Abstract
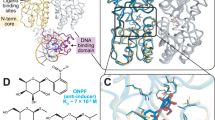
The Zn2Cys6 DNA-binding domain has been identified by sequence homology in approximately forty fungal proteins, including the K. lactis LAC9 transcriptional activator. Using 1H NMR spectroscopy, we have determined the solution structure of a cadmium-substituted form of the LAC9 DNA-binding domain. We have complemented this approach by applying a series of 113Cd-1H NMR experiments, including several novel heteroTOCSY-based techniques. The DNA-binding domain forms a core of two α-helix/extended strand segments around the Cd2 binuclear cluster, with a network of amide proton-cysteinyl Sγ hydrogen bonds stablizing the cluster. Comparison with other Zn2Cys6 domain structures provides insight into the common structural elements used in metal coordination and DNA binding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Pan, T. & Coleman, J.E. Structure and function of the Zn(II) binding site within the DNA-binding domain of the GAL4 transcription factor. Proc. natn. Acad. Sci. U.S.A. 86, 3145–3149 (1989).
Gardner, K.H., Pan, T., Narula, S., Rivera, E. & Coleman, J.E. Structure of the binuclear metal-binding site in the GAL4 transcription factor. Biochemistry 30, 11292–11302 (1991).
Baleja, J.D., Marmorstein, R., Harrison, S.C. & Wagner, G. Solution structure of the DNA binding domain of Cd2-GAL4 from S, cerevisiae. Nature 356, 450–453 (1992).
Kraulis, P.J., Raine, A.R.C. Gadhavi, P.L. & Laue, E.D. Structure of the DNA-binding domain of zinc GAL4. Nature 356, 448–450 (1992).
Marmorstein, R., Carey, M., Ptashne, M. & Harrison, S.C. DNA recognition by GAL4: structure of a protein-DNA complex. Nature 356, 408–414 (1992).
Marmorstein, R. & Harrison, S.C. Crystal structure of a PPR1-DNA complex: DNA recognition by proteins containing a Zn2Cys6 binuclear cluster. Genes Dev. 8, 2504–2512 (1994).
Johnston, M. A model fungal gene regulatory mechanism: the GALgenes of Saccharomyces Cerevisiae. Microbiol. Rev. 51, 458–476 (1987).
Dickson, R.C. & Riley, M.I. in Yeast genetic engineering (eds Barr, P.J., Brake, A.J. & Valenzuela, P.) 19–40 (Butterworths, Boston; 1989).
Salmeron, J.M.J. & Johnston, S.A. Analysis of the Kluyveromyces lactis positive regulatory gene LAC9 reveals functional homology to, but sequence divergence from, the Saccharomyces cerevisiae GALA gene. Nucl. Acids Res. 14, 7767–7781 (1986).
Halvorsen, Y.-D.C., Nandabalan, K. & Dickson, R.C. LAC9 DNA-binding domain coordinates two zinc atoms per monomer and contacts DNA as a dimer. J. biol. Chem. 265, 13283–13289 (1990).
Pan, T., Halvorsen, Y.-D., Dickson, R.C. & Coleman, J.E. The transcription factor LAC9 from Kluyveromyces lactis-like GAL4 from Saccharomyces cerevisiae forms a Zn(II)2Cys6 binuclear cluster. J. biol. Chem. 265, 21427 (1990).
Gardner, K.H. & Coleman, J.E. 113Cd-1H heteroTOCSY: a method for determining metal-protein connectivities. J. biomol. NMR 4, 761–774 (1994).
Rodgers, K.R. & Coleman, J.E. DNA binding and bending by the transcription factors GAL4(62*) and GAL4(149*). Prot. Sci. 3, 608–619 (1994).
Mau, T., Baleja, J.D. & Wagner, G. Effects of DNA binding and metal substitution on the dynamics of the GAL4 DNA-binding domain as studied by amide proton exchange. Prot. Sci. 1, 1403–1412 (1992).
Shirakawa, M. et al. Assignment of 1H, 15N, 13C resonances, identification of elements of secondary structure and determination of the global fold of the DNA-binding domain of GAL4. Biochemistry 32, 2144–2153 (1993).
Wishart, D.S., Sykes, B.D. & Richards, F.M. The chemical shift index: a fast and simple method for the assignment of protein secondary structure through NMR spectroscopy. Biochemistry 31, 1647–1651 (1992).
Frey, M.H. et al. Polypeptide-metal connectivities in metallothionein 2 by novel 1H-113Cd two-dimensional NMR experiments. J. Am. chem. Soc. 107, 6847–6851 (1985).
Kellogg, G.W. & Schweitzer, B.I. Two- and three-dimensional 31P-driven NMR procedures for complete assignment of backbone resonances in oligodeoxyribonucleotides. J. Biomol. NMR 3, 577–595 (1993).
Otting, G. & Wüthrich, K. Heteronuclear filters in two-dimensional [1H, 1H]-NMR spectroscopy: combined use with isotope labelling for studies of macromolecular conformation and intermolecular interactions. Quart. Rev. Biophys. 23, 39–96 (1990).
Schwabe, J.W.R. & Klug, A. Zinc mining for protein domains. Nature struct. Biol. 1, 345–349 (1994).
Blake, P.R. et al. Quantitative measurement of small through-hydrogen-bond and ‘through-space’ 1H-113Cd and 1H-199Hg J couplings in metal-substituted rubredoxin from Pyrococcus furiousus. J. biomol. NMR 2, 527–533 (1992).
Pérez-Alvarado, G.C. et al. Structure of the carboxy-terminal LIM domain from the cysteine rich protein CRP. Nature struct. Biol. 1, 388–398 (1994).
