Abstract
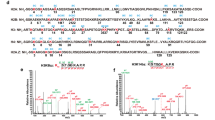
Yeast ESA1 is a member of the MYST subfamily of histone acetyltransferases (HATs), which use acetyl-coenzyme A (CoA) to acetylate specific Lys residues within histones to regulate gene expression. The structure of an ESA1–CoA complex reveals structural similarity to the catalytic core of the GCN5/PCAF subfamily of HAT proteins. Here we report additional structural and functional studies on ESA1 that demonstrate that histone acetylation proceeds through an acetyl-cysteine enzyme intermediate. This Cys residue is strictly conserved within the MYST members, suggesting a common mode of catalysis by this HAT subfamily. However, this mode of catalysis differs dramatically from the GCN5/PCAF subfamily, which mediate direct nucleophilic attack of the acetyl-CoA cofactor by the enzyme-deprotonated substrate lysine of the histone. These results demonstrate that different HAT subfamilies can use distinct catalytic mechanisms, which have implications for their distinct biological roles and for the development of HAT-specific inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Allfrey, V.G., Faulkner, R. & Mirsky, A.E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl. Acad. Sci. USA 51, 786–794 (1964).
Turner, B.M. Histone acetylation as an epigenetic determinant of long-term transcriptional competence. Cell. Mol. Life Sci. 54, 21–31 (1998).
Brownell, J.E. & Allis, C.D. Special HATs for special occasions: linking histone acetylation to chromatin assembly and gene activation. Curr. Opin. Genet. Dev. 6, 176–184 (1996).
Brownell, J.E. et al. Tetrahymena histone acetyltransferase A: a homolog of yeast GCN5p linking histone acetylation to gene activation. Cell 84, 843–851 (1996).
Smith, E.R. et al. ESA1 is a histone acetyltransferase that is essential for growth in yeast. Proc. Natl. Acad. Sci. USA 95, 3561–3565 (1998).
Bannister, A.J. & Kouzarides, T. The CBP co-activator is a histone acetyltransferase. Nature 384, 641–643 (1996).
Ogryzko, V.V., Schiltz, R.L., Russanova, V., Howard, B.H. & Nakatani, Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87, 953–959 (1996).
Grant, P.A. & Berger, S.L. Histone acetyltransferase complexes. Semin. Cell. Dev. Biol. 10, 169–177 (1999).
Grant, P.A. et al. Yeast GCN5 functions in two multisubunit complexes to acetylate nucleosomal histones: characterization of an Ada complex and the SAGA (Spt/Ada) complex. Genes Dev. 11, 1640–1650 (1997).
Allard, S. et al. NuA4, an essential transcription adaptor/histone H4 acetyltransferase complex containing ESA1p and the ATM-related cofactor Tra1p. EMBO J. 18, 5108–5119 (1999).
John, S. et al. The Something About Silencing protein, Sas3, is the catalytic subunit of NuA3, a yTAF(II)30-containing HAT complex that interacts with the Spt16 subunit of the yeast CP (Cdc68/Pob3)–FACT complex. Genes Dev. 14, 1196–1208 (2000).
Clarke, A.S., Lowell, J.E., Jacobson, S.J. & Pillus, L. ESA1p is an essential histone acetyltransferase required for cell cycle progression. Mol. Cell. Biol. 19, 2515–2526 (1999).
Borrow, J. et al. The translocation t(8;l6)(p11, p13) of acute myeloid leukaemia fuses a putative acetyltransferase to the CREB binding protein. Nature Genet. 14, 33–41 (1996).
Kamine, J., Elangovan, B., Subramanian, T., Coleman, D. & Chinnadurai, G. Identification of a cellular protein that specifically interacts with the essential cysteine region of the HIV-1 tat transactivator. Virology 216, 357–366 (1996).
Akhtar, A. & Becker, P.B. Activation of transcription through histone H4 acetylation by MOF, an acetyltransferase essential for dosage compensation in Drosophila. Mol. Cell 5, 367–375 (2000).
Reifsnyder, C., Lowell, J., Clarke, A. & Pillus, L. Yeast SAS silencing genes and human genes associated with AML and HIV-1 Tat interactions are homologous with acetyltransferases. Nature Genet. 14, 44–49 (1996).
Ehrenhofer-Murray, A.E., Rivier, D.H. & Rine, J. The role of Sas2, an acetyltransferase homologue of Saccharomyces cerevisiae, in silencing and ORC function. Genetics 145, 923–934 (1997).
Reid, J.L., Iyer, V.R., Brown, P.O. & Struhl, K. Coordinate regulation of yeast ribosomal protein genes is associated with targeted recruitment of ESA1 histone acetylase. Mol. Cell 6, 1297–1307 (2000).
Iizuka, M. & Stillman, B. Histone acetyltransferase HBO1 interacts with the ORC1 subunit of the human initiator protein. J. Biol. Chem. 274, 23027–23034 (1999).
Thomas, T., Voss, A.K., Chowdhury, K. & Gruss, P. Querkopf, a MYST family histone acetyltransferase, is required for normal cerebral cortex development. Development 127, 2537–2548 (2000).
