Abstract
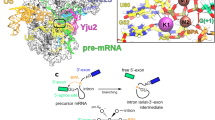
Spliceosomes catalyze the maturation of precursor mRNAs in organisms ranging from yeast to humans. Their catalytic core comprises three small nuclear RNAs (U2, U5 and U6) involved in substrate positioning and catalysis. It has been postulated, but never shown experimentally, that the U2–U6 complex adopts at least two conformations that reflect different activation states. We have used single-molecule fluorescence to probe the structural dynamics of a protein-free RNA complex modeling U2–U6 from yeast and mutants of highly conserved regions of U2–U6. Our data show the presence of at least three distinct conformations in equilibrium. The minimal folding pathway consists of a two-step process with an obligatory intermediate. The first step is strongly magnesium dependent, and we provide evidence suggesting that the second step corresponds to the formation of the genetically conserved helix IB. Site-specific mutations in the highly conserved AGC triad and the U80 base in U6 suggest that the observed conformational dynamics correlate with residues that have an important role in splicing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kalnina, Z., Zayakin, P., Silina, K. & Line, A. Alterations of pre-mRNA splicing in cancer. Genes Chromosom. Cancer 42, 342–357 (2005).
Licatalosi, D.D. & Darnell, R.B. Splicing regulation in neurologic disease. Neuron 52, 93–101 (2006).
Stark, H. & Luhrmann, R. Cryo-electron microscopy of spliceosomal components. Annu. Rev. Biophys. Biomol. Struct. 35, 435–457 (2006).
Black, D.L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Brow, D.A. Allosteric cascade of spliceosome activation. Annu. Rev. Genet. 36, 333–360 (2002).
Parker, R., Siliciano, P.G. & Guthrie, C. Recognition of the TACTAAC box during mRNA splicing in yeast involves base pairing to the U2-like snRNA. Cell 49, 229–239 (1987).
Lesser, C.F. & Guthrie, C. Mutations in U6 snRNA that alter splice site specificity: implications for the active site. Science 262, 1982–1988 (1993).
O'Keefe, R.T., Norman, C. & Newman, A.J. The invariant U5 snRNA loop 1 sequence is dispensable for the first catalytic step of pre-mRNA splicing in yeast. Cell 86, 679–689 (1996).
Ségault, V. et al. Conserved loop I of U5 small nuclear RNA is dispensable for both catalytic steps of pre-mRNA splicing in HeLa nuclear extracts. Mol. Cell. Biol. 19, 2782–2790 (1999).
Valadkhan, S. & Manley, J.L. Splicing-related catalysis by protein-free snRNAs. Nature 413, 701–707 (2001).
Valadkhan, S., Mohammadi, A., Jaladat, Y. & Geisler, S. Protein-free small nuclear RNAs catalyze a two-step splicing reaction. Proc. Natl. Acad. Sci. USA 106, 11901–11906 (2009).
Sontheimer, E.J., Gordon, P.M. & Piccirilli, J.A. Metal ion catalysis during group II intron self-splicing: parallels with the spliceosome. Genes Dev. 13, 1729–1741 (1999).
Steiner, M., Karunatilaka, K.S., Sigel, R.K. & Rueda, D. Single-molecule studies of group II intron ribozymes. Proc. Natl. Acad. Sci. USA 105, 13853–13858 (2008).
Toor, N., Keating, K.S., Taylor, S.D. & Pyle, A.M. Crystal structure of a self-spliced group II intron. Science 320, 77–82 (2008).
Abelson, J. Is the spliceosome a ribonucleoprotein enzyme? Nat. Struct. Mol. Biol. 15, 1235–1237 (2008).
Madhani, H.D. & Guthrie, C. A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell 71, 803–817 (1992).
Hilliker, A.K. & Staley, J.P. Multiple functions for the invariant AGC triad of U6 snRNA. RNA 10, 921–928 (2004).
Mefford, M.A. & Staley, J.P. Evidence that U2/U6 helix I promotes both catalytic steps of pre-mRNA splicing and rearranges in between these steps. RNA 15, 1386–1397 (2009).
Sashital, D.G., Cornilescu, G., McManus, C.J., Brow, D.A. & Butcher, S.E. U2–U6 RNA folding reveals a group II intron-like domain and a four-helix junction. Nat. Struct. Mol. Biol. 11, 1237–1242 (2004).
Sun, J.S. & Manley, J.L. A novel U2–U6 snRNA structure is necessary for mammalian mRNA splicing. Genes Dev. 9, 843–854 (1995).
Rhode, B.M., Hartmuth, K., Westhof, E. & Luhrmann, R. Proximity of conserved U6 and U2 snRNA elements to the 5′ splice site region in activated spliceosomes. EMBO J. 25, 2475–2486 (2006).
Zhao, R. & Rueda, D. RNA folding dynamics by single-molecule fluorescence resonance energy transfer. Methods 10.1016/j.ymeth.2009.04.017 (2009).