Brünger, A.T. XPLOR 3.1 Manual (Yale University Press, New Haven, 1992).
Presta, L.G. & Rose, G.D. Helix signals in proteins. Science 240, 1632–1641 (1988).
Zerbe, O., Pountney, D.L., von Philipsborn, W. & Vasák, M. Vicinal 113Cd, 1Hβ-cysteine coupling in Cd-substituted metalloproteins follows a Karplus-type dependence. J. Am. chem. Soc. 116, 377–378 (1994).
Adman, E., Watenpaugh, K.D. & Jensen, L.H. NH⋯S hydrogen bonds in Peptococcus aerogenes ferredoxin, Clostridium pasteurianum rubredoxin, and Chromatium high potential iron protein. Proc. natn. Acad. Sci. U.S.A. 72, 4854–4858 (1975).
Basile, L.A. & Coleman, J.E. Optical activity associated with the sulfur to metal charge transfer bands of Zn and Cd GAL4. Prot. Sci. 1, 617–624 (1992).
Halvorsen, Y.-D.C., Nandabalan, K. & Dickson, R.C. Identification of base and backbone contacts used for DNA sequence recognition and high-affinity binding by LAC9, a transcription activator containing a C6 zinc finger. Molec. cell. Biol. 11, 1777–1784 (1991).
Wüthrich, K. NMR of proteins and nucleic acids (John Wiley & Sons, New York, 1986).
Pan, T. & Coleman, J.E. The DNA binding domain of GAL4 forms a binuclear metal ion complex. Biochemistry 29, 3623–3629 (1990).
Schweitzer, B.I., Gardner, K.H. & Tucker-Kellogg, G. HeteroTOCSY-based experiments for measuring heteronuclear relaxation in nucleic acids and proteins. J. Biomol. NMR in the press.
Marion, D. & Wüthrich, K. Application of phase sensitive two-dimensional correlated spectroscopy (COSY) for measurements of 1H-1H spin-spin coupling constants in proteins. Biochem. biophys. Res. Commun. 113, 967–974 (1983).
Macura, S., Huang, Y., Suter, D. & Ernst, R.R. Two-dimensional chemical exchange and cross-relaxation spectroscopy of coupled nuclear spins. J. magn. Reson 43, 259–281 (1981).
Brown, S.C., Weber, P.L. & Mueller, L. Toward complete 1H NMR spectra in proteins. J. magn. Reson. 77, 166–169 (1988).
Cavanagh, J. & Rance, M. Sensitivity improvement in isotropic mixing (TOCSY) experiments. J. magn. Reson. 88, 72–85 (1990).
Shaka, A.J., Lee, C.J. & Pines, A. Iterative schemes for bilinear operators; applications to spin decoupling. J. magn. Reson. 77, 274–293 (1988).
Derome, A.E. & Williamson, M.P. Rapid-pulsing artifacts in double-quantum-filtered COSY. J. magn. Reson. 88, 177–185 (1990).
Mueller, L. PE-COSY, A Simple Alternative to E-COSY. J. magn. Reson. 72, 191–196 (1987).
Kim, Y. & Prestegard, J.H. Measurement of vicinal couplings from cross peaks in COSY spectra. J. magn. Reson. 84, 9–13 (1989).
Manoleras, N. & Norton, R.S. Spectral processing methods for the removal of t1 noise and solvent artifacts from NMR spectra. j. Biomol. NMR 2, 485–494 (1992).
Nilges, M., Clore, G.M. & Gronenborn, A.M. 1H-NMR stereospecific assignments by conformational data-base searches. Biopolymers 29, 813–822 (1990).
Wüthrich, K., Billeter, M. & Braun, W. Pseudo-structures for the 20 common amino acids for use in studies of protein conformations by measurements of intramolecular proton-proton distance constraints with nuclear magnetic resonance. J. molec. Biol. 169, 949–961 (1983).
Clore, G.M., Gronenborn, A.M., Nilges, M. & Ryan, C.A. Three-dimensional structure of potato carboxypeptidase inhibitor in solution. A study using nuclear magnetic resonance, distance geometry, and restrained molecular dynamics. Biochemistry 26, 8012–8023 (1987).
Kraulis, P.J. MOLSCRIPT: A Program To Produce Both Detailed and Schematic Plots of Protein Structures. J. appl. Crystallogr. 24, 946–950 (1991).
Bacon, D.J. & Anderson, W.F. A Fast Algorithm for Rendering Space-Filling Molecule Pictures. J. molec. Graphics 6, 219–220 (1988).
Merritt, E.A. & Murphy, M.E.P., Raster3D Version 2.0 - A Program for Photorealistic Molecular Graphics. Acta. crystallogr. D50, 869–873 (1994).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Gardner, K., Anderson, S. & Coleman, J. Solution structure of the Kluyveromyces lactis LAC9 Cd2Cys6 DNA-binding domain. Nat Struct Mol Biol 2, 898–905 (1995). https://doi.org/10.1038/nsb1095-898
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb1095-898
This article is cited by
-
Protein and metal cluster structure of the wheat metallothionein domain γ-Ec-1: the second part of the puzzle
JBIC Journal of Biological Inorganic Chemistry (2011)
-
Linkers made to measure
Nature Structural Biology (1997)
-
Structure and mobility of the PUT3 dimer
Natural Structural Biology (1997)
-
Crystal structure of a PUT3–DNA complex reveals a novel mechanism for DMA recognition by a protein containing a Zn2Cys6 binuclear cluster
Nature Structural Biology (1997)
-
Crystal structure of the RAG1 dimerization domain reveals multiple zinc-binding motifs including a novel zinc binuclear cluster
Nature Structural Biology (1997)