Abstract
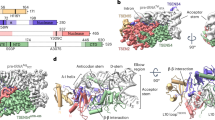
Muscleblind-like (MBNL) proteins, regulators of developmentally programmed alternative splicing, harbor tandem CCCH zinc-finger (ZnF) domains that target pre-mRNAs containing YGCU(U/G)Y sequence elements (where Y is a pyrimidine). In myotonic dystrophy, reduced levels of MBNL proteins lead to aberrant alternative splicing of a subset of pre-mRNAs. The crystal structure of MBNL1 ZnF3/4 bound to r(CGCUGU) establishes that both ZnF3 and ZnF4 target GC steps, with site-specific recognition mediated by a network of hydrogen bonds formed primarily with main chain groups of the protein. The relative alignment of ZnF3 and ZnF4 domains is dictated by the topology of the interdomain linker, with a resulting antiparallel orientation of bound GC elements, supportive of a chain-reversal loop trajectory for MBNL1-bound pre-mRNA targets. We anticipate that MBNL1-mediated targeting of looped RNA segments proximal to splice-site junctions could contribute to pre-mRNA alternative-splicing regulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Referenced accessions
GenBank/EMBL/DDBJ
References
Black, D.L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003).
Pascual, M., Vicente, M., Monferrer, L. & Artero, R. The Muscleblind family of proteins: an emerging class of regulators of developmentally programmed alternative splicing. Differentiation 74, 65–80 (2006).
Ho, T.H. et al. Muscleblind proteins regulate alternative splicing. EMBO J. 23, 3103–3112 (2004).
Lin, X. et al. Failure of MBNL1-dependent post-natal splicing transitions in myotonic dystrophy. Hum. Mol. Genet. 15, 2087–2097 (2006).
Osborne, R.J. & Thornton, C.A. RNA-dominant diseases. Hum. Mol. Genet. 15, R162–R169 (2006).
Ranum, L.P. & Cooper, T.A. RNA-mediated neuromuscular disorders. Annu. Rev. Neurosci. 29, 259–277 (2006).
Brook, J.D. et al. Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 68, 799–808 (1992).
Liquori, C.L. et al. Myotonic dystrophy type 2 caused by a CCTG expansion in intron 1 of ZNF9. Science 293, 864–867 (2001).
Fardaei, M. et al. Three proteins, MBNL, MBLL and MBXL, co-localize in vivo with nuclear foci of expanded-repeat transcripts in DM1 and DM2 cells. Hum. Mol. Genet. 11, 805–814 (2002).
Kino, Y. et al. Muscleblind protein, MBNL1/EXP, binds specifically to CHHG repeats. Hum. Mol. Genet. 13, 495–507 (2004).
Miller, J.W. et al. Recruitment of human muscleblind proteins to (CUG)n expansions associated with myotonic dystrophy. EMBO J. 19, 4439–4448 (2000).
Kanadia, R.N. et al. A muscleblind knockout model for myotonic dystrophy. Science 302, 1978–1980 (2003).
Kanadia, R.N. et al. Reversal of RNA missplicing and myotonia after muscleblind overexpression in a mouse poly(CUG) model for myotonic dystrophy. Proc. Natl. Acad. Sci. USA 103, 11748–11753 (2006).
Paul, S. et al. Interaction of muscleblind, CUG-BP1 and hnRNP H proteins in DM1-associated aberrant IR splicing. EMBO J. 25, 4271–4283 (2006).
Philips, A.V., Timchenko, L.T. & Cooper, T.A. Disruption of splicing regulated by a CUG-binding protein in myotonic dystrophy. Science 280, 737–741 (1998).
Savkur, R.S., Philips, A.V. & Cooper, T.A. Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat. Genet. 29, 40–47 (2001).
Squillace, R.M., Chenault, D.M. & Wang, E.H. Inhibition of muscle differentiation by the novel muscleblind-related protein CHCR. Dev. Biol. 250, 218–230 (2002).
Adereth, Y., Dammai, V., Kose, N., Li, R. & Hsu, T. RNA-dependent integrin α3 protein localization regulated by the Muscleblind-like protein MLP1. Nat. Cell Biol. 7, 1240–1247 (2005).
Hudson, B.P., Martinez-Yamout, M.A., Dyson, H.J. & Wright, P.E. Recognition of the mRNA AU-rich element by the zinc finger domain of TIS11d. Nat. Struct. Mol. Biol. 11, 257–264 (2004).
Varani, L. et al. The NMR structure of the 38 kDa U1A protein – PIE RNA complex reveals the basis of cooperativity in regulation of polyadenylation by human U1A protein. Nat. Struct. Biol. 7, 329–335 (2000).
