Abstract
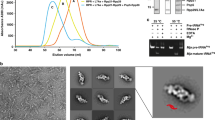
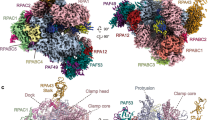
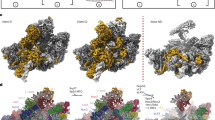
Box H/ACA small nucleolar and Cajal body ribonucleoprotein particles comprise the most complex pseudouridine synthases and are essential for ribosome and spliceosome maturation. The multistep and multicomponent-mediated enzyme mechanism remains only partially understood. Here we report a crystal structure at 2.35 Å of a substrate-bound functional archaeal enzyme containing three of the four proteins, Cbf5, Nop10 and L7Ae, and a box H/ACA RNA that reveals detailed information about the protein-only active site. The substrate RNA, containing 5-fluorouridine at the modification position, is fully docked and catalytically rearranged by the enzyme in a manner similar to that seen in two stand-alone pseudouridine synthases. Structural analysis provides a mechanism for plasticity in the diversity of guide RNA sequences used and identifies a substrate-anchoring loop of Cbf5 that also interacts with Gar1 in unliganded structures. Activity analyses of mutated proteins and RNAs support the structural findings and further suggest a role of the Cbf5 loop in regulation of enzyme activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Maden, B.E. The numerous modified nucleotides in eukaryotic ribosomal RNA. Prog. Nucleic Acid Res. Mol. Biol. 39, 241–303 (1990).
Limbach, P.A., Crain, P.F. & McCloskey, J.A. Summary: the modified nucleosides of RNA. Nucleic Acids Res. 22, 2183–2196 (1994).
Grosjean, H. & Benne, R. Modification and editing of RNA (ASM, Washington, DC, 1998).
Charette, M. & Gray, M.W. Pseudouridine in RNA: what, where, how, and why. IUBMB Life 49, 341–351 (2000).
Ofengand, J. Ribosomal RNA pseudouridines and pseudouridine synthases. FEBS Lett. 514, 17–25 (2002).
Koonin, E.V. Pseudouridine synthases: four families of enzymes containing a putative uridine-binding motif also conserved in dUTPases and dCTP deaminases. Nucleic Acids Res. 24, 2411–2415 (1996).
Kaya, Y. & Ofengand, J. A novel unanticipated type of pseudouridine synthase with homologs in bacteria, archaea, and eukarya. RNA 9, 711–721 (2003).
Decatur, W.A. & Fournier, M.J. RNA-guided nucleotide modification of ribosomal and other RNAs. J. Biol. Chem. 278, 695–698 (2003).
Ganot, P., Bortolin, M.L. & Kiss, T. Site-specific pseudouridine formation in preribosomal RNA is guided by small nucleolar RNAs. Cell 89, 799–809 (1997).
Meier, U.T. The many facets of H/ACA ribonucleoproteins. Chromosoma 114, 1–14 (2005).
Yu, Y.T., Terns, R.M. & Terns, M.P. Fine-tuning of RNA functions by modification and editing. in Topics in Current Genetics (ed. Grosjean, H.) 223–262 (Springer, New York, USA, 2005).
Balakin, A.G., Smith, L. & Fournier, M.J. The RNA world of the nucleolus: two major families of small RNAs defined by different box elements with related functions. Cell 86, 823–834 (1996).
Ganot, P., Caizergues-Ferrer, M. & Kiss, T. The family of box ACA small nucleolar RNAs is defined by an evolutionarily conserved secondary structure and ubiquitous sequence elements essential for RNA accumulation. Genes Dev. 11, 941–956 (1997).
Watkins, N.J. et al. Cbf5p, a potential pseudouridine synthase, and Nhp2p, a putative RNA-binding protein, are present together with Gar1p in all H BOX/ACA-motif snoRNPs and constitute a common bipartite structure. RNA 4, 1549–1568 (1998).
Henras, A. et al. Nhp2p and Nop10p are essential for the function of H/ACA snoRNPs. EMBO J. 17, 7078–7090 (1998).
Lafontaine, D.L., Bousquet, A.C., Henry,Y., Caizergues, F.M. & Tollervey, D. The box H+ ACA snoRNAs carry Cbf5p, the putative rRNA pseudouridine synthase. Genes Dev. 12, 527–537 (1998).
Morrissey, J.P. & Tollervey, D. Yeast snR30 is a small nucleolar RNA required for 18S rRNA synthesis. Mol. Cell. Biol. 13, 2469–2477 (1993).
Eliceiri, G.L. The vertebrate E1/U17 small nucleolar ribonucleoprotein particle. J. Cell. Biochem. 98, 486–495 (2006).
