Abstract
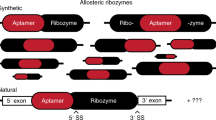
RNA transcripts containing the hammerhead ribozyme have been engineered to self-destruct in the presence of specific nucleoside 3′,5′-cyclic monophosphate compounds. These RNA molecular switches were created by a new combinatorial strategy termed 'allosteric selection,' which favors the emergence of ribozymes that rapidly self-cleave only when incubated with their corresponding effector compounds. Representative RNAs exhibit 5,000-fold activation upon cGMP or cAMP addition, display precise molecular recognition characteristics, and operate with catalytic rates that match those exhibited by unaltered ribozymes. These findings demonstrate that a vast number of ligand-responsive ribozymes with dynamic structural characteristics can be generated in a massively parallel fashion. Moreover, optimized allosteric ribozymes could serve as highly selective sensors of chemical agents or as unique genetic control elements for the programmed destruction of cellular RNAs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Breaker, R.R. In vitro selection of catalytic polynucleotides. Chem. Rev. 97, 371–390 ( 1997).
Pan, T. Novel and variant ribozymes obtained through in vitro selection. Curr. Opin. Chem. Biol. 1, 17–25 (1997).
Frauendorf, C. & Jäschke, A. Catalysis of organic reactions by RNA. Angew. Chem. Int. Edn. Engl. 37, 1378–1381 (1998).
Kurz, M. & Breaker, R.R. In vitro selection of nucleic acid enzymes. Curr. Top. Microbiol. Immunol. 243, 137–156 (1999).
White, H.B. Coenzymes as fossils of an earlier metabolic state. J. Mol. Evol. 7, 101–104 ( 1976).
Benner, S.A. et al. Natural selection, protein engineering, and the last riboorganism: rational building in biochemistry. Cold Spring Harbor Symp. Quant. Biol. 52, 53–63 ( 1987).
Benner, S.A., Ellington, A.D. & Tauer, A. Modern metabolism as a palimpsest of the RNA world. Proc. Natl. Acad. Sci. USA 86, 7054– 7058 (1989).
Hirao, I. & Ellington, A.D. Re-creating the RNA world. Curr. Biol. 5, 1017–1022 (1995).
Soukup, G.A. & Breaker, R.R. Allosteric ribozymes. In Ribozymes: biology and biotechnology. (eds Gaur, R.K. & Krupp, G.) (Eaton Publishing, Natick, Massachusetts; 1999).
Porta, H. & Lizardi, P.M. An allosteric hammerhead ribozyme. Bio/Technol. 13, 161–164 (1995).
Kuwabara, T. et al. A novel allosterically trans-activated ribozyme, the maxizyme, with exceptional specificity in vitro and in vivo. Mol. Cell 2, 617–627 ( 1998).
Westhof, E., Masquida, B. & Jaeger, L. RNA tectonics: towards RNA design. Folding & Design 1, R78–R88 (1996).
Batey, R.T. & Doudna, J.A. The parallel universe of RNA folding. 5, 337–340 ( 1998).
Gold, L., Polisky, B., Uhlenbeck, O. & Yarus, M. Diversity of oligonucleotide functions. Annu. Rev. Biochem. 64, 763–797 (1995).
Osborne, S.E. & Ellington, A.D. Nucleic acid selection and the challenge of combinatorial chemistry. Chem. Rev. 97, 349–370 (1997).
Tang, J. & Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 4, 453– 459 (1997).
Soukup, G.A. & Breaker, R.R. Engineering precision RNA molecular switches. Proc. Natl. Acad. Sci. USA 96, 3584–3589 (1999).
Soukup, G.A. & Breaker, R.R. Design of allosteric hammerhead ribozymes activated by ligand-induced structure stabilization. Structure Folding & Design 7, 783– 791 (1999).
Perutz, M. Mechanisms of cooperativity and allosteric regulation in proteins. (Cambridge University Press, New York, New York; 1994).
Araki, M., Okuno, Y., Hara, Y. & Sugiura, Y. Allosteric regulation of a ribozyme activity through ligand-induced conformational change. Nucleic Acids Res. 26, 3379–3384 (1998).
Patel, D.J. et al. Structure, recognition and adaptive binding in RNA aptamer complexes. J. Mol. Biol. 272, 645– 664 (1997).
Robertson, M.P. & Ellington, A.D. In vitro selection of an allosteric ribozyme that transduces analytes to amplicons. Nature Biotechnol. 17, 62– 66 (1999).
Nerbonne, J.M., Richard, S., Nargeot, J. & Lester, H.A. New photoactivatable cyclic nucleotides produce intracellular jumps in cyclic AMP and cyclic GMP concentrations. Nature 310, 74– 76 (1984).
Ho, H.C., Teo, T.S., Desai, R. & Wang, J.H. Catalytic and regulatory properties of two forms of bovine heart cyclic nucleotide phosphodiesterase. Biochim. Biophys. Acta 429, 461– 473 (1976).
Tang, J. & Breaker, R.R. Examination of the catalytic fitness of the hammerhead ribozyme by in vitro selection. RNA 3, 914–925 (1997).
Tang, J. & Breaker, R.R. Mechanism for allosteric inhibition of an ATP-sensitive ribozyme. Nucleic Acids Res. 26, 4214–4221 (1998).
Lin, E.C.C. & Lynch, A.S. Regulation of gene expression in Escherichia coli. (R.G. Landes Co, Austin, Texas; 1996).
Denison, C. & Kodadek, T. Small-molecule-based strategies for controlling gene expression. Chem. Biol. 5, R129–R145 (1998).
Werstuck, G. & Green, M.R. Controlling gene expression in living cells through small molecule–RNA interactions. Science 282, 296–298 (1998).
Hertel, K.J. et al. Numbering system for the hammerhead. Nucleic Acids Res. 20, 3252 (1992).
Acknowledgements
We thank J. Shin for contributions to Figures 3 and 8, and we thank members of the Breaker laboratory for helpful discussions. We also thank T.E. Shrader at the Albert Einstein College of Medicine for the kind gift of the construct to express His-tagged T7 RNAP. Funding for this work was provided by grants from the National Institutes of Health (NIH), the Defense Advanced Research Projects Agency (DARPA) and the Yale Diabetes and Endocrinology Research Center (DERC). M.K. was supported by Sankyo Co. Ltd. of Japan and G.A.S. was supported by a Seessel postdoctoral fellowship from Yale University. R.R.B. is the recipient of a Hellman family fellowship and a fellowship from the David and Lucile Packard Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Koizumi, M., Soukup, G., Kerr, J. et al. Allosteric selection of ribozymes that respond to the second messengers cGMP and cAMP. Nat Struct Mol Biol 6, 1062–1071 (1999). https://doi.org/10.1038/14947
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/14947
This article is cited by
-
Engineering synthetic RNA devices for cell control
Nature Reviews Genetics (2022)
-
A multiplexed, automated evolution pipeline enables scalable discovery and characterization of biosensors
Nature Communications (2021)
-
Reprogramming eukaryotic translation with ligand-responsive synthetic RNA switches
Nature Methods (2016)
-
Identification of the Same Na+-Specific DNAzyme Motif from Two In Vitro Selections Under Different Conditions
Journal of Molecular Evolution (2015)
-
Aptamers for allosteric regulation
Nature Chemical Biology (2011)