Abstract
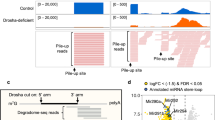
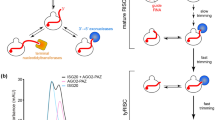
microRNAs (miRNAs) are generated from long primary (pri-) RNA polymerase II (Pol II)–derived transcripts by two RNase III processing reactions: Drosha cleavage of nuclear pri-miRNAs and Dicer cleavage of cytoplasmic pre-miRNAs. Here we show that Drosha cleavage occurs during transcription acting on both independently transcribed and intron-encoded miRNAs. We also show that both 5′-3′ and 3′-5′ exonucleases associate with the sites where co-transcriptional Drosha cleavage occurs, promoting intron degradation before splicing. We finally demonstrate that miRNAs can also derive from 3` flanking transcripts of Pol II genes. Our results demonstrate that multiple miRNA-containing transcripts are co-transcriptionally cleaved during their synthesis and suggest that exonucleolytic degradation from Drosha cleavage sites in pre-mRNAs may influence the splicing and maturation of numerous mRNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kim, V.N. MicroRNA biogenesis: coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 6, 376–385 (2005).
Basyuk, E., Suavet, F., Doglio, A., Bordonné, R. & Bertrand, E. Human let-7 stem-loop precursors harbour features of RNase III cleavage products. Nucleic Acids Res. 31, 6593–6597 (2003).
Denli, A.M., Tops, B.B.J., Plasterk, R.H.A., Ketting, R.F. & Hannon, G.J. Processing of primary microRNAs by the microprocessor complex. Nature 432, 231–235 (2004).
Gregory, R.I. et al. The Microprocessor complex mediates the genesis of microRNAs. Nature 432, 235–240 (2004).
Han, J. et al. Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell 125, 887–901 (2006).
Lund, E., Güttinger, S., Calado, A., Dahlberg, J.E. & Kutay, U. Nuclear export of microRNA precursors. Science 303, 95–98 (2004).
Valencia-Sanchez, M.A., Liu, J., Hannon, G.J. & Parker, R. Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev. 20, 515–524 (2006).
Hannon, G.J. RNA interference. Nature 418, 244–251 (2002).
Schwarz, D.S. et al. Asymmetry in the assembly of the RNAi enzyme complex. Cell 115, 199–208 (2003).
Lee, Y., Jeon, K., Lee, J.T., Kim, S. & Kim, V.N. MicroRNA maturation: stepwise processing and subcellular localization. EMBO J. 21, 4663–4670 (2002).
Saini, H.K., Griffiths-Jones, S. & Enright, A.J. Genomic analysis of human microRNA transcripts. Proc. Natl. Acad. Sci. USA 104, 17719–17724 (2007).
Lagos-Quintana, M., Rauhut, R., Meyer, J., Borkhardt, A. & Tuschl, T. New microRNAs from mouse and human. RNA 9, 175–179 (2003).
Rodriguez, A., Griffiths-Jones, S., Ashurst, J.L. & Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 14, 1902–1910 (2004).
Kim, Y.K. & Kim, V.N. Processing of intronic microRNAs. EMBO J. 26, 775–783 (2007).
Baskerville, S. & Bartel, D.P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247 (2005).
Smalheiser, N.R. EST analyses predict the existence of a population of chimeric microRNA precursor-mRNA transcripts expressed in normal human and mouse tissues. Genome Biol. 4, 403 (2003).
Qu, L.H. et al. U24, a novel intron-encoded small nucleolar RNA with two 12 nt long, phylogenetically conserved complementarities to 28S rRNA. Nucleic Acids Res. 23, 2669–2676 (1995).
de Turris, V. et al. TOP promoter elements control the relative ratio of intron-encoded snoRNA versus spliced mRNA biosynthesis. J. Mol. Biol. 344, 383–394 (2004).
Hirose, T., Shu, M.D. & Steitz, J.A. Splicing-dependent and -independent modes of assembly for intron-encoded box C/D snoRNPs in mammalian cells. Mol. Cell 12, 113–123 (2003).
Richard, P., Kiss, A.M., Darzacq, X. & Kiss, T. Cotranscriptional recognition of human intronic box H/ACA snoRNAs occurs in a splicing-independent manner. Mol. Cell. Biol. 26, 2540–2549 (2006).