Mitchell, J.R., Cheng, J. & Collins, K. A box H/ACA small nucleolar RNA-like domain at the human telomerase RNA 3′ end. Mol. Cell. Biol. 19, 567–576 (1999).
Collins, K. The biogenesis and regulation of telomerase holoenzymes. Nat. Rev. Mol. Cell Biol. 7, 484–494 (2006).
Lukowiak, A.A., Narayanan, A., Li, Z.H., Terns, R.M. & Terns, M.P. The snoRNA domain of vertebrate telomerase RNA functions to localize the RNA within the nucleus. RNA 7, 1833–1844 (2001).
Heiss, N.S. et al. X-linked dyskeratosis congenita is caused by mutations in a highly conserved gene with putative nucleolar functions. Nat. Genet. 19, 32–38 (1998).
Walne, A.J. et al. Genetic heterogeneity in autosomal recessive dyskeratosis congenita with one subtype due to mutations in the telomerase-associated protein NOP10. Hum. Mol. Genet. 16, 1619–1629 (2007).
Vulliamy, T. et al. Mutations in the telomerase component NHP2 cause the premature ageing syndrome dyskeratosis congenita. Proc. Natl. Acad. Sci. USA 105, 8073–8078 (2008).
Spedaliere, C.J., Ginter, J.M., Johnston, M.V. & Mueller, E.G. The pseudouridine synthases: revisiting a mechanism that seemed settled. J. Am. Chem. Soc. 126, 12758–12759 (2004).
Hamma, T. & Ferre-D'Amare, A.R. Pseudouridine synthases. Chem. Biol. 13, 1125–1135 (2006).
Phannachet, K., Elias, Y. & Huang, R.H. Dissecting the roles of a strictly conserved tyrosine in substrate recognition and catalysis by pseudouridine 55 synthase. Biochemistry 44, 15488–15494 (2005).
Roovers, M. et al. Formation of the conserved pseudouridine at position 55 in archaeal tRNA. Nucleic Acids Res. 34, 4293–4301 (2006).
Schattner, P. et al. Genome-wide searching for pseudouridylation guide snoRNAs: analysis of the Saccharomyces cerevisiae genome. Nucleic Acids Res. 32, 4281–4296 (2004).
Decatur, W.A. & Fournier, M.J. rRNA modifications and ribosome function. Trends Biochem. Sci. 27, 344–351 (2002).
Baker, D.L. et al. RNA-guided RNA modification: functional organization of the archaeal H/ACA RNP. Genes Dev. 19, 1238–1248 (2005).
Wang, C. & Meier, U.T. Architecture and assembly of mammalian H/ACA small nucleolar and telomerase ribonucleoproteins. EMBO J. 23, 1857–1867 (2004).
Charpentier, B., Muller, S. & Branlant, C. Reconstitution of archaeal H/ACA small ribonucleoprotein complexes active in pseudouridylation. Nucleic Acids Res. 33, 3133–3144 (2005).
Li, L. & Ye, K. Crystal structure of an H/ACA box ribonucleoprotein particle. Nature 443, 302–307 (2006).
Li, H. Unveiling substrate RNA binding to H/ACA RNPs: one side fits all. Curr. Opin. Struct. Biol. 18, 78–85 (2008).
Liang, B., Xue, S., Terns, R.M., Terns, M.P. & Li, H. Substrate RNA positioning in the archaeal H/ACA ribonucleoprotein complex. Nat. Struct. Mol. Biol. 14, 1189–1195 (2007).
Rashid, R. et al. Crystal structure of a Cbf5-Nop10-Gar1 complex and implications in RNA-guided pseudouridylation and dyskeratosis congenita. Mol. Cell 21, 249–260 (2006).
Ye, K. H/ACA guide RNAs, proteins and complexes. Curr. Opin. Struct. Biol. 17, 287–292 (2007).
Hamma, T., Reichow, S.L., Varani, G. & Ferre-D'Amare, A.R. The Cbf5-Nop10 complex is a molecular bracket that organizes box H/ACA RNPs. Nat. Struct. Mol. Biol. 12, 1101–1107 (2005).
Manival, X. et al. Crystal structure determination and site-directed mutagenesis of the Pyrococcus abyssi aCBF5-aNOP10 complex reveal crucial roles of the C-terminal domains of both proteins in H/ACA sRNP activity. Nucleic Acids Res. 34, 826–839 (2006).
Liang, B. et al. Long-distance placement of substrate RNA by H/ACA proteins. RNA 14, 2086–2094 (2008).
Klein, R.J., Misulovin, Z. & Eddy, S.R. Noncoding RNA genes identified in AT-rich hyperthermophiles. Proc. Natl. Acad. Sci. USA 99, 7542–7547 (2002).
