Abstract
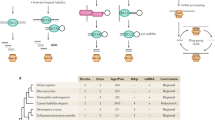
In the fission yeast Schizosaccharomyces pombe, the RNA interference (RNAi) machinery is required to generate small interfering RNAs (siRNAs) that mediate heterochromatic gene silencing. Efficient silencing also requires the TRAMP complex, which contains the noncanonical Cid14 poly(A) polymerase and targets aberrant RNAs for degradation. Here we use high-throughput sequencing to analyze Argonaute-associated small RNAs (sRNAs) in both the presence and absence of Cid14. Most sRNAs in fission yeast start with a 5′ uracil, and we argue these are loaded most efficiently into Argonaute. In wild-type cells most sRNAs match to repeated regions of the genome, whereas in cid14Δ cells the sRNA profile changes to include major new classes of sRNAs originating from ribosomal RNAs and a tRNA. Thus, Cid14 prevents certain abundant RNAs from becoming substrates for the RNAi machinery, thereby freeing the RNAi machinery to act on its proper targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Hannon, G.J. RNA interference. Nature 418, 244–251 (2002).
Hamilton, A.J. & Baulcombe, D.C. A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286, 950–952 (1999).
Hammond, S.M., Bernstein, E., Beach, D. & Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–296 (2000).
Zamore, P.D., Tuschl, T., Sharp, P.A. & Bartel, D.P. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25–33 (2000).
Elbashir, S.M., Lendeckel, W. & Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Baulcombe, D. RNA silencing in plants. Nature 431, 356–363 (2004).
Sijen, T. et al. On the role of RNA amplification in dsRNA-triggered gene silencing. Cell 107, 465–476 (2001).
Zaratiegui, M., Irvine, D.V. & Martienssen, R.A. Noncoding RNAs and gene silencing. Cell 128, 763–776 (2007).
Buhler, M. & Moazed, D. Transcription and RNAi in heterochromatic gene silencing. Nat. Struct. Mol. Biol. 14, 1041–1048 (2007).
Volpe, T.A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Verdel, A. et al. RNAi-mediated targeting of heterochromatin by the RITS complex. Science 303, 672–676 (2004).
Buker, S.M. et al. Two different Argonaute complexes are required for siRNA generation and heterochromatin assembly in fission yeast. Nat. Struct. Mol. Biol. 14, 200–207 (2007).
Reinhart, B.J. & Bartel, D.P. Small RNAs correspond to centromere heterochromatic repeats. Science 297, 1831 (2002).
Cam, H.P. et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat. Genet. 37, 809–819 (2005).
Motamedi, M.R. et al. Two RNAi complexes, RITS and RDRC, physically interact and localize to noncoding centromeric RNAs. Cell 119, 789–802 (2004).
Buhler, M., Verdel, A. & Moazed, D. Tethering RITS to a nascent transcript initiates RNAi- and heterochromatin-dependent gene silencing. Cell 125, 873–886 (2006).
Sadaie, M., Iida, T., Urano, T. & Nakayama, J. A chromodomain protein, Chp1, is required for the establishment of heterochromatin in fission yeast. EMBO J. 23, 3825–3835 (2004).
Buhler, M., Haas, W., Gygi, S.P. & Moazed, D. RNAi-dependent and -independent RNA turnover mechanisms contribute to heterochromatic gene silencing. Cell 129, 707–721 (2007).
Chen, E.S. et al. Cell cycle control of centromeric repeat transcription and heterochromatin assembly. Nature 451, 734–737 (2008).
LaCava, J. et al. RNA degradation by the exosome is promoted by a nuclear polyadenylation complex. Cell 121, 713–724 (2005).
Stevenson, A.L. & Norbury, C.J. The Cid1 family of non-canonical poly(A) polymerases. Yeast 23, 991–1000 (2006).
Win, T.Z. et al. Requirement of fission yeast Cid14 in polyadenylation of rRNAs. Mol. Cell. Biol. 26, 1710–1721 (2006).
Vanacova, S. et al. A new yeast poly(A) polymerase complex involved in RNA quality control. PLoS Biol. 3, e189 (2005).
Wyers, F. et al. Cryptic Pol II transcripts are degraded by a nuclear quality control pathway involving a new poly(A) polymerase. Cell 121, 725–737 (2005).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Rajagopalan, R., Vaucheret, H., Trejo, J. & Bartel, D.P. A diverse and evolutionarily fluid set of microRNAs in Arabidopsis thaliana. Genes Dev. 20, 3407–3425 (2006).
Ambros, V., Lee, R.C., Lavanway, A., Williams, P.T. & Jewell, D. MicroRNAs and other tiny endogenous RNAs in C. elegans. Curr. Biol. 13, 807–818 (2003).
Ruby, J.G. et al. Large-scale sequencing reveals 21U-RNAs and additional microRNAs and endogenous siRNAs in C. elegans. Cell 127, 1193–1207 (2006).
Lau, N.C., Lim, L.P., Weinstein, E.G. & Bartel, D.P. An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862 (2001).
Reinhart, B.J., Weinstein, E.G., Rhoades, M.W., Bartel, B. & Bartel, D.P. MicroRNAs in plants. Genes Dev. 16, 1616–1626 (2002).
Aravin, A.A. et al. The small RNA profile during Drosophila melanogaster development. Dev. Cell 5, 337–350 (2003).
Aravin, A. et al. A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442, 203–207 (2006).
Lau, N.C. et al. Characterization of the piRNA complex from rat testes. Science 313, 363–367 (2006).
