Abstract
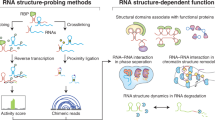
A detailed understanding of the functions and interactions of biological macromolecules requires knowledge of their molecular structures. Structural genomics, the systematic determination of all macromolecular structures represented in a genome, is focused at present exclusively on proteins. It is clear, however, that RNA molecules play a variety of significant roles in cells, including protein synthesis and targeting, many forms of RNA processing and splicing, RNA editing and modification, and chromosome end maintenance. To comprehensively understand the biology of a cell, it will ultimately be necessary to know the identity of all encoded RNAs, the molecules with which they interact and the molecular structures of these complexes. This report focuses on the feasibility of structural genomics of RNA, approaches to determining RNA structures and the potential usefulness of an RNA structural database for both predicting folds and deciphering biological functions of RNA molecules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mitchell, J.R., Cheng, J. & Collins, K. Mol. Cell. Biol. 19, 567– 576 (1999).
Moore, P.B. Annu. Rev. Biochem. 68, 287–300 (1999).
Leontis, N.B. & Westhof, E. Quart. Rev. Biophys. 31, 399–455 (1998).
Schmitz, U. et al. RNA 5, 1419–1429 (1999).
Batey, R.T., Rambo, R.P., Lucast, L., Rha, B. & Doudna, J.A. Science 287, 1232– 1239 (2000).
Jovine, L. et al. Structure 8, 527–540 (2000).
Ban, N., Nissen, P., Hansen, J., Moore, P.B. & Steitz, T.A. Science 289, 905– 920 (2000).
Ferre-D'Amare, A.R., Zhou, K. & Doudna, J.A. Nature 395, 567– 574 (1998).
Cate, J.H. et al. Science 273, 1678–1685 (1996).
Butcher, S.E., Dieckmann, T. & Feigon, J. EMBO J. 16, 7490– 7299 (1997).
Bourdeau, V., Ferbeyre, G., Pageau, M., Paquin, B. & Cedergren R. Nucleic Acids. Res. 27, 4457– 4467 (1999).
Lowe, T.M. & Eddy, S.R. Science 283, 1168–1171 (1999).
Omer, A.D. et al. Science 288, 517–522 (2000).
Latham, J.A. & Cech, T.R. Science 245, 276–282 (1989).
Christian, E.L. & Yarus, M. J. Mol. Biol. 228, 743–758 (1992).
Harris, M.E., Kazantsev, A.V., Chen, J.L. & Pace, N.R. RNA 3, 561–576 ( 1997).
Ryder, S.P., Ortoleva-Donnelly, L., Kosek, A.B. & Strobel, S.A. Methods Enzymol. 317, 92–109 (2000).
Stowell, M.H., Miyazawa, A. & Unwin, N. Curr. Opin. Struct. Biol. 8, 595–600 (1998).
Ferre-D'Amare, A.R. & Doudna, J.A. Nucleic Acids Res. 24, 977–978 ( 1996).
Doudna, J.A., Grosshans, C., Gooding, A. & Kundrot, C.E. Proc. Natl. Acad. Sci. USA 90, 7829– 7833 (1993).
Scott, W.G. et al. J. Mol. Biol. 250, 327– 332 (1995).
Golden, B.L., Podell, E.R., Gooding, A.R. & Cech, T.R. J. Mol. Biol. 270, 711–723 (1997).
Ferre-D'Amare, A.R., Zhou, K., Doudna, J.A. J. Mol. Biol. 279, 621–631 (1998).
Ferre-D'Amare, A.R. & Doudna, J.A. J. Mol. Biol. 295, 541–556 ( 2000).
Burley, S.K. et al. Nature Genet. 23, 151– 157 (1999).
Michel, F. & Westhof, E. J. Mol. Biol. 216, 585–610 (1990).
Golden, B.L., Gooding, A.R., Podell, E.R. & Cech, T.R. Science 282, 259–264 ( 1998).
Tanner, N.K. et al. Curr. Biol. 4, 488– 498 (1994).
Schluenzen, F. et al. Cell 102, 615 ( 2000).
Wimberly, B.T. et al. Nature 407, 327–339 (2000).
Carter, A.P. et al. Nature 407, 340–348 (2000).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Doudna, J. Structural genomics of RNA. Nat Struct Mol Biol 7 (Suppl 11), 954–956 (2000). https://doi.org/10.1038/80729
Issue Date:
DOI: https://doi.org/10.1038/80729
This article is cited by
-
Thermodynamics of unfolding mechanisms of mouse mammary tumor virus pseudoknot from a coarse-grained loop-entropy model
Journal of Biological Physics (2022)
-
QRNAS: software tool for refinement of nucleic acid structures
BMC Structural Biology (2019)
-
Nature inspired optimization algorithm for prediction of “minimum free energy” “RNA secondary structure”
Journal of Reliable Intelligent Environments (2019)
-
Bioinformatic Analysis of the Contribution of Primer Sequences to Aptamer Structures
Journal of Molecular Evolution (2008)