Abstract
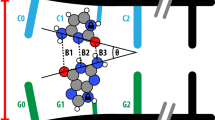
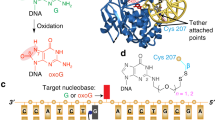
The major product of oxidative base damage is 8-oxo-7,8-dihydro-2′-deoxyguanine (8odG). The coding potential of this lesion is modulated by its glycosidic torsion angle that controls whether its Watson-Crick or Hoogsteen edge is used for base pairing. The 2.0-Å structure of DNA polymerase (pol) β bound with 8odGTP opposite template adenine indicates that the modified nucleotide assumes the mutagenic syn conformation and that the nonmutagenic anti conformation would be incompatible with efficient DNA synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Ames, B.N. & Gold, L.S. Mutat. Res. 250, 3–16 (1991).
Kouchakdjian, M. et al. Biochemistry 30, 1403–1412 (1991).
McAuley-Hecht, K.E. et al. Biochemistry 33, 10266–10270 (1994).
Maki, H. & Sekiguchi, M. Nature 355, 273–275 (1992).
Beard, W.A. & Wilson, S.H. Chem. Rev. 106, 361–382 (2006).
Batra, V.K., Beard, W.A., Shock, D.D., Pedersen, L.C. & Wilson, S.H. Mol. Cell 30, 315–324 (2008).
Krahn, J.M., Beard, W.A. & Wilson, S.H. Structure 12, 1823–1832 (2004).
Miller, H., Prasad, R., Wilson, S.H., Johnson, F. & Grollman, A.P. Biochemistry 39, 1029–1033 (2000).
Krahn, J.M., Beard, W.A., Miller, H., Grollman, A.P. & Wilson, S.H. Structure 11, 121–127 (2003).
Batra, V.K. et al. Structure 14, 757–766 (2006).
Brown, J.A., Duym, W.W., Fowler, J.D. & Suo, Z. J. Mol. Biol. 367, 1258–1269 (2007).
Sucato, C.A. et al. Biochemistry 46, 461–471 (2007).
Culp, S.J., Cho, B.P., Kadlubar, F.F. & Evans, F.E. Chem. Res. Toxicol. 2, 416–422 (1989).
Uesugi, S. & Ikehara, M. J. Am. Chem. Soc. 99, 3250–3253 (1977).
Hsu, G.W., Ober, M., Carell, T. & Beese, L.S. Nature 431, 217–221 (2004).
Rechkoblit, O. et al. PLoS Biol. 4, e11 (2006).
Beard, W.A. & Wilson, S.H. Structure 11, 489–496 (2003).
Acknowledgements
This research was supported by Research Project Number Z01-ES050158 in the Intramural Research Program of the US National Institutes of Health, US National Institute of Environmental Health Sciences and was in association with the US National Institutes of Health Grant 1U19CA105010.
Author information
Authors and Affiliations
Contributions
E.W.H. and R.P. expressed and purified protein; L.C.P. contributed to structure refinement; V.K.B. prepared, crystallized and solved the structure of the 8odGTP–dA pol β complex; V.K.B., W.A.B. and S.H.W. analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1–3 and Supplementary Methods (PDF 3685 kb)
Rights and permissions
About this article
Cite this article
Batra, V., Beard, W., Hou, E. et al. Mutagenic conformation of 8-oxo-7,8-dihydro-2′-dGTP in the confines of a DNA polymerase active site. Nat Struct Mol Biol 17, 889–890 (2010). https://doi.org/10.1038/nsmb.1852
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1852
This article is cited by
-
Watching a double strand break repair polymerase insert a pro-mutagenic oxidized nucleotide
Nature Communications (2021)
-
Outlier analyses of the Protein Data Bank archive using a probability-density-ranking approach
Scientific Data (2018)
-
How DNA polymerases catalyse replication and repair with contrasting fidelity
Nature Reviews Chemistry (2017)
-
Oxidized nucleotide insertion by pol β confounds ligation during base excision repair
Nature Communications (2017)
-
The 8-oxo-dGTP interaction with human DNA polymerase β: two patterns of ligand behavior
Structural Chemistry (2016)