Yang, P.K. et al. Cotranscriptional recruitment of the pseudouridylsynthetase Cbf5p and of the RNA binding protein Naf1p during H/ACA snoRNP assembly. Mol. Cell. Biol. 25, 3295–3304 (2005).
Fazi, F. et al. A minicircuitry comprised of microRNA-223 and transcription factors NFI-A and C/EBPα regulates human granulopoiesis. Cell 123, 819–831 (2005).
Lagos-Quintana, M., Rauhut, R., Lendeckel, W. & Tuschl, T. Identification of novel genes coding for small expressed RNAs. Science 294, 853–858 (2001).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
Glover-Cutter, K., Kim, S., Espinosa, J. & Bentley, D.L. RNA polymerase II pauses and associates with pre-mRNA processing factors at both ends of genes. Nat. Struct. Mol. Biol. 15, 71–78 (2008).
Yeom, K.H., Lee, Y., Han, J., Suh, M.R. & Kim, V.N. Characterization of DGCR8/Pasha, the essential cofactor for Drosha in primary miRNA processing. Nucleic Acids Res. 34, 4622–4629 (2006).
Dye, M.J., Gromak, N. & Proudfoot, N.J. Exon tethering in transcription by RNA polymerase II. Mol. Cell 21, 849–859 (2006).
Gromak, N., Talotti, G., Proudfoot, N.J. & Pagani, F. Modulating alternative splicing by cotranscriptional cleavage of nascent intronic RNA. RNA 14, 359–366 (2007).
Kluiver, J. et al. Regulation of pri-microRNA BI transcription and processing in Burkitt lynphoma. Oncogene 26, 3769–3776 (2007).
Thomson, J.M. et al. Extensive post-transcriptional regulation of microRNA and its implication for cancer. Genes Dev. 20, 2202–2207 (2006).
West, S., Gromak, N. & Proudfoot, N.J. Human 5′ → 3′ exonuclease Xrn2 promotes transcription termination at co-transcriptional cleavage sites. Nature 432, 522–525 (2004).
West, S., Proudfoot, N.J. & Dye, M. Molecular dissection of mammalian RNA polymerase II transcriptional termination. Mol. Cell 29, 600–610 (2008).
West, S., Gromak, N., Norbury, C.J. & Proudfoot, N.J. Adenylation and exosome-mediated degradation of cotranscriptionally cleaved pre-messenger RNA in human cells. Mol. Cell 21, 437–443 (2006).
Wuarin, J. & Schibler, U. Physical isolation of nascent RNA chains transcribed by RNA polymerase II: evidence for cotranscriptional splicing. Mol. Cell. Biol. 14, 7219–7225 (1994).
Zhou, H. & Lin, K. Excess of microRNAs in large and very 5′ biased introns. Biochem. Biophys. Res. Commun. 368, 709–715 (2008).
Kampa, D. et al. Novel RNAs identified from an in-depth analysis of the transcriptome of human chromosomes 21 and 22. Genome Res. 14, 331–342 (2004).
Clark, T.A. et al. Discovery of tissue-specific exons using comprehensive human exon microarrays. Genome Biol. 8, R64 (2007).
Proudfoot, N.J., Furger, A. & Dye, M.J. Integrating mRNA processing with transcription. Cell 108, 501–512 (2002).
Ryman, K., Fong, N., Bratt, E., Bentley, D.L. & Ohman, M. The C-terminal domain of RNA Pol II helps ensure that editing precedes splicing of the GluR-B transcript. RNA 13, 1071–1078 (2007).
Kim, M. et al. The yeast Rat1 exonuclease promotes transcription termination by RNA polymerase II. Nature 432, 517–522 (2004).
Gromak, N., West, S. & Proudfoot, N.J. Pause sites promote transcriptional termination of mammalian RNA polymerase II. Mol. Cell. Biol. 26, 3986–3996 (2006).
Dye, M.J., Gromak, N., Haussecker, D., West, S. & Proudfoot, N.J. Turnover and function of noncoding RNA polymerase II transcripts. Cold Spring Harb. Symp. Quant. Biol. 71, 275–284 (2006).
Danin-Kreiselman, M., Lee, C.Y. & Chanfreau, G. RNAse III-mediated degradation of unspliced pre-mRNAs and lariat introns. Mol. Cell 11, 1279–1289 (2003).
Kiss, T. SnoRNP biogenesis meets Pre-mRNA splicing. Mol. Cell 23, 775–776 (2006).
Shiohama, A., Sasaki, T., Noda, S., Minoshima, S. & Shimizu, N. Nucleolar localization of DGCR8 and identification of eleven DGCR8-associated proteins. Exp. Cell Res. 313, 4196–4207 (2007).
Guil, S. & Cáceres, J.F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat. Struct. Mol. Biol. 14, 591–596 (2007).
Wagner, E.J. & Garcia-Blanco, M.A. RNAi-mediated PTB depletion leads to enhanced exon definition. Mol. Cell 10, 943–949 (2002).
Forsberg, E.C. et al. Developmentally dynamic histone acetylation pattern of a tissue-specific chromatin domain. Proc. Natl. Acad. Sci. USA 97, 14494–14499 (2000).
Ballarino, M., Morlando, M., Pagano, F., Fatica, A. & Bozzoni, I. The cotranscriptional assembly of snoRNPs controls the biosynthesis of H/ACA snoRNAs in Saccharomyces cerevisiae. Mol. Cell. Biol. 25, 5396–5403 (2005).
Haussecker, D. & Proudfoot, N. J. Dicer-dependent turnover of intergenic transcripts from the human β-globin gene cluster. Mol. Cell. Biol. 25, 9724–9733 (2005).
Dye, M.J. & Proudfoot, N.J. Terminal exon definition occurs cotranscriptionally and promotes termination of RNA polymerase II. Mol. Cell 3, 371–378 (1999).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Megraw, M., Sethupathy, P., Corda, B. & Hatzigeorgiou, A.G. miRGen: a database for the study of animal microRNA genomic organization and function. Nucleic Acids Res. 35, D149–D155 (2007).
Dye, M.J. & Proudfoot, N.J. Multiple transcript cleavage precedes polymerase release in termination by RNA polymerase II. Cell 105, 669–681 (2001).
Pawlicki, J.M. & Steitz, J.A. Primary microRNA transcript retention at sites of transcription leads to enhanced microRNA production. J. Cell Biol. 182, 61–76 (2008).
Acknowledgements
We are grateful to D. Cacchiarelli for the bioinformatics analysis and to M. Dye and S. West for helpful advice and discussion and for sharing unpublished data. M.M. was supported by an EMBO Long term fellowship. This work was partly supported by the 6th Framework Programme of the European Commission, SIROCCO and RIGHT Integrated Projects (LSHG-CT-2006-037900 and LSHB-CT-2004 005276) and Centro di Eccellenza BEMM to I.B. and by the ESF project “NuRNASu” to I.B. and N.J.P. The Proudfoot laboratory is supported by a Wellcome Trust Programme Grant.
Author information
Authors and Affiliations
Contributions
M.M. performed the experimental work except that M.B. carried out experiments on intergenic miRNA host genes together with F.P., and N.G. carried out experiments on exonuclease involvement. M.M., I.B., N.G. and N.J.P. wrote the paper. All authors read and agree with the paper's contents.
Note: Supplementary information is available on the Nature Structural & Molecular Biology website.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, and Supplementary Table 1 (PDF 152 kb)
Rights and permissions
About this article
Cite this article
Morlando, M., Ballarino, M., Gromak, N. et al. Primary microRNA transcripts are processed co-transcriptionally. Nat Struct Mol Biol 15, 902–909 (2008). https://doi.org/10.1038/nsmb.1475
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1475
This article is cited by
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)
-
Crosstalk Between m6A RNA Methylation and miRNA Biogenesis in Cancer: An Unholy Nexus
Molecular Biotechnology (2023)
-
The dysregulation of miRNAs in epilepsy and their regulatory role in inflammation and apoptosis
Functional & Integrative Genomics (2023)
-
R-loops at microRNA encoding loci promote co-transcriptional processing of pri-miRNAs in plants
Nature Plants (2022)
-
Arabidopsis RBV is a conserved WD40 repeat protein that promotes microRNA biogenesis and ARGONAUTE1 loading
Nature Communications (2022)