Yan, Y., Barlev, N.A., Haley, R.H., Berger, S.L. & Marmorstein, R. Crystal structure of yeast ESA1 suggests a unified mechanism of catalysis and substrate binding by histone acetyltransferases. Mol. Cell 6, 1195–1205 (2000).
Clements, A. et al. Crystal structure of the histone acetyltransferase domain of the human P/CAF transcriptional regulator bound to coenzyme-A. EMBO J. 18, 3521–3532 (1999).
Rojas, J.R. et al. Structure of the Tetrahymena GCN5 bound to coenzyme-A and a histone H3 peptide. Nature 401, 93–98 (1999).
Trievel, R.C. et al. Crystal structure and mechanism of histone acetylation of the yeast GCN5 transcriptional coactivator. Proc. Natl. Acad. Sci. USA 96, 8931–8936 (1999).
Lo, W.-S. et al. Phosphorylation of serine 10 in histone H3 is functionally linked in vitro and in vivo to GCN5-mediated acetylation at lysine 14. Mol. Cell 5, 917–926 (2000).
Claiborne, A., Mallett, T.C., Yeh, J.I., Luba, J. & Parsonage, D. Structural, redox, and mechanistic parameters for cysteine-sulfenic acid function in catalysis and regulation. Adv. Protein Chem. 58, 215–276 (2001).
Creaven, M. et al. Control of the histone-acetyltransferase activity of Tip60 by the HIV-1 transactivator protein, Tat. Biochemistry 38, 8826–8830 (1999).
Wang, L., Liu, L. & Berger, S.L. Critical residues for histone acetylation by GCN5, functioning in Ada and SAGA complexes, are also required for transcriptional function in vivo. Genes Dev. 12, 640–653 (1998).
Tanner, K.G., Langer, M.R. & Denu, J.M. Kinetic mechanism of human histone acetyltransferase P/CAF. Biochemistry 39, 11961–11969 (2000).
Tanner, K.G., Langer, M.R., Kim, Y. & Denu, J.M. Kinetic mechanism of the histone acetyltransferase GCN5 from yeast. J. Biol. Chem. 275, 22048–22055 (2000).
Lau, O.D. et al. p300/CBP-associated factor histone acetyltransferase processing of a peptide substrate. Kinetic analysis of the catalytic mechanism. J. Biol. Chem. 275, 21953–21959 (2000).
Trievel, R.C., Li, F.-Y. & Marmorstein, R. Application of a novel fluorescence histone acetyltransferase enzyme assay to study the substrate specificity of human PCAF. Anal. Biochem. 287, 319–328 (2000).
Segel, I.H. Enzyme Kinetics: Behavior and Analysis of Rapid Equilibrium and Steady-state Enzyme Systems (John Wiley & Sons; 1993).
Thompson, P.R., Kurooka, H., Nakatani, Y. & Cole, P.A. Transcriptional coactivator protein p300. Kinetic characterization of its histone acetyltransferase activity. J. Biol. Chem. 276, 33721–33729 (2001).
Lau, O.D. et al. HATs off: Selective synthetic inhibitors of the histone acetyltransferases p300 and PCAF. Mol. Cell 5, 539–595 (2000).
Cleland, W.W. Statistical analysis of enzyme kinetic data. Methods Enzymol. 63, 103–138 (1979).
Chong, J.M. & Speicher, D.W. Determination of disulfide bond assignments and N-glycosylation sites of the human gastrointestinal carcinoma antigen GA733-2 (CO17-1A, EGP, KS1-4, KSA, and Ep-CAM). J. Biol. Chem. 276, 5804–5813 (2001).
Otwinowski, Z. in Proceedings of the CCP4 Study Weekend: Data Collection and Processing. (eds Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK, 1993).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Brünger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.Y. & Cowen, S.W. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brünger, A.T. & Krukowski, A. Slow-cooling protocols for crystallographic refinement by simulated annealing. Acta Crystallogr. A 46, 585–593 (1990).
Rice, L.M. & Brünger, A.T. Torsion angle dynamics: reduced variable conformational sampling enhances crystallographic structure refinement. Proteins 19, 277–290 (1994).
Laskowski, R.A., MacArthur, M.A., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Read, R.J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A 42, 140–149 (1986).
Acknowledgements
We thank S. Berger, N. Barlev and W. Lo for help with the HAT assay and valuable discussions; N. DiFlorio in the Wistar Microchemistry facility for help with the mass spectrometry analysis; A. Weaver for help on beamline F2 at CHESS; T. Penning and V. Heredia for help with the Cleland program; K. Zhao for PCAF HAT domain recombinant protein; and D. Christianson, A. Clements, R. Venkataramani and A. Poux for useful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yan, Y., Harper, S., Speicher, D. et al. The catalytic mechanism of the ESA1 histone acetyltransferase involves a self-acetylated intermediate. Nat Struct Mol Biol 9, 862–869 (2002). https://doi.org/10.1038/nsb849
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb849