Abstract
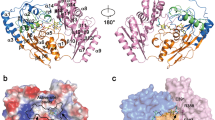
5-Aminolaevulinate dehydratase (ALAD) is a homo-octameric metallo-enzyme that catalyses the formation of porphobilinogen from 5-aminolaevulinic acid. The structure of the yeast enzyme has been solved to 2.3 Å resolution, revealing that each subunit adopts a TIM barrel fold with a 39 residue N-terminal arm. Pairs of monomers wrap their arms around each other to form compact dimers and these associate to form a 422 symmetric octamer. All eight active sites are on the surface of the octamer and possess two lysine residues (210 and 263), one of which, Lys 263, forms a Schiff base link to the substrate. The two lysine side chains are close to two zinc binding sites one of which is formed by three cysteine residues (133,135 and 143) while the other involves Cys 234 and His 142. ALAD has features at its active site that are common to both metallo- and Schiff base-aldolases and therefore represents an intriguing combination of both classes of enzyme. Lead ions, which inhibit ALAD potently, replace the zinc bound to the enzyme's unique triple-cysteine site.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Jordan, P.M. Biosynthesis of tetrapyrroles. In New Comprehensive Biochemistry Vol. 19 (Ed. Neuberger, A. & Van Deenen, L.L.N) 1–65 (Elsevier, Amsterdam; 1991).
Jordan, P.M. Highlights in haem biosynthesis. Curr. Opin. Struct Biol. 4, 902–911 (1994).
Warren, M.J. & Scott, A.I. Tetrapyrrole assembly and modification into the ligands of biologically functional cofactors. Trends Biochem. Sci. 15, 486–491 (1990).
Jaffe, E.K. Porphobilinogen synthase, the first source of heme's asymmetry. J. Bioenerg. Biomemb. 27, 169–179 (1995).
Doss, M. et al. New types of hepatic porphyria with porphobilinogen synthase defect and intermittant acute clinical manifestation. Klin. Wochenschr. 57, 1123–1127 (1979).
Simons, T.J.B. The affinity of human erythrocyte porphobilinogen synthase for Zn2+ and Pb2+. Eur. J. Biochem. 234, 178–183 (1995).
Guo, G.G., Gu, M. & Etlinger, J.D. 240-kDa proteasome inhibitor (CF-2) is identical to 5-aminolevulinic acid dehydratase. J. Biol. Chem. 269, 12399–12402 (1994).
Coux, O., Tanaka, K. & Goldberg, A.L. Structure and functions of the 20S and 26S proteasomes. Ann. Rev. Biochem. 65, 801–847 (1996).
Hester, G. et al. The crystal structure of fructose-1,6-bisphosphate aldolase from Drosophila melanogaster at 2.5 Å resolution. FEBS Lett. 292, 237–242 (1991).
Gamblin, S.J. et al. Activity and specificity of human aldolases. J. Mol. Biol. 219, 573–576 (1991).
Blom, N.S. et al. Novel active site in E. coli fructose-1, 6-bisphosphate aldolase. Nature Struct. Biol. 3, 856–862 (1996).
Cooper, S.J. et al. The crystal structure of a class II fructose-1,6-bisphosphate aldolase shows a novel metal-binding active site embedded in a familiar fold. Structure 4, 1303–1315 (1996).
Mancia, F. et al. Howcoenzyme B12 radicals are generated: the crystal structure of methylmalonyl-coenzyme A mutase at 2 Å resolution. Structure 4, 339–350 (1996).
Wu, W. et al. The quaternary structure of 5-aminolevulinic acid dehydratase from bovine liver. Proc. Natl. Acad. Sci. USA. 71, 1767–1770 (1974).
Pilz, I. et al. Small angle X-ray scattering study of bovine porphobilinogen synthase. Biol. Chem. Hoppe-Seyler 369, 1099–1103 (1988).
Senior, N. PhD Thesis, University of London, UK (1996).
Boese, Q.F. et al. 5-aminolevulinic acid dehydratase in pea. Identification of an unusual metal-binding domain in the plant enzyme. J. Biol. Chem. 266, 17060–17066.
Dent, A. et al. Two different zinc sites in bovine 5-aminolevulinate dehydratase distinguished by extended X-ray absorption fine structure. Biochemistry 29, 7822–7828.
Erskine, P.T. PhD Thesis, University of London (1997).
Blom, N. and Sysgusch, J. Product binding and the role of the C-terminal region in class I D-fructose-1,6-bisphosphate aldolase. Nature Struct. Biol. 4, 36–39 (1997).
Neier, R. Chemical synthesis of porphobilinogen and studies of its biosynthesis. Adv. Nitrogen Heterocycles 2, 35–146 (1996).
Wetmur, J.G. et al. Human 5-aminolevulinate dehydratase: nucleotide sequence of a full length cDNA clone. Proc. Natl. Acad. Sci. USA. 83, 7703–7707 (1986).
Wetmur, J.G. et al. Molecular characterisation of the human 5-aminolevulinate dehydratase 2 (ALAD2) allele: implications for molecular screening of individuals for genetic susceptibility to lead poisoning. Am. J. Hum. Genet. 49, 757–763 (1991).
Ishida, N. et al. Cloning and expression of the defective genes from a patient with 5-aminolevulinate dehydratase porphyria. J. Clin. Invest. 89, 1431–1437 (1992).
Plewinska, M. et al. 5-Aminolevulinate dehydratase deficient porphyria: identification of the molecular lesions in a severely affected homozygote. Am. J. Hum. Genet. 49, 167–174 (1991).
Brennan, M.J.W. & Cantrill, R.C. 5-Aminolaevulinic acid is a potent agonist for GABA autoreceptors. Nature 280, 514–515 (1979).
Groll, M. et al. Structure of 20S proteasome from yeast at 2.4 Å resolution. Nature 386, 463–471 (1997).
Senior, N. et al. Comparative studies of the 5-aminolevulinic acid dehydratase from P. sativum, E. coli and S. cerevisiae. Biochem. J. 320, 401–412 (1996).
Erskine, P.T. et al. Crystallisation of 5-aminolaevulinic acid dehydratase from Escherichia coli and Saccharomyces cerevisiae and preliminary X-ray characterisation of the crystals. Prot. Sci. 6, 1–3. (1997).
CCP4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D50, 760–763 (1994).
Jones, T.A. et al. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Kraulis, P.J. MOLSCRIPT: a program to produce detailed and schematic plots of protein structure. J. Appl. Crystallogr. 24, 946–950 (1991).
Evans, S.V. Setor - hardware lighted 3-dimensional solid model representations of macromolecules. J. Mol. Graphics. 11, 134 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Erskine, P., Senior, N., Awan, S. et al. X-ray structure of 5-aminolaevulinate dehydratase, a hybrid aldolase. Nat Struct Mol Biol 4, 1025–1031 (1997). https://doi.org/10.1038/nsb1297-1025
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1297-1025
This article is cited by
-
Biochemical and molecular characterization of a novel porphobilinogen synthase from Corynebacterium glutamicum
World Journal of Microbiology and Biotechnology (2023)
-
Heme biosynthesis depends on previously unrecognized acquisition of iron-sulfur cofactors in human amino-levulinic acid dehydratase
Nature Communications (2020)
-
Antimicrobial activity of metals: mechanisms, molecular targets and applications
Nature Reviews Microbiology (2013)
-
Redox and metal-regulated oligomeric state for human porphobilinogen synthase activation
Amino Acids (2011)