Abstract
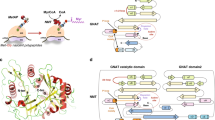
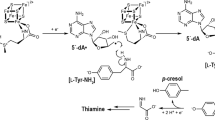
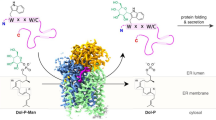
N-myristoyltransferase (Nmt) attaches myristate to the N-terminal glycine of many important eukaryotic and viral proteins. It is a target for anti-fungal and anti-viral therapy. We have determined the structure, to 2.9 Å resolution, of a ternary complex of Saccharomyces cerevisiae Nmt1p with bound myristoylCoA and peptide substrate analogs. The model reveals structural features that define the enzyme's substrate specificities and regulate the ordered binding and release of substrates and products. A novel catalytic mechanism is proposed involving deprotonation of the N-terminal ammonium of a peptide substrate by the enzyme's C-terminal backbone carboxylate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Towler, D.A. et al. Purification and characterization of myristoylCoA:protein N-myristoyltransferase. Proc. Natl. Acad. Sci. USA 84, 2708–2712 (1987).
Boutin, J. Myristoylation. Cell. Signal. 9, 15–35 (1997).
Bhatnagar, R. S. & Gordon, J. I. Understanding covalent modifications of proteins by lipids: where cell biology and biophysics mingle. Trends Cell Biol. 7, 14–18 (1997).
Bryant, M. L. et al. Incorporation of 12-methoxydodecanoate into the human immunodeficiency virus I gag polyprotein precursor inhibits its proteolytic processing and viral production in a chronically infected human lymphoid cell line. Proc. Natl. Acad. Sci. USA 88, 2055–2059 (1991).
Duronio, R. J., Towler, D. A., Heuckeroth, R. O. & Gordon, J. I. Disruption of the yeast N-myristoyl transferase gene causes recessive lethality. Science 243, 796–800 (1989).
Weinberg, R. A. et al. Genetic studies reveal that myristoylCoA:protein N- myristoyltransferase is an essential enzyme in Candida albicans. Mol. Microbiol. 16, 241–250 (1995).
Lodge, J. K., Jackson-Machelski, E., Toffaletti, D. L., Perfect, J. R. & Gordon, J. I. Targeted gene replacement demonstrates that myristoylCoA:protein N-myristoyltransferase is essential for viability of Cryptococcus neoformans. Proc. Natl. Acad. Sci. USA 91, 12008–12012 (1994).
Sikorski, J. A. et al. Selective peptidic and peptidomimetic inhibitors of Candida albicans myristoylCoA:protein N-myristoyltransferase: A new approach to antifungal therapy. Biopolymers 43, 43–71 (1997).
White, T. C., Marr, K. A. & Bowden, P. A. Clinical, cellular, and molecular factors that contribute to antifungal drug resistance. Clin. Microbiol. Rev. 11, 382–402 (1998).
Rudnick, D. A. et al. Kinetic and structural evidence for a sequential ordered Bi Bimechanism of catalysis by Saccharomyces cerevisiae myristoylCoA:protein N myristoyltransferase. J. Biol. Chem. 266, 9732–9739 (1991).
Bhatnagar, R. S. et al. Titration calorimetric analysis of acylCoA recognition by myristoylCoA:protein N-myristoyltransferase. Biochemistry 36, 6700–6708 (1997).
Rocque, W. J., McWherter, C. A., Wood, D. C. & Gordon, J. I. A comparative analysis of the kinetic mechanism and peptide substrate specificity of human and Saccharomyces cerevisiae myristoylCoA:protein N-myristoyltransferase. J. Biol. Chem. 268, 9964–9971 (1993).
Weston, S. A. et al. Crystal structure of the anti-fungal targed N-myristoyltransferase. Nature Struct. Biol. 5, 213–221 (1998).
Paige, L. A., Zheng, G-Q., DeFrees, S. A., Cassady, J. M. & Geahlen, R. L. S-(2- oxopentadecyl)-CoA, a nonhydrolyzable analog of myristoylCoA, is a potent inhibitor of myristoylCoA:protein N-myristoyltransferase. J. Med. Chem., 32, 1665–1667 (1989).
Devadas, B. et al. Design and synthesis of potent and selective dipeptide inhibitors of Candida albicans myristoylCoA:protein N-myristoyltransferase. J. Med. Chem., 38, 1837–1840 (1995).
Rudnick, D. A. et al. Analogs of palmitoylCoA that are substrates for myristoylCoA:protein N-myristoyltransferase. Proc. Natl. Acad. Sci. USA 89, 10507–10511 (1992).
