Abstract
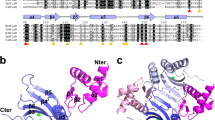
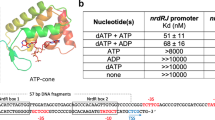
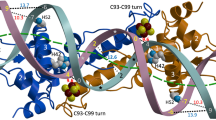
NikR is a metal-responsive transcription factor that controls nickel uptake in Escherichia coli by regulating expression of a nickel-specific ATP-binding cassette (ABC) transporter. We have determined the first two structures of NikR: the full-length apo repressor at a resolution of 2.3 Å and the nickel-bound C-terminal regulatory domain at a resolution of 1.4 Å. NikR is the only known metal-responsive member of the ribbon-helix-helix family of transcription factors, and its structure has a quaternary arrangement consisting of two dimeric DNA-binding domains separated by a tetrameric regulatory domain that binds nickel. The position of the C-terminal regulatory domain enforces a large spacing between the contacts that each NikR DNA-binding domain can make with the nik operator. The regulatory domain of NikR contains four nickel-binding sites at the tetramer interface, each exhibiting a novel square-planar coordination by three histidines and one cysteine side chain.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Maroney, M.J. Structure/function relationships in nickel metallobiochemistry. Curr. Opin. Chem. Biol. 3, 188–199 (1999).
Chivers, P.T. & Sauer, R.T. Regulation of high affinity nickel uptake in bacteria. Ni2+-dependent interaction of NikR with wild-type and mutant operator sites. J. Biol. Chem. 275, 19735–19741 (2000).
Navarro, C., Wu, L.F. & Mandrand-Berthelot, M.A. The nik operon of Escherichia coli encodes a periplasmic binding-protein-dependent transport system for nickel. Mol. Microbiol. 9, 1181–1191 (1993).
Chivers, P.T. & Sauer, R.T. NikR repressor: high-affinity nickel binding to the C-terminal domain regulates binding to operator DNA. Chem. Biol. 9, 1141–1148 (2002).
Carrington, P.E., Chivers, P.T., Al-Mjeni, F., Sauer, R.T. & Maroney, M.J. Nickel coordination is regulated by the DNA-bound state of NikR. Nat. Struct. Biol. 10, 126–130 (2003).
Chivers, P.T. & Sauer, R.T. NikR is a ribbon-helix-helix DNA-binding protein. Protein Sci. 8, 2494–2500 (1999).
Breg, J.N., van Opheusden, J.H., Burgering, M.J., Boelens, R. & Kaptein, R. Structure of Arc repressor in solution: evidence for a family of β-sheet DNA-binding proteins. Nature 346, 586–589 (1990).
Burgering, M.J. et al. Solution structure of dimeric Mnt repressor (1–76). Biochemistry 33, 15036–15045 (1994).
Gomis-Ruth, F.X. et al. The structure of plasmid-encoded transcriptional repressor CopG unliganded and bound to its operator. EMBO J. 17, 7404–7415 (1998).
Rafferty, J.B., Somers, W.S., Saint-Girons, I. & Phillips, S.E. Three-dimensional crystal structures of Escherichia coli met repressor with and without corepressor. Nature 341, 705–710 (1989).
Murayama, K., Orth, P., de la Hoz, A.B., Alonso, J.C. & Saenger, W. Crystal structure of omega transcriptional repressor encoded by Streptococcus pyogenes plasmid pSM19035 at 1.5 Å resolution. J. Mol. Biol. 314, 789–796 (2001).
Doukov, T.I., Iverson, T.M., Seravalli, J., Ragsdale, S.W. & Drennan, C.L. A Ni-Fe-Cu center in a bifunctional carbon monoxide dehydrogenase/acetyl-CoA synthase. Science 298, 567–572 (2002).
Chipman, D.M. & Shaanan, B. The ACT domain family. Curr. Opin. Struct. Biol. 11, 694–700 (2001).
Schuller, D.J., Grant, G.A. & Banaszak, L.J. The allosteric ligand site in the Vmax-type cooperative enzyme phosphoglycerate dehydrogenase. Nat. Struct. Biol. 2, 69–76 (1995).
