as Lys48 linkages and may be involved in vital cellular processes. Here we report the generation of Lys11-linked polyubiquitin in vitro, for which the Lys11-specific E2 enzyme UBE2S was fused to a ubiquitin binding domain. Crystallographic and NMR analyses of Lys11-linked diubiquitin reveal that Lys11-linked chains adopt compact conformations in which Ile44 is solvent exposed. Furthermore, we identify the OTU family deubiquitinase Cezanne as the first deubiquitinase with Lys11-linkage preference. Our data highlight the intrinsic specificity of the ubiquitin system that extends to Lys11-linked chains and emphasize that differentially linked polyubiquitin chains must be regarded as independent post-translational modifications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Komander, D. The emerging complexity of protein ubiquitination. Biochem. Soc. Trans. 37, 937–953 (2009).
Chen, Z.J. & Sun, L.J. Nonproteolytic functions of ubiquitin in cell signaling. Mol. Cell 33, 275–286 (2009).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Xu, P. et al. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 137, 133–145 (2009).
Peng, J. et al. A proteomics approach to understanding protein ubiquitination. Nat. Biotechnol. 21, 921–926 (2003).
Dye, B.T. & Schulman, B.A. Structural mechanisms underlying posttranslational modification by ubiquitin-like proteins. Annu. Rev. Biophys. Biomol. Struct. 36, 131–150 (2007).
Ye, Y. & Rape, M. Building ubiquitin chains: E2 enzymes at work. Nat. Rev. Mol. Cell Biol. 10, 755–764 (2009).
Hofmann, R.M. & Pickart, C.M. Noncanonical MMS2-encoded ubiquitin-conjugating enzyme functions in assembly of novel polyubiquitin chains for DNA repair. Cell 96, 645–653 (1999).
Chen, Z. & Pickart, C.M. A 25-kilodalton ubiquitin carrier protein (E2) catalyzes multi-ubiquitin chain synthesis via lysine 48 of ubiquitin. J. Biol. Chem. 265, 21835–21842 (1990).
Baboshina, O.V. & Haas, A.L. Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J. Biol. Chem. 271, 2823–2831 (1996).
Jin, L., Williamson, A., Banerjee, S., Philipp, I. & Rape, M. Mechanism of ubiquitin-chain formation by the human anaphase-promoting complex. Cell 133, 653–665 (2008).
Williamson, A. et al. Identification of a physiological E2 module for the human anaphase-promoting complex. Proc. Natl. Acad. Sci. USA 106, 18213–18218 (2009).
Garnett, M.J. et al. UBE2S elongates ubiquitin chains on APC/C substrates to promote mitotic exit. Nat. Cell Biol. 11, 1363–1369 (2009).
Wu, T. et al. UBE2S drives elongation of K11-linked ubiquitin chains by the anaphase-promoting complex. Proc. Natl. Acad. Sci. USA 107, 1355–1360 (2010).
Rodrigo-Brenni, M.C. & Morgan, D.O. Sequential E2s drive polyubiquitin chain assembly on APC targets. Cell 130, 127–139 (2007).
Alexandru, G. et al. UBXD7 binds multiple ubiquitin ligases and implicates p97 in HIF1alpha turnover. Cell 134, 804–816 (2008).
Eddins, M.J., Carlile, C.M., Gomez, K.M., Pickart, C.M. & Wolberger, C. Mms2-Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nat. Struct. Mol. Biol. 13, 915–920 (2006).
Reyes-Turcu, F.E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208 (2006).
McCullough, J., Clague, M.J. & Urbe, S. AMSH is an endosome-associated ubiquitin isopeptidase. J. Cell Biol. 166, 487–492 (2004).
Varadan, R. et al. Solution conformation of Lys63-linked di-ubiquitin chain provides clues to functional diversity of polyubiquitin signaling. J. Biol. Chem. 279, 7055–7063 (2004).
Varadan, R., Walker, O., Pickart, C. & Fushman, D. Structural properties of polyubiquitin chains in solution. J. Mol. Biol. 324, 637–647 (2002).
Tenno, T. et al. Structural basis for distinct roles of Lys63- and Lys48-linked polyubiquitin chains. Genes Cells 9, 865–875 (2004).
Zhang, N. et al. Structure of the s5a:k48-linked diubiquitin complex and its interactions with rpn13. Mol. Cell 35, 280–290 (2009).
Varadan, R., Assfalg, M., Raasi, S., Pickart, C. & Fushman, D. Structural determinants for selective recognition of a Lys48-linked polyubiquitin chain by a UBA domain. Mol. Cell 18, 687–698 (2005).
Raasi, S., Varadan, R., Fushman, D. & Pickart, C.M. Diverse polyubiquitin interaction properties of ubiquitin-associated domains. Nat. Struct. Mol. Biol. 12, 708–714 (2005).
Komander, D., Clague, M.J. & Urbe, S. Breaking the chains: structure and function of the deubiquitinases. Nat. Rev. Mol. Cell Biol. 10, 550–563 (2009).
Komander, D. et al. Molecular discrimination of structurally equivalent Lys 63-linked and linear polyubiquitin chains. EMBO Rep. 10, 466–473 (2009).
Komander, D. et al. The structure of the CYLD USP domain explains its specificity for Lys63-linked polyubiquitin and reveals a B box module. Mol. Cell 29, 451–464 (2008).