Kielkopf, C.L., Rodionova, N.A., Green, M.R. & Burley, S.K. A novel peptide recognition mode revealed by the X-ray structure of a core U2AF35/U2AF65 heterodimer. Cell 106, 595–605 (2001).
Selenko, P. et al. Structural basis for the molecular recognition between human splicing factors U2AF65 and SF1/mBBP. Mol. Cell 11, 965–976 (2003).
D'Souza, V. & Summers, M.F. Structural basis for packaging the dimeric genome of Moloney murine leukaemia virus. Nature 431, 586–590 (2004).
De Guzman, R.N. et al. Structure of the HIV-1 nucleocapsid protein bound to the SL3 Ψ-RNA recognition element. Science 279, 384–388 (1998).
Lee, B.M. et al. Induced fit and “lock and key” recognition of 5S RNA by zinc fingers of transcription factor IIIA. J. Mol. Biol. 357, 275–291 (2006).
Lu, D., Searles, M.A. & Klug, A. Crystal structure of a zinc-finger-RNA complex reveals two modes of molecular recognition. Nature 426, 96–100 (2003).
Warf, M.B. & Berglund, J.A. MBNL binds similar RNA structures in the CUG repeats of myotonic dystrophy and its pre-mRNA substrate cardiac troponin T. RNA 13, 2238–2251 (2007).
Oberstrass, F.C. et al. Structure of PTB bound to RNA: specific binding and implications for splicing regulation. Science 309, 2054–2057 (2005).
Ule, J. et al. An RNA map predicting Nova-dependent splicing regulation. Nature 444, 580–586 (2006).
Martinez-Contreras, R. et al. Intronic binding sites for hnRNP A/B and hnRNP F/H proteins stimulate pre-mRNA splicing. PLoS Biol. 4, e21 (2006).
Marley, J., Lu, M. & Bracken, C. A method for efficient isotopic labeling of recombinant proteins. J. Biomol. NMR 20, 71–75 (2001).
Otwinowski, Z. & Minor, W. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Schneider, T.R. & Sheldrick, G.M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002).
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Collaborative Computational Project, Number 4. Collaborative computational project number 4. The CCP4 suite: programmes for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Cambillau, C. & Roussel, A. Turbo Frodo, Version OpenGL.1 〈http://www.afmb.univ-mrs.fr/-TURBO-〉(1997).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Poveda, J.A., Prieto, M., Encinar, J.A., Gonzalez-Ros, J.M. & Mateo, C.R. Intrinsic tyrosine fluorescence as a tool to study the interaction of the shaker B “ball” peptide with anionic membranes. Biochemistry 42, 7124–7132 (2003).
Kurganov, B.I., Sugrobova, N.P. & Yakovlev, V.A. Estimation of dissociation constant of enzyme-ligand complex from fluorometric data by “difference” method. FEBS Lett. 19, 308–310 (1972).
Acknowledgements
This research was supported by the US National Institutes of Health grant CA049982 to D.J.P. We thank S. Ilin for help with NMR data collection and O. Rechkoblit for help with X-ray synchrotron data collection of one of the data sets. We would like to thank the staff of NE-CAT beam line at the Advanced Photon Source, Argonne National Laboratory, supported by the US Department of Energy, for assistance with data collection.
Author information
Authors and Affiliations
Contributions
The biochemical and structural research was undertaken by M.T. under the supervision of D.J.P. Both M.T. and D.J.P. contributed to the writing of the paper.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 3128 kb)
Rights and permissions
About this article
Cite this article
Teplova, M., Patel, D. Structural insights into RNA recognition by the alternative-splicing regulator muscleblind-like MBNL1. Nat Struct Mol Biol 15, 1343–1351 (2008). https://doi.org/10.1038/nsmb.1519
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1519
This article is cited by
-
UTP11 promotes the growth of hepatocellular carcinoma by enhancing the mRNA stability of Oct4
BMC Cancer (2024)
-
Global tissue transcriptomic analysis to improve genome annotation and unravel skin pigmentation in goldfish
Scientific Reports (2021)
-
Elucidation of the aberrant 3′ splice site selection by cancer-associated mutations on the U2AF1
Nature Communications (2020)
-
Hybrid splicing minigene and antisense oligonucleotides as efficient tools to determine functional protein/RNA interactions
Scientific Reports (2017)
-
Recognition of distinct RNA motifs by the clustered CCCH zinc fingers of neuronal protein Unkempt
Nature Structural & Molecular Biology (2016)