Abstract
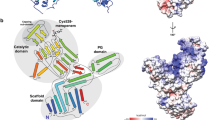
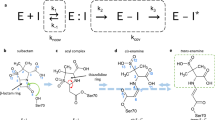
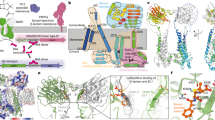
The structure of the 28 kDa β-lactamase inhibitor protein-II (BLIP-II) in complex with the TEM-1 β-lactamase has been determined to 2.3 Å resolution. BLIP-II is a secreted protein produced by the soil bacterium Streptomyces exfoliatus SMF19 and is able to bind and inhibit TEM-1 with subnanomolar affinity. BLIP-II is a seven-bladed β-propeller with a unique blade motif consisting of only three antiparallel β-strands. The overall fold is highly similar to the core structure of the human regulator of chromosome condensation (RCC1). Although BLIP-II does not share the same fold with BLIP, the first β-lactamase inhibitor protein for which structural data was available, a comparison of the two complexes reveals a number of similarities and provides further insights into key components of the TEM-1–BLIP and TEM-1–BLIP-II interfaces. Our preliminary results from gene knock-out studies and scanning electron microscopy also reveal a critical role of BLIP-II in sporulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Doran, J.C., Leskiw, B.K., Aippersbach, S. & Jensen, S.E. J. Bacteriol. 172, 4904–4918 (1990).
Strynadka, N.C.J. et al. Nature 368, 657–660 (1994).
Kim, M.K. & Lee, K.J. App. Env. Microbiol. 60, 1029–1032 (1994).
Massova, I. & Mobashery, S. Antimicrob. Agents Chemother. 42, 1–17 (1998).
Therrien, C. & Levesque, R.C. FEMS Microbiol. Rev. 24, 251–262 (2000).
Blasquez, J. Baquero, M.F., Canton, R., Alos, R. & Bagnew, F. Antimicrob. Agents Chemother. 37, 2059–2063 (1993).
Strynadka, N.C.J., Jensen, S.E., Alzari, P.M. & James, M.N.G. Nature Struct. Biol. 3, 290–297 (1996).
Kang, S.G., Park, H.U., Lee, H.S., Kim, H.T. & Lee, K.J. J. Biol. Chem. 275, 16851–16856 (2000).
Park, H.U. & Lee, K.J. Microbiology 144, 2161–2167 (1998).
Ohtsubo, M. et al. Genes Dev. 1, 585–593 (1987).
Cole, C.N. & Hammell, C.M. Curr. Biol. 8, 368–372 (1998).
Moore, J.D. BioEssays 23, 77–85 (2001).
Strynadka, N.C.J. et al. Nature 359, 700–705 (1992).
Renault, L. et al. Nature 392, 97–101 (1998).
Fülöp, V. & Jones, D.T. Curr. Opin. Struct. Biol. 9, 715–721 (1999).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Sawyer, L. & James, M.N.G. Nature 295, 79–80 (1982).
Dennis, M.S. & Lazarus, R.A. J. Biol. Chem. 269, 22129–22136 (1994).
Petrosino, J., Rudgers, G., Gilbert, H. & Palzkill, T. J. Biol. Chem. 274, 2394–2400 (1999).
Renault, L., Kuhlmann, J., Henkel, A. & Wittinghofer, A. Cell 105, 245–255 (2001).
Zhu, Y.F., Curran, I.H., Joris, B., Ghuysen, J.M., & Lampen, J.O. J. Bacteriol. 172, 1137–1141 (1990).
Zhang, H.Z., Hackbarth, C.J., Chansky, K.M. & Chambers, H.F. Science 291, 1962–1965 (2001).
Otwinowski, Z. In Proceedings of the CCP4 study weekend: data collection and processing (eds Sawyer, L., Issacs, N. & Bailey, S.) 56–62 (Daresbury Laboratory, Warrington; 1993).
Collaborative Computational Project, Number 4. Acta Crystallogr. D 50, 760–763 (1994).
Navaza, J. Acta Crystallogr. A 50, 30–42 (1996).
McRee, D.E. J. Mol. Graphics 10, 44–47 (1992).
Cohen, G.H. J. Appl. Crystallogr. 30, 1160–1161 (1997).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Fonzé, E., Vermeire, M., Nguyen-Distèche, M., Brasseur, R. & Charlier, P. J. Biol. Chem. 274, 21853–21860 (1999).
Acknowledgements
We thank S. Mosimann, Y. Luo and R. Pfuetzner for helpful suggestions and S. He for ESMS analysis. N.C.J.S. is supported by the Canadian Institutes of Health Research (Canada). K.J.L. is grateful to the KISTEP for a research grant from the International Joint R & D Project. N.C.J.S. is a CIHR scholar, a Burroughs Wellcome New Investigator and a HHMI International Scholar. D.L. is a recipient of a CIHR doctoral research award. S.G.K. is a recipient of a post-doctoral research award from the KOSEF.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Lim, D., Park, H., De Castro, L. et al. Crystal structure and kinetic analysis of β-lactamase inhibitor protein-II in complex with TEM-1 β-lactamase. Nat Struct Mol Biol 8, 848–852 (2001). https://doi.org/10.1038/nsb1001-848
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb1001-848
This article is cited by
-
Modeling and fitting protein-protein complexes to predict change of binding energy
Scientific Reports (2016)
-
An interrupted beta-propeller and protein disorder: structural bioinformatics insights into the N-terminus of alsin
Journal of Molecular Modeling (2009)
-
Molecular analysis of the CRINKLY4 gene family in Arabidopsis thaliana
Planta (2005)