Tran, H., Hamada, F., Schwarz-Romond, T. & Bienz, M. Trabid, a new positive regulator of Wnt-induced transcription with preference for binding and cleaving K63-linked ubiquitin chains. Genes Dev. 22, 528–542 (2008).
Komander, D. & Barford, D. Structure of the A20 OTU domain and mechanistic insights into deubiquitination. Biochem. J. 409, 77–85 (2008).
Edelmann, M.J. et al. Structural basis and specificity of human otubain 1-mediated deubiquitination. Biochem. J. 418, 379–390 (2009).
Winborn, B.J. et al. The deubiquitinating enzyme ataxin-3, a polyglutamine disease protein, edits Lys63 linkages in mixed linkage ubiquitin chains. J. Biol. Chem. 283, 26436–26443 (2008).
Hymowitz, S.G. & Wertz, I.E. A20: from ubiquitin editing to tumour suppression. Nat. Rev. Cancer 10, 332–341 (2010).
Skaug, B., Jiang, X. & Chen, Z.J. The role of ubiquitin in NF-κB regulatory pathways. Annu. Rev. Biochem. 78, 769–796 (2009).
Enesa, K. et al. NF-κB suppression by the deubiquitinating enzyme Cezanne: a novel negative feedback loop in pro-inflammatory signaling. J. Biol. Chem. 283, 7036–7045 (2008).
Fushman, D. & Walker, O. Exploring the linkage dependence of polyubiquitin conformations using molecular modeling. J. Mol. Biol. 395, 803–814 (2010).
van Wijk, S.J. et al. A comprehensive framework of E2-RING E3 interactions of the human ubiquitin-proteasome system. Mol. Syst. Biol. 5, 295 (2009).
Markson, G. et al. Analysis of the human E2 ubiquitin conjugating enzyme protein interaction network. Genome Res. 19, 1905–1911 (2009).
Xia, Z.P. et al. Direct activation of protein kinases by unanchored polyubiquitin chains. Nature 461, 114–119 (2009).
Nijman, S.M. et al. A genomic and functional inventory of deubiquitinating enzymes. Cell 123, 773–786 (2005).
Neidhardt, F.C., Bloch, P.L. & Smith, D.F. Culture medium for enterobacteria. J. Bacteriol. 119, 736–747 (1974).
Pickart, C.M. & Raasi, S. Controlled synthesis of polyubiquitin chains. Methods Enzymol. 399, 21–36 (2005).
Mori, S., Abeygunawardana, C., Johnson, M.O. & van Zijl, P.C. Improved sensitivity of HSQC spectra of exchanging protons at short interscan delays using a new fast HSQC (FHSQC) detection scheme that avoids water saturation. J. Magn. Reson. B. 108, 94–98 (1995).
Hajduk, P.J. et al. NMR-based discovery of lead inhibitors that block DNA binding of the human papillomavirus E2 protein. J. Med. Chem. 40, 3144–3150 (1997).
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006).
Vagin, A. & Teplyakov, A. An approach to multi-copy search in molecular replacement. Acta Crystallogr. D Biol. Crystallogr. 56, 1622–1624 (2000).
Vijay-Kumar, S., Bugg, C.E. & Cook, W.J. Structure of ubiquitin refined at 1.8 Å resolution. J. Mol. Biol. 194, 531–544 (1987).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Acknowledgements
We would like to thank E. Stephens, S. Peak-Chew and F. Begum for MS analysis, M. Allen for help with isotope labeling, T. Nicholson (ENZO LifeSciences) for providing ubiquitin mutants, P. Cohen (Univ. of Dundee), S. Urbe, M. Clague (Univ. of Liverpool), K.D. Wilkinson (Emory Univ.), S. Todi (Univ. Of Michigan) and B. Kessler (Oxford Univ.) for constructs, T. Tenno (Kobe Univ.) and M. Shirakawa (Kyoto Univ.) for kindly providing NMR data for Lys48- and Lys63-linked diubiquitin and members of the Komander lab, especially Y. Kulathu, Y. Ye, M. Akutsu, S. Virdee and A. Eslambolchi for providing reagents and many discussions. A.B. is a MRC Career Development Fellow.
Author information
Authors and Affiliations
Contributions
A.B. designed, performed and analyzed all experiments; S.M.V.F. performed solution measurements and analyzed the data; D.K. designed research, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Table 1, Supplementary Discussion, Supplementary Methods and Supplementary Data (PDF 5381 kb)
Rights and permissions
About this article
Cite this article
Bremm, A., Freund, S. & Komander, D. Lys11-linked ubiquitin chains adopt compact conformations and are preferentially hydrolyzed by the deubiquitinase Cezanne. Nat Struct Mol Biol 17, 939–947 (2010). https://doi.org/10.1038/nsmb.1873
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1873
This article is cited by
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Deubiquitinases in cancer
Nature Reviews Cancer (2023)
-
A structural basis for the diverse linkage specificities within the ZUFSP deubiquitinase family
Nature Communications (2022)
-
A widely distributed family of eukaryotic and bacterial deubiquitinases related to herpesviral large tegument proteins
Nature Communications (2022)
-
Beyond K48 and K63: non-canonical protein ubiquitination
Cellular & Molecular Biology Letters (2021)