Towler, D. A., Gordon, J. I., Adams, S. P. & Glaser, L. The biology and enzymology of eukaryotic protein acylation. Annu. Rev. Biochem. 57, 69–99 (1988).
Bairoch, A., Bucher, P. & Hofmann, K. The PROSITE database, its status in 1997. Nucleic Acids Res. 25, 217–221 (1997).
Zhang, L., Jackson-Machelski, E. & Gordon, J. I. Genetic and biochemical studies of S. cerevisiae myristoylCoA:protein N-myristoyltransferase mutants. J. Biol. Chem. 271, 33131–33140 (1996).
Peseckis, S.M. & Resh, M.D. Fatty acyl transfer by N-myristyl transferase is dependent upon conserved cysteine and histidine residues. J. Biol. Chem. 269, 30888–30892 (1994).
Rudnick, D. A., Johnson, R. L. & Gordon, J. I. Studies of the catalytic activities and substrate specificities of Saccharomyces cerevisiae myristoylCoA:protein N-myristoyltransferase deletion mutants and human/yeast Nmt chimeras in Eschericia coli and S. cerevisiae . J. Biol. Chem. 267, 23852–23861 (1992).
Zheng, W., Johnston, S. A. & Joshua-Tor, L. The unusual active site of Gal6/bleomycin hydrolase can act as a carboxypeptidase, aminopeptidase, and peptide ligase. Cell 93, 103–109 (1998).
Hendrickson, W. A., Horton, J. R. & LeMaster, D. Selenomethionine proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
Otwinowski, Z. in Proceedings of the CCP4 study weekend: data collection and processing. (eds Sawyers, L. Isaacs, N. & Bailey, S.) 56–62 (SERC Daresbury Laboratory, Warrington, UK; 1993).
Terwilliger, T. C., Kim, S.-H. & Eisenberg, D. Generalized method of determining heavy-atom positions using the difference Patterson function. Acta Crystallogr. A 43, 1–5 (1987).
De La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Meth. Enz. 276, 472–493 (1997).
Abrahams, J. P. & Leslie, A. G. W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D52, 32–42 (1996).
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Terwilliger, T. C. & Eisenberg, D. Unbiased Three-Dimensional Refinement of Heavy-Atom Parameters by Correlation of Origin-Removed Patterson Functions. Acta Crystallogr. A39, 813–817 (1983).
Hall, T. M. T., Porter, J. A., Beachy, P. A. & Leahy, D. J. A potential catalytic site revealed by the 1.7Å crystal structure of the amino-terminal signalling domain of Sonic hedgehog. Nature 378, 212–216 (1995).
Brünger, A. T. et al. Crystallography and NMR system: a new software system for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998).
Brünger, A. T. The free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–474 (1992).
Carson, M. Ribbons. Meth. Enz. 277, 493–505 (1997).
Nicholls, A. & Honig, B. A rapid finite difference algorithm, utilizing successive over- relaxation to solve the Poisson-Boltzman equation. J. Comput. Chem. 12, 435–445 (1991).
Engh, R. A. & Huber, R. Accurate bond and angle parameters for X-ray protein structure refinement. Acta Crystallogr. A47, 392–400 (1991).
Acknowledgements
We thank B. Devadas, J. Sikorski and C. McWherter (G.D. Searle) for SC-58272, C. Ogata at NSLS, and M. Soltis, and P. Kuhn at SSRL for help with data collection, and J. Kuriyan for helpful comments. This work was supported in part by grants from the NIH and Monsanto.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bhatnagar, R., Fütterer, K., Farazi, T. et al. Structure of N-myristoyltransferase with bound myristoylCoA and peptide substrate analogs. Nat Struct Mol Biol 5, 1091–1097 (1998). https://doi.org/10.1038/4202
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/4202
This article is cited by
-
Molecular basis of antibiotic self-resistance in a bee larvae pathogen
Nature Communications (2022)
-
Drug discovery in leishmaniasis using protein lipidation as a target
Biophysical Reviews (2021)
-
High-resolution snapshots of human N-myristoyltransferase in action illuminate a mechanism promoting N-terminal Lys and Gly myristoylation
Nature Communications (2020)
-
Validation of N-myristoyltransferase as an antimalarial drug target using an integrated chemical biology approach
Nature Chemistry (2014)
-
An improved method and cost effective strategy for soluble expression and purification of human N-myristoyltransferase 1 in E. coli
Molecular and Cellular Biochemistry (2014)