Cho, Y., Sharma, V. & Sacchettini, J.C. Crystal structure of ATP phosphoribosyltransferase from Mycobacterium tuberculosis. J. Biol. Chem. 278, 8333–8339 (2003).
Kobe, B. et al. Structural basis of autoregulation of phenylalanine hydroxylase. Nat. Struct. Biol. 6, 442–448 (1999).
Leonard, P.M. et al. Crystal structure of the Lrp-like transcriptional regulator from the archaeon Pyrococcus furiosus. EMBO J. 20, 990–997 (2001).
Ettema, T.J., Brinkman, A.B., Tani, T.H., Rafferty, J.B. & Van Der Oost, J. A novel ligand-binding domain involved in regulation of amino acid metabolism in prokaryotes. J. Biol. Chem. 277, 37464–37468 (2002).
Schneider, T.R. A genetic algorithm for the identification of conformationally invariant regions in protein molecules. Acta Crystallogr. D 58, 195–208 (2002).
Sheldrick, G.M. & Schneider, T.R. SHELXL: high-resolution refinement. Methods Enzymol. 277, 319–343 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
LeMaster, D.M. & Richards, F.M. 1H-15N heteronuclear NMR studies of Escherichia coli thioredoxin in samples isotopically labeled by residue type. Biochemistry 24, 7263–7268 (1985).
Hunt, J.B., Neece, S.H. & Ginsburg, A. The use of 4-(2-pyridylazo)resorcinol in studies of zinc release from Escherichia coli aspartate transcarbamoylase. Anal. Biochem. 146, 150–157 (1985).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Cowtan, K.D. & Main, P. Phase combination and cross validation in iterated density-modification calculations. Acta Crystallogr. D 52, 43–48 (1996).
McRee, D.E. XtalView/Xfit—a versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Lamzin, V.S. & Wilson, K.S. Automated refinement of protein models. Acta Crystallogr. D 49, 129–147 (1993).
Terwilliger, T.C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 1174–1178 (1999).
Carson, M. Ribbons. Methods Enzymol. 277, 493–505 (1997).
Raumann, B.E., Rould, M.A., Pabo, C.O. & Sauer, R.T. DNA recognition by β-sheets in the Arc repressor-operator crystal structure. Nature 367, 754–757 (1994).
Acknowledgements
This research is supported in part by the US National Institutes of Health (NIH) (R.T.S., C.L.D.), Searle Scholars Program (C.L.D.), Cecil and Ida Green Career Development Fund (C.L.D.), Lester Wolfe Predoctoral Fellowship (E.R.S.), and the Gray Fund for Undergraduate Research (Y.G.). Data were collected at the National Synchrotron Light Source (NSLS), Advanced Light Source (ALS), Advanced Photon Source (APS), and Stanford Synchrotron Radiation Laboratory (SSRL) synchrotrons. Synchrotron facilities are funded by the US Department of Energy (ALS 5.0.2, NSLS X25, SSRL), NIH National Center of Research Resources (APS NE-CAT 8BM, NSLS X25), and the US National Institute of General Medical Sciences (NSLS X25).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Schreiter, E., Sintchak, M., Guo, Y. et al. Crystal structure of the nickel-responsive transcription factor NikR. Nat Struct Mol Biol 10, 794–799 (2003). https://doi.org/10.1038/nsb985
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb985
This article is cited by
-
Grating-coupled interferometry reveals binding kinetics and affinities of Ni ions to genetically engineered protein layers
Scientific Reports (2020)
-
The lineage and diversity of putative amino acid sensor ACR proteins in plants
Amino Acids (2020)
-
Bacterial sensors define intracellular free energies for correct enzyme metalation
Nature Chemical Biology (2019)
-
Comprehensive mapping of the Helicobacter pylori NikR regulon provides new insights in bacterial nickel responses
Scientific Reports (2017)
-
Surface plasmon resonance and isothermal titration calorimetry to monitor the Ni(II)-dependent binding of Helicobacter pylori NikR to DNA
Analytical and Bioanalytical Chemistry (2016)