Hershey, J.W.B. & Merrick, W.C. The pathway and mechanism of initiation of protein synthesis. in Translational Control of Gene Expression (eds. Sonenberg, N., Hershey, J.W.B. & Mathews, M.B.) 48 (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2000).
Fabrizio, P. & Abelson, J. Thiophosphates in yeast U6 snRNA specifically affect pre-mRNA splicing in vitro. Nucleic Acids Res. 20, 3659–3664 (1992).
Sontheimer, E.J., Sun, S. & Piccirilli, J.A. Metal ion catalysis during splicing of premessenger RNA. Nature 388, 801–805 (1997).
Yean, S.L., Wuenschell, G., Termini, J. & Lin, R.J. Metal-ion coordination by U6 small nuclear RNA contributes to catalysis in the spliceosome. Nature 408, 881–884 (2000).
Huppler, A., Nikstad, L.J., Allmann, A.M., Brow, D.A. & Butcher, S.E. Metal binding and base ionization in the U6 RNA intramolecular stem-loop structure. Nat. Struct. Biol. 9, 431–435 (2002).
Yuan, F. et al. Use of a novel Forster resonance energy transfer method to identify locations of site-bound metal ions in the U2–U6 snRNA complex. Nucleic Acids Res. 35, 2833–2845 (2007).
Alemán, E.A., Lamichhane, R. & Rueda, D. Exploring RNA folding one molecule at a time. Curr. Opin. Chem. Biol. 12, 647–654 (2008).
Nir, E. et al. Shot-noise limited single-molecule FRET histograms: comparison between theory and experiments. J. Phys. Chem. B 110, 22103–22124 (2006).
Kobitski, A.Y., Nierth, A., Helm, M., Jaschke, A. & Nienhaus, G.U. Mg2+-dependent folding of a Diels-Alderase ribozyme probed by single-molecule FRET analysis. Nucleic Acids Res. 35, 2047–2059 (2007).
McKinney, S.A., Freeman, A.D., Lilley, D.M. & Ha, T. Observing spontaneous branch migration of Holliday junctions one step at a time. Proc. Natl. Acad. Sci. USA 102, 5715–5720 (2005).
Sashital, D.G., Allmann, A.M., Van Doren, S.R. & Butcher, S.E. Structural basis for a lethal mutation in U6 RNA. Biochemistry 42, 1470–1477 (2003).
Reiter, N.J., Nikstad, L.J., Allmann, A.M., Johnson, R.J. & Butcher, S.E. Structure of the U6 RNA intramolecular stem-loop harboring an S(P)-phosphorothioate modification. RNA 9, 533–542 (2003).
Moore, M.J. & Sharp, P.A. Evidence for two active sites in the spliceosome provided by stereochemistry of pre-mRNA splicing. Nature 365, 364–368 (1993).
Gordon, P.M., Sontheimer, E.J. & Piccirilli, J.A. Metal ion catalysis during the exon-ligation step of nuclear pre-mRNA splicing: extending the parallels between the spliceosome and group II introns. RNA 6, 199–205 (2000).
Sontheimer, E.J. The spliceosome shows its metal. Nat. Struct. Biol. 8, 11–13 (2001).
Sun, J.S. & Manley, J.L. The human U6 snRNA intramolecular helix: structural constraints and lack of sequence specificity. RNA 3, 514–526 (1997).
Rueda, D. & Walter, N.G. Fluorescent energy transfer readout of an aptazyme-based biosensor. Methods Mol. Biol. 335, 289–310 (2006).
Rueda, D., Wick, K., McDowell, S.E. & Walter, N.G. Diffusely bound Mg2+ ions slightly reorient stems I and II of the hammerhead ribozyme to increase the probability of formation of the catalytic core. Biochemistry 42, 9924–9936 (2003).
Acknowledgements
We would like to thank S. Butcher and D. Brow for many helpful and stimulating discussions and for commenting on the manuscript. This work was supported by the NIH (R01 GM085116) and an NSF CAREER award to D.R. (MCB-0747285).
Author information
Authors and Affiliations
Contributions
Z.G. performed experiments, analyzed data and wrote the manuscript; K.S.K. performed experiments and edited the manuscript; D.R. designed experiments and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 981 kb)
Rights and permissions
About this article
Cite this article
Guo, Z., Karunatilaka, K. & Rueda, D. Single-molecule analysis of protein-free U2–U6 snRNAs. Nat Struct Mol Biol 16, 1154–1159 (2009). https://doi.org/10.1038/nsmb.1672
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1672
This article is cited by
-
Impact of bisphenol-A on the spliceosome and meiosis of sperm in the testis of adolescent mice
BMC Veterinary Research (2022)
-
Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells
Nature Communications (2018)
-
Genome-wide identification and functional prediction of novel and fungi-responsive lincRNAs in Triticum aestivum
BMC Genomics (2016)