Abstract
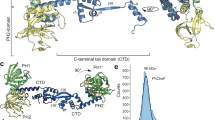
The protein CheZ, which has the last unknown structure in the Escherichia coli chemotaxis pathway, stimulates the dephosphorylation of the response regulator CheY by an unknown mechanism. Here we report the co-crystal structure of CheZ with CheY, Mg2+ and the phosphoryl analog, BeF3−. The predominant structural feature of the CheZ dimer is a long four-helix bundle composed of two helices from each monomer. The side chain of Gln 147 of CheZ inserts into the CheY active site and is essential to the dephosphorylation activity of CheZ. Gln 147 may orient a water molecule for nucleophilic attack, similar to the role of the conserved Gln residue in the RAS family of GTPases. Similarities between the CheY–CheZ and Spo0F–Spo0B structures suggest a general mode of interaction for modulation of response regulator phosphorylation chemistry.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bren, A. & Eisenbach, M. J. Bacteriol. 182, 6865–6873 (2000).
Bourret, R.B. & Stock, A.M. J. Biol. Chem. 277, 9625–9628 (2002).
Yi, T.M., Huang, Y., Simon, M.I. & Doyle, J. Proc. Natl. Acad. Sci. USA 97, 4649–4653 (2000).
Duke, T.A., Novere, N.L. & Bray, D. J. Mol. Biol. 308, 541–553 (2001).
Stock, A.M., Robinson, V.L. & Goudreau, P.N. Annu. Rev. Biochem. 69, 183–215 (2000).
Segall, J.E., Manson, M.D. & Berg, H.C. Nature 296, 855–857 (1982).
Stock, A.M. & Stock, J.B. J. Bacteriol. 169, 3301–3311 (1987).
Boesch, K.C., Silversmith, R.E. & Bourret, R.B. J. Bacteriol. 182, 3544–3552 (2000).
Sanna, M.G. & Simon, M.I. J. Bacteriol. 178, 6275–6280 (1996).
McEvoy, M.M., Bren, A., Eisenbach, M. & Dahlquist, F.W. J. Mol. Biol. 289, 1423–1433 (1999).
Blat, Y. & Eisenbach, M. Biochemistry 35, 5679–5683 (1996).
Sanna, M.G., Swanson, R.V., Bourret, R.B. & Simon, M.I. Mol. Microbiol. 15, 1069–1079 (1995).
Zhu, X., Volz, K. & Matsumura, P. J. Biol. Chem. 272, 23758–23764 (1997).
Lukat, G.S., Stock, A.M. & Stock, J.B. Biochemistry 29, 5436–5442 (1990).
Stock, A.M. et al. Biochemistry 32, 13375–13380 (1993).
Blat, Y., Gillespie, B., Bren, A., Dahlquist, F.W. & Eisenbach, M. J. Mol. Biol. 284, 1191–1199 (1998).
Wang, H. & Matsumura, P. Mol. Microbiol. 19, 695–703 (1996).
Yan, D. et al. Proc. Natl. Acad. Sci. USA 96, 14789–14794 (1999).
Lee, S.Y. et al. J. Biol. Chem. 276, 16425–16431 (2001).
Blat, Y. & Eisenbach, M. J. Biol. Chem. 271, 1226–1231 (1996).
Saul, F.A. & Alzari, P.M. Methods Mol. Biol. 66, 11–23 (1996).
Silversmith, R.E., Smith, J.G., Guanga, G.P., Les, J.T. & Bourret, R.B. J. Biol. Chem. 276, 18478–18484 (2001).
Lee, S.Y. et al. Nature Struct. Biol. 8, 52–56 (2001).
Robinson, V.L., Buckler, D. & Stock, A.M. Nature Struct. Biol. 7, 626–633 (2000).
Lukat, G.S., Lee, B.H., Mottonen, J.M., Stock, A.M. & Stock, J.B. J. Biol. Chem. 266, 8348–8354 (1991).
Maegley, K.A., Admiraal, S.J. & Herschlag, D. Proc. Natl. Acad. Sci. USA 93, 8160–8166 (1996).
Lowy, D.R. & Willumsen, B.M. Annu. Rev. Biochem. 62, 851–891 (1993).
Scheffzek, K. et al. Science 277, 333–338 (1997).
Zapf, J., Sen, U., Madhusudan, Hoch, J.A. & Varughese, K.I. Structure 8, 851–862 (2000).
Mourey, L. et al. J. Biol. Chem. 276, 31074–31082 (2001).
Reizer, J., Reizer, A., Perego, M. & Saier, M.H.J. Microb. Comp. Genomics 2, 103–111 (1997).
Hess, J.F., Bourret, R.B. & Simon, M.I. Methods Enzymol. 200, 188–204 (1991).
Bourret, R.B., Hess, J.F. & Simon, M.I. Proc. Natl. Acad. Sci. USA 87, 41–45 (1990).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1996).
Terwilliger, T.C. & Berendzen, J. Acta Crystallogr. D 55, 849–861 (1999).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–911 (1998).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110–119 (1991).
Esnouf, R.M. Acta Crystallogr. D 55, 938–940 (1999).
Nicholls, A., Sharp, K. & Honig, B. Proteins 11, 281–296 (1991).
Carson, M. J. Appl. Crystallogr. 24, 958–961 (1991).
Acknowledgements
We thank J. Snyder and M. Kimple (UNC Chapel Hill) for help collecting the native data set; D. Wemmer and S.-Y. Lee (UC Berkeley) for sharing the BeF3− parameter and topology files and the coordinates for the CheY–BeF3−-FliM structure before they were publicly accessible; G. Zhang (National Jewish Medical and Research Center) and H. Ke, D. Worthylake and M. Redinbo (UNC Chapel Hill) for helpful discussion; L. Betts (UNC X-ray facility) for technical support; X. Chen, M. Churchill and the X-ray Core Facility at the University of Colorado Health Science Center (UCHSC) for their support for the completion of R.Z.'s project at UCHSC; and the Brookhaven beamline staff for help with data collection.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Zhao, R., Collins, E., Bourret, R. et al. Structure and catalytic mechanism of the E. coli chemotaxis phosphatase CheZ. Nat Struct Mol Biol 9, 570–575 (2002). https://doi.org/10.1038/nsb816
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb816