Abstract
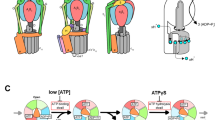
Rotation of the central shaft γ subunit in a molecular motor F1-ATPase is assumed to correlate with and probably be driven by domain motions of the three catalytic β subunits. Here we observe directly these β motions through an attached fluorophore, concomitantly with 80° and 40° substep rotations of γ in the same single molecules. We show the sequence of conformations that each β subunit undergoes in three-step bending, a ∼40° counterclockwise turn followed by two ∼20° clockwise turns, occurring in synchronization with two substep rotations of γ. The results indicate that most previous crystal structures mimic the conformational set of three β subunits in the catalytic dwells. Moreover, a previously undescribed set of β conformations, open, closed and partially closed, is revealed in the ATP-waiting dwells. The present study thus bridges the gap between the chemical and mechanical steps in F1-ATPase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Noji, H., Yasuda, R., Yoshida, M. & Kinosita, K. Jr. Direct observation of the rotation of F1-ATPase. Nature 386, 299–302 (1997).
Yasuda, R., Noji, H., Kinosita, K. Jr & Yoshida, M. F1-ATPase is a highly efficient molecular motor that rotates with discrete 120° steps. Cell 93, 1117–1124 (1998).
Yasuda, R., Noji, H., Yoshida, M., Kinosita, K. Jr & Itoh, H. Resolution of distinct rotational substeps by submillisecond kinetic analysis of F1-ATPase. Nature 410, 898–904 (2001).
Shimabukuro, K. et al. Catalysis and rotation of F1 motor: cleavage of ATP at the catalytic site occurs in 1 ms before 40° substep rotation. Proc. Natl. Acad. Sci. USA 100, 14731–14736 (2003).
Adachi, K. et al. Coupling of rotation and catalysis in F1-ATPase revealed by single-molecule imaging and manipulation. Cell 130, 309–321 (2007).
Watanabe, R., Iino, R., Shimabukuro, K., Yoshida, M. & Noji, H. Temperature-sensitive reaction intermediate of F1-ATPase. EMBO Rep. 9, 84–90 (2008).
Furuike, S. et al. Temperature dependence of the rotation and hydrolysis activities of F1-ATPase. Biophys. J. 95, 761–770 (2008).
Ariga, T., Muneyuki, E. & Yoshida, M. F1-ATPase rotates by an asymmetric, sequential mechanism using all three catalytic subunits. Nat. Struct. Mol. Biol. 14, 841–846 (2007).
Nishizaka, T. et al. Chemomechanical coupling in F1-ATPase revealed by simultaneous observation of nucleotide kinetics and rotation. Nat. Struct. Mol. Biol. 11, 142–148 (2004).
Weber, J. & Senior, A.E. ATP synthase: what we know about ATP hydrolysis and what we do not know about ATP synthesis. Biochim. Biophys. Acta 1458, 300–309 (2000).
Ono, S. et al. Origin of apparent negative cooperativity of F1-ATPase. Biochim. Biophys. Acta 1607, 35–44 (2003).
Ren, H., Bandyopadhyay, S. & Allison, W.S. The α3(βMet222Ser/Tyr345Trp)3γ subcomplex of the TF1-ATPase does not hydolyze ATP at a significant rate until the substrate binds to the catalytic site of the lowest affinity. Biochemistry 45, 6222–6230 (2006).
Kayalar, C., Rosing, J. & Boyer, P.D. An alternating site sequence for oxidative phosphorylation suggested by measurement of substrate binding patterns and exchange reaction inhibitions. J. Biol. Chem. 252, 2486–2491 (1977).
Abrahams, J.P., Leslie, A.G., Lutter, R. & Walker, J.E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994).
Ren, H., Dou, C., Stelzer, M.S. & Allison, W.S. Oxidation of the α3(βD311C/R333C)3γ subcomplex of the thermophilic Bacillus PS3 F1-ATPase indicates that only two β subunits can exist in the closed conformation simultaneously. J. Biol. Chem. 274, 31366–31372 (1999).
Tsunoda, S.P., Muneyuki, E., Amano, T., Yoshida, M. & Noji, H. Cross-linking of two β subunits in the closed conformation in F1-ATPase. J. Biol. Chem. 274, 5701–5706 (1999).
Tozawa, K. et al. Functions and ATP-binding responses of the twelve histidine residues in the TF1-ATPase β subunit. J. Biochem. 130, 527–533 (2001).
Yagi, H., Tsujimoto, T., Yamazaki, T., Yoshida, M. & Akutsu, H. Conformational change of H+-ATPase β monomer revealed on segmental isotope labeling NMR spectroscopy. J. Am. Chem. Soc. 126, 16632–16638 (2004).
Masaike, T., Muneyuki, E., Noji, H., Kinosita, K. Jr & Yoshida, M. F1-ATPase changes its conformations upon phosphate release. J. Biol. Chem. 277, 21643–21649 (2002).
Masaike, T., Suzuki, T., Tsunoda, S.P., Konno, H. & Yoshida, M. Probing conformations of the β subunit of F0F1-ATP synthase in catalysis. Biochem. Biophys. Res. Commun. 342, 800–807 (2006).
Wang, H. & Oster, G. Energy transduction in the F1 motor of ATP synthase. Nature 396, 279–282 (1998).
Sun, S.X., Wang, H. & Oster, G. Asymmetry in the F1-ATPase and its implications for the rotational cycle. Biophys. J. 86, 1373–1384 (2004).
Forkey, J.N., Quinlan, M.E., Shaw, M.A., Corrie, J.E. & Goldman, Y.E. Three-dimensional structural dynamics of myosin V by single-molecule fluorescence polarization. Nature 422, 399–404 (2003).
Tomishige, M., Stuurman, N. & Vale, R.D. Single-molecule observations of neck linker conformational changes in the kinesin motor protein. Nat. Struct. Mol. Biol. 13, 887–894 (2006).
Yildiz, A. et al. Myosin V walks hand-over-hand: single fluorophore imaging with 1.5-nm localization. Science 300, 2061–2065 (2003).
Mori, T., Vale, R.D. & Tomishige, M. How kinesin waits between steps. Nature 450, 750–754 (2007).
Iino, R., Rondelez, Y., Yoshida, M. & Noji, H. Chemomechanical coupling in single-molecule F-type ATP synthase. J. Bioenerg. Biomembr. 37, 451–454 (2005).
Zimmermann, B., Diez, M., Zarrabi, N., Graber, P. & Borsch, M. Movements of the ε-subunit during catalysis and activation in single membrane-bound H+-ATP synthase. EMBO J. 24, 2053–2063 (2005).
Kagawa, Y. & Hamamoto, T. The energy transmission in ATP synthase: from the γ-c rotor to the α3β3 oligomer fixed by OSCP-b stator via the β DELSEED sequence. J. Bioenerg. Biomembr. 28, 421–431 (1996).
Hara, K.Y. et al. The role of the DELSEED motif of the β subunit in rotation of F1-ATPase. J. Biol. Chem. 275, 14260–14263 (2000).
Corrie, J.E. et al. Dynamic measurement of myosin light-chain-domain tilt and twist in muscle contraction. Nature 400, 425–430 (1999).
Ha, T., Enderle, T., Chemla, D.S., Selvin, P.R. & Weiss, S. Single molecule dynamics studied by polarization modulation. Phys. Rev. Lett. 77, 3979–3982 (1996).
Amano, T., Tozawa, K., Yoshida, M. & Murakami, H. Spatial precision of a catalytic carboxylate of F1-ATPase β subunit probed by introducing different carboxylate-containing side chains. FEBS Lett. 348, 93–98 (1994).
Ariga, T., Masaike, T., Noji, H. & Yoshida, M. Stepping rotation of F1-ATPase with one, two, or three altered catalytic sites that bind ATP only slowly. J. Biol. Chem. 277, 24870–24874 (2002).
Gibbons, C., Montgomery, M.G., Leslie, A.G. & Walker, J.E. The structure of the central stalk in bovine F1-ATPase at 2.4 Å resolution. Nat. Struct. Biol. 7, 1055–1061 (2000).
Abrahams, J.P. et al. The structure of bovine F1-ATPase complexed with the peptide antibiotic efrapeptin. Proc. Natl. Acad. Sci. USA 93, 9420–9424 (1996).
van Raaij, M.J., Abrahams, J.P., Leslie, A.G. & Walker, J.E. The structure of bovine F1-ATPase complexed with the antibiotic inhibitor aurovertin B. Proc. Natl. Acad. Sci. USA 93, 6913–6917 (1996).
Orriss, G.L., Leslie, A.G., Braig, K. & Walker, J.E. Bovine F1-ATPase covalently inhibited with 4-chloro-7-nitrobenzofurazan: the structure provides further support for a rotary catalytic mechanism. Structure 6, 831–837 (1998).
Braig, K., Menz, R.I., Montgomery, M.G., Leslie, A.G. & Walker, J.E. Structure of bovine mitochondrial F1-ATPase inhibited by Mg2+ ADP and aluminium fluoride. Structure 8, 567–573 (2000).
Kagawa, R., Montgomery, M.G., Braig, K., Leslie, A.G. & Walker, J.E. The structure of bovine F1-ATPase inhibited by ADP and beryllium fluoride. EMBO J. 23, 2734–2744 (2004).
Bowler, M.W., Montgomery, M.G., Leslie, A.G. & Walker, J.E. How azide inhibits ATP hydrolysis by the F-ATPases. Proc. Natl. Acad. Sci. USA 103, 8646–8649 (2006).
Bowler, M.W., Montgomery, M.G., Leslie, A.G. & Walker, J.E. Ground state structure of F1-ATPase from bovine heart mitochondria at 1.9 Å resolution. J. Biol. Chem. 282, 14238–14242 (2007).
Weber, J. & Senior, A.E. Catalytic mechanism of F1-ATPase. Biochim. Biophys. Acta 1319, 19–58 (1997).
Nakamoto, R.K., Baylis Scanlon, J.A. & Al-Shawi, M.K. The rotary mechanism of the ATP synthase. Arch. Biochem. Biophys. 476, 43–50 (2008).
Yasuda, R. et al. The ATP-waiting conformation of rotating F1-ATPase revealed by single-pair fluorescence resonance energy transfer. Proc. Natl. Acad. Sci. USA 100, 9314–9318 (2003).
Menz, R.I., Walker, J.E. & Leslie, A.G. Structure of bovine mitochondrial F1-ATPase with nucleotide bound to all three catalytic sites: implications for the mechanism of rotary catalysis. Cell 106, 331–341 (2001).
Furuike, S. et al. Axle-less F1-ATPase rotates in the correct direction. Science 319, 955–958 (2008).
Murakami, S., Nakashima, R., Yamashita, E., Matsumoto, T. & Yamaguchi, A. Crystal structures of a multidrug transporter reveal a functionally rotating mechanism. Nature 443, 173–179 (2006).
Suno, R. et al. Structure of the whole cytosolic region of ATP-dependent protease FtsH. Mol. Cell 22, 575–585 (2006).
Hirono-Hara, Y. et al. Pause and rotation of F1-ATPase during catalysis. Proc. Natl. Acad. Sci. USA 98, 13649–13654 (2001).
Kabaleeswaran, V., Puri, N., Walker, J.E., Leslie, A.G. & Mueller, D.M. Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1-ATPase. EMBO J. 25, 5433–5442 (2006).
Acknowledgements
We are grateful to S. Kimura for technical assistance, T. Amano for advice on preparation and biochemical studies of the β subunit, H. Noji, T. Ariga, R. Iino, H. Ueno, K. Adachi and M. Takeda for discussion on the rotation assay, J. Corrie for discussion on fluorescent probes, Y. Matsumoto and T. Shibahara for advice on MS, S. Hayashi and M. Ikeguchi for discussion on crystal structures, A.P. Dominey for critically reading the manuscript, J. Yajima for discussion, K. Abe for the objective lens and our colleagues for discussion, technical advice and assistance. This study was supported in part by the ERATO Yoshida ATP System Project of Japan Science and Technology Agency, a Grant-in-Aid for Scientific Research on Priority Areas (No. 18074008 to T.N. and T.M.), a Grant-in-Aid for Young Scientists (B) (No.18770138 to T.M.) from the Ministry of Education, Culture, Sports, Science and Technology of Japan and from the Japan Society for the Promotion of Science, Saneyoshi Scholarship (No.1817 to T.M.), the Grant Program for Natural Science from the Asahi Glass Foundation to T.M. and a grant from the New Energy and Industrial Technology Development Organization (NEDO) to T.N.
Author information
Authors and Affiliations
Contributions
T.M., F.K.-H. and T.N. performed the experiments; K.O., M.Y. and T.N. supervised the project; T.M. wrote the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 1250 kb)
Supplementary Video 1
The video shows motions of the C-terminal domain in a single catalytic subunit β upon rotational substeps of the γ-shaft in F1-ATPase shown in Fig. 2b-f. A fluorophore linked to β (right column) and polystyrene beads attached to γ (left column) are simultaneously observed. In different angles of γ and chemical states of β (0°, 120°, 200° and 240° from top to bottom rows), the fluorophore shows phase shifts in spite of synchronized polarization of the excitation laser. It indicates domain motions of β represented by angular motions of the C-terminal helix upon rotational substeps of γ. Colored boxes show peak frames obtained by sinusoidal fitting of fluorescence intensity during each dwell. Images were recorded at 30 frames per second, and are played at 5 frames per second (1/6 speed) for 45 frames (3 cycles of polarization of the excitation laser). The images of beads were rotated with bilinear interpolation so that the beads point to the 12 o'clock direction at the 0° dwell. All contrast manipulations and enhancements for clarity were strictly linear in each column. Frame sizes are 0.57 μm × 0.52 μm (height × width) for the beads window (left frames, inner size) and 0.97 μm × 0.97 μm (height × width) for the fluorescence window (right frames, inner size). To see the phase shift of the fluorescence intensity clearly, it is recommended to loop the movie and play continuously. (MOV 769 kb)
Rights and permissions
About this article
Cite this article
Masaike, T., Koyama-Horibe, F., Oiwa, K. et al. Cooperative three-step motions in catalytic subunits of F1-ATPase correlate with 80° and 40° substep rotations. Nat Struct Mol Biol 15, 1326–1333 (2008). https://doi.org/10.1038/nsmb.1510
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1510
This article is cited by
-
Angle change of the A-domain in a single SERCA1a molecule detected by defocused orientation imaging
Scientific Reports (2021)
-
Direct visualization of virus removal process in hollow fiber membrane using an optical microscope
Scientific Reports (2021)
-
The catalytic dwell in ATPases is not crucial for movement against applied torque
Nature Chemistry (2020)
-
Single-molecule pull-out manipulation of the shaft of the rotary motor F1-ATPase
Scientific Reports (2019)
-
Motor torque measurement of Halobacterium salinarum archaellar suggests a general model for ATP-driven rotary motors
Communications Biology (2019)