Girard, A., Sachidanandam, R., Hannon, G.J. & Carmell, M.A. A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442, 199–202 (2006).
Grivna, S.T., Beyret, E., Wang, Z. & Lin, H. A novel class of small RNAs in mouse spermatogenic cells. Genes Dev. 20, 1709–1714 (2006).
Watanabe, T. et al. Identification and characterization of two novel classes of small RNAs in the mouse germline: retrotransposon-derived siRNAs in oocytes and germline small RNAs in testes. Genes Dev. 20, 1732–1743 (2006).
Rand, T.A., Petersen, S., Du, F. & Wang, X. Argonaute2 cleaves the anti-guide strand of siRNA during RISC activation. Cell 123, 621–629 (2005).
Matranga, C., Tomari, Y., Shin, C., Bartel, D.P. & Zamore, P.D. Passenger-strand cleavage facilitates assembly of siRNA into Ago2-containing RNAi enzyme complexes. Cell 123, 607–620 (2005).
Irvine, D.V. et al. Argonaute slicing is required for heterochromatic silencing and spreading. Science 313, 1134–1137 (2006).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Allshire, R.C., Javerzat, J.P., Redhead, N.J. & Cranston, G. Position effect variegation at fission yeast centromeres. Cell 76, 157–169 (1994).
Scott, K.C., Merrett, S.L. & Willard, H.F. A heterochromatin barrier partitions the fission yeast centromere into discrete chromatin domains. Curr. Biol. 16, 119–129 (2006).
Good, L., Intine, R.V. & Nazar, R.N. The ribosomal-RNA-processing pathway in Schizosaccharomyces pombe. Eur. J. Biochem. 247, 314–321 (1997).
Wang, S.W., Stevenson, A.L., Kearsey, S.E., Watt, S. & Bahler, J. Global role for polyadenylation-assisted nuclear RNA degradation in posttranscriptional gene silencing. Mol. Cell. Biol. 28, 656–665 (2008).
Hong, E.J., Villen, J., Gerace, E.L., Gygi, S.P. & Moazed, D. A cullin E3 ubiquitin ligase complex associates with Rik1 and the Clr4 histone H3–K9 methyltransferase and is required for RNAi-mediated heterochromatin formation. RNA Biol. 2, 106–111 (2005).
Li, F. et al. Two novel proteins, Dos1 and Dos2, interact with Rik1 to regulate heterochromatic RNA interference and histone modification. Curr. Biol. 15, 1448–1457 (2005).
Verdel, A. & Moazed, D. RNAi-directed assembly of heterochromatin in fission yeast. FEBS Lett. 579, 5872–5878 (2005).
Justesen, J., Hartmann, R. & Kjeldgaard, N.O. Gene structure and function of the 2′-5′-oligoadenylate synthetase family. Cell. Mol. Life Sci. 57, 1593–1612 (2000).
Chen, C.C. et al. A member of the polymerase β nucleotidyltransferase superfamily is required for RNA interference in C. elegans. Curr. Biol. 15, 378–383 (2005).
Lee, S.R. & Collins, K. Physical and functional coupling of RNA-dependent RNA polymerase and Dicer in the biogenesis of endogenous siRNAs. Nat. Struct. Mol. Biol. 14, 604–610 (2007).
Gullerova, M. & Proudfoot, N.J. Cohesin complex promotes transcriptional termination between convergent genes in S. pombe. Cell 132, 983–995 (2008).
Hansen, K.R. et al. Global effects on gene expression in fission yeast by silencing and RNA interference machineries. Mol. Cell. Biol. 25, 590–601 (2005).
Ho, C.K., Wang, L.K., Lima, C.D. & Shuman, S. Structure and mechanism of RNA ligase. Structure 12, 327–339 (2004).
Acknowledgements
We thank W. Johnston and S. Buker for reagents and members of the Moazed and Bartel laboratories for helpful discussions. We also thank H. Grosshans for comments on the manuscript. M.B. was supported by a Swiss National Science Foundation postdoctoral fellowship and is currently supported by the Novartis Research Foundation and the Swiss National Science Foundation SNF Professorship. This work was supported by grants from the US National Institutes of Health (D.M.) and the Howard Hughes Medical Institute (D.P.B.).
Author information
Authors and Affiliations
Contributions
M.B. performed the experimental work with yeast; N.S. performed the computational analysis of sequencing reads; All authors contributed to the design of the study and preparation of the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1, Supplementary Tables 1–5 and Supplmentary Results (PDF 699 kb)
Rights and permissions
About this article
Cite this article
Bühler, M., Spies, N., Bartel, D. et al. TRAMP-mediated RNA surveillance prevents spurious entry of RNAs into the Schizosaccharomyces pombe siRNA pathway. Nat Struct Mol Biol 15, 1015–1023 (2008). https://doi.org/10.1038/nsmb.1481
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1481
This article is cited by
-
Roles and regulation of tRNA-derived small RNAs in animals
Nature Reviews Molecular Cell Biology (2024)
-
The inner nuclear membrane protein Lem2 coordinates RNA degradation at the nuclear periphery
Nature Structural & Molecular Biology (2022)
-
FIERY1 promotes microRNA accumulation by suppressing rRNA-derived small interfering RNAs in Arabidopsis
Nature Communications (2019)
-
MINTmap: fast and exhaustive profiling of nuclear and mitochondrial tRNA fragments from short RNA-seq data
Scientific Reports (2017)
-
Tailing and degradation of Argonaute-bound small RNAs protect the genome from uncontrolled RNAi
Nature Communications (2017)