Rozhdestvensky, T.S. et al. Binding of L7Ae protein to the K-turn of archaeal snoRNAs: a shared RNA binding motif for C/D and H/ACA box snoRNAs in Archaea. Nucleic Acids Res. 31, 869–877 (2003).
Wu, H. & Feigon, J. H/ACA small nucleolar RNA pseudouridylation pockets bind substrate RNA to form three-way junctions that position the target U for modification. Proc. Natl. Acad. Sci. USA 104, 6655–6660 (2007).
Jin, H., Loria, J.P. & Moore, P.B. Solution structure of an rRNA substrate bound to the pseudouridylation pocket of a box H/ACA snoRNA. Mol. Cell 26, 205–215 (2007).
Hoang, C. & Ferre-D'Amare, A.R. Cocrystal structure of a tRNA Psi55 pseudouridine synthase: nucleotide flipping by an RNA-modifying enzyme. Cell 107, 929–939 (2001).
Spedaliere, C.J. & Mueller, E.G. Not all pseudouridine synthases are potently inhibited by RNA containing 5-fluorouridine. RNA 10, 192–199 (2004).
Hoang, C. et al. Crystal structure of pseudouridine synthase RluA: indirect sequence readout through protein-induced RNA structure. Mol. Cell 24, 535–545 (2006).
Baker, D.L. et al. Determination of protein-rna interaction sites in the Cbf5-H/ACA Guide RNA complex by mass spectrometric protein footprinting. Biochemistry 47, 1500–1510 (2008).
Muller, S., Fourmann, J.B., Loegler, C., Charpentier, B. & Branlant, C. Identification of determinants in the protein partners aCBF5 and aNOP10 necessary for the tRNA:{Psi}55-synthase and RNA-guided RNA:{Psi}-synthase activities. Nucleic Acids Res. 35, 5610–5624 (2007).
Darzacq, X. et al. Stepwise RNP assembly at the site of H/ACA RNA transcription in human cells. J. Cell Biol. 173, 207–218 (2006).
McKenna, S.A. et al. Purification and characterization of transcribed RNAs using gel filtration chromatography. Nat. Protocols 2, 3270–3277 (2007).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Murshudov, G.N., Vagin, A.A., Lebedev, A., Wilson, K.S. & Dodson, E.J. Efficient anisotropic refinement of macromolecular structures using FFT. Acta Crystallogr. D Biol. Crystallogr. 55, 247–255 (1999).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for binding protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Winn, M.D., Isupov, M.N. & Murshudov, G.N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 57, 122–133 (2001).
Acknowledgements
This work was supported in part by the US National Institutes of Health grants R01 GM66958-01 (H.L.) and R01 GM54682 (M.T. and R.T.). B.L. (0815118E) and J.Z. (0815267E) are supported by the American Heart Association, Greater Southeast Affiliate. X-ray diffraction data were collected by the Southeast Regional Collaborative Access Team (SER-CAT) 22-ID beamline at the Advanced Photon Source (APS), Argonne National Laboratory. Supporting institutions for APS beamlines may be found at http://necat.chem.cornell.edu and http://www.ser-cat.org/members.html. Use of the APS was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. W-31-109-Eng-38.
Author information
Authors and Affiliations
Contributions
B.L. determined the crystal structure; J.Z. prepared samples; E.K. performed fluorescence assays; M.P.T. and R.M.T. prepared the manuscript; H.L. supervised the project and prepared the manuscript.
Corresponding author
Supplementary information
Supplementary Figures and Tables
Supplementary Tables 1 and 2 and Supplementary Figures 1 and 2 (PDF 1194 kb)
Rights and permissions
About this article
Cite this article
Liang, B., Zhou, J., Kahen, E. et al. Structure of a functional ribonucleoprotein pseudouridine synthase bound to a substrate RNA. Nat Struct Mol Biol 16, 740–746 (2009). https://doi.org/10.1038/nsmb.1624
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1624
This article is cited by
-
The Fate of Duplicated Enzymes in Prokaryotes: The Case of Isomerases
Journal of Molecular Evolution (2023)
-
Contribution of protein Gar1 to the RNA-guided and RNA-independent rRNA:Ψ-synthase activities of the archaeal Cbf5 protein
Scientific Reports (2018)
-
RNA-dependent pseudouridylation catalyzed by box H/ACA RNPs
Frontiers in Biology (2018)
-
Quantum chemical calculations support pseudouridine synthase reaction through a glycal intermediate and provide details of the mechanism
Theoretical Chemistry Accounts (2018)
-
Crystal structure of an ASCH protein from Zymomonas mobilis and its ribonuclease activity specific for single-stranded RNA
Scientific Reports (2017)