Abstract
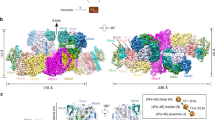
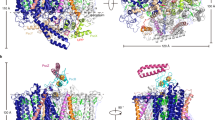
Catalase-peroxidase is a member of the class I peroxidase superfamily. The enzyme exhibits both catalase and peroxidase activities to remove the harmful peroxide molecule from the living cell. The 2.0 Å crystal structure of the catalase-peroxidase from Haloarcula marismortui (HmCP) reveals that the enzyme is a dimer of two identical subunits. Each subunit is composed of two structurally homologous domains with a topology similar to that of class I peroxidase. The active site of HmCP is in the N-terminal domain. Although the arrangement of the catalytic residues and the cofactor heme b in the active site is virtually identical to that of class I peroxidases, the heme moiety is buried inside the domain, similar to that in a typical catalase. In the vicinity of the active site, novel covalent bonds are formed among the side chains of three residues, including that of a tryptophan on the distal side of the heme. Together with the C-terminal domain, these covalent bonds fix two long loops on the surface of the enzyme that cover the substrate access channel to the active site. These features provide an explanation for the dual activities of this enzyme.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Cannac-Caffrey, V. et al. Biochimie 80, 1003–1011 (1998).
Zamocky, M., Janecek, S. & Koller, F. Gene 256, 169–182 (2000).
Welinder, K.G. Biochim. Biophys. Acta 1080, 215–220 (1992).
Murthy, M.R.N., Reid, T.J. III, Sicignano, A., Tanaka, N. & Rossmann, M.G. J. Mol. Biol. 152, 465–499 (1981).
Bravo, J. et al. Structure 3, 491–502 (1995).
Cendrin, F., Jouve, H.M., Gaillard, J., Thibault, P. & Zaccai, G. Biochim. Biophys. Acta 1209, 1–9 (1994).
Zamocky, M., Regelsberer, G., Jakopitsch, C. & Obinger C. FEBS Lett. 492, 177–182 (2001).
Finzel, B.C., Poulos, T.L. & Kraut, J. J. Biol. Chem. 259, 13027–13036 (1984).
Patterson, W.R. & Poulos, T.L. Biochemistry 34, 4331–4341 (1995).
Kleywegt, G.J. & Jones, T.A. Acta Crystallogr. D 50, 178–185 (1994).
Wengenack, N.L. et al. J. Infect. Dis. 176, 722–727 (1997).
Sivaraja, M., Goodin, D.B., Smith, M. & Hoffman, B.M. Science. 245, 738–740 (1989).
Chouchane, S., Lippai, I. & Magliozzo R.S. Biochemistry 39, 9975–9983 (2000)
Hillar, A. et al. Biochemistry 39, 5868–5875 (2000).
Ramaswamy, S. & Musser, J.M. Tuber. Lung Dis. 79, 3–29 (1998).
Yamada, Y. et al. Acta Crystallogr. D 57, 1157–1158 (2001).
Leslie, A.G.W. Proceedings of the CCP4 study weekend (eds Sawyer, L., Isaacs, N. & Bailey, S.) 44–51 (SERC Daresbury Laboratory, Warrington; 1993).
Evans, P.R. Proceedings of the CCP4 study weekend (eds Wilson, K.S., Davies, G., Ashton, A.W. & Bailey, S.) 97–102 (SERC Daresbury Laboratory, Warrington; 1997).
Otwinowski, Z. Proceedings of the CCP4 study weekend (eds Wolf, W., Evans, P.R. & Leslie, A.G.W.) 80–86 (SERC Daresbury Laboratory, Warrington; 1991).
Tanaka, N. Acta Crystallogr. A 33, 191–193 (1977).
Cowtan, K. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. J. Appl. Crystallogr. 26, 283–291 (1993).
Guex, N. & Peitsch, M.C. Electrophoresis 18, 2714–2723 (1997).
Gouet, P., Courcelle, E., Stuart, D.I. & Metoz, F. Bioinformatics 15, 305–308 (1999).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Esnouf, R.M. J. Mol. Graph. Model. 15, 132–134 (1997).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins 11, 281–282 (1991).
Acknowledgements
The present research was undertaken with the approval of the Photon Factory Advisory Committee, Japan, and the Japan Synchrotron Radiation Research Institute (JASRI). The authors wish to express their thanks to the staff at the Photon factory and SPring-8 for their help and the use of the diffractometer. The project was partly supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan; the ACT-JST Program, Japan Science and Technology Corporation; and research grant from Salt-Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Yamada, Y., Fujiwara, T., Sato, T. et al. The 2.0 Å crystal structure of catalase-peroxidase from Haloarcula marismortui. Nat Struct Mol Biol 9, 691–695 (2002). https://doi.org/10.1038/nsb834
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb834
This article is cited by
-
Catalase from the Antarctic Fungus Aspergillus fumigatus I-9–Biosynthesis and Gene Characterization
Indian Journal of Microbiology (2023)
-
The structure–function correlation of Cicer arietinum catalase 4 (Ca catalase 4): the key scavenger enzyme during chickpea–Fusarium interplay
Proceedings of the Indian National Science Academy (2023)
-
Peroxygenase reactions catalyzed by cytochromes P450
JBIC Journal of Biological Inorganic Chemistry (2014)
-
Multifunctional enzymes in archaea: promiscuity and moonlight
Extremophiles (2013)
-
Characterization of alcohol dehydrogenase from the haloalkaliphilic archaeon Natronomonas pharaonis
Extremophiles (2008)