Abstract
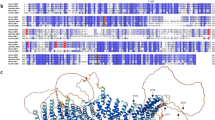
p115RhoGEF, a guanine nucleotide exchange factor for Rho GTPase, is also a GTPase activating protein (GAP) for G12 and G13 heterotrimeric Gα subunits. Near its N-terminus, p115RhoGEF contains a domain (rgRGS) with remote sequence identity to RGS (regulators of G protein signaling) domains. The rgRGS domain is necessary but not sufficient for the GAP activity of p115RhoGEF. The 1.9 Å resolution crystal structure of the rgRGS domain shows structural similarity to RGS domains but possesses a C-terminal extension that folds into a layer of helices that pack against the hydrophobic core of the domain. Mutagenesis experiments show that rgRGS may form interactions with Gα13 that are analogous to those in complexes of RGS proteins with their Gα substrates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Cerione, R.A. & Zheng, Y. Curr. Opin. Cell. Biol. 8, 216–222 (1996).
Lemmon, M.A. & Ferguson, K.M. Curr. Top. Microbiol. Immunol. 228, 39–74 (1998).
Hart, M.J. et al. Science 280, 2112–2114 (1998).
Whitehead, I.P. et al. J. Biol. Chem. 271, 18643–18650 (1996).
Kourlas, P.J. et al. Proc. Natl. Acad. Sci. USA 97, 2145–2150 (2000).
Fukuhara, S., Murga, C., Zohar, M., Igishi, T. & Gutkind, J.S. J. Biol. Chem. 274, 5868–5879 (1999).
Jackson, M. et al. Nature 410, 89–93 (2001).
Kozasa, T. et al. Science 280, 2109–2111 (1998).
Ross, E.M. & Wilkie, T.M. Annu. Rev. Biochem. 69, 795–827 (2000).
Berman, D.M., Kozasa, T. & Gilman, A.G. J. Biol. Chem. 271, 27209–27212 (1996).
Tesmer, J.J.G., Berman, D.M., Gilman, A.G. & Sprang, S.R. Cell 89, 251–261 (1997).
Slep, K.C. et al. Nature 409, 1071–1077 (2001).
Wells, C.D. et al. J. Biol. Chem. in the press (2001).
de Alba, E., De Vries, L., Farquhar, M.G. & Tjandra, N. J. Mol. Biol. 291, 927–939 (1999).
Spink, K.E., Polakis, P. & Weis, W.I. EMBO J. 19, 2270–2279 (2000).
Srinivasa, S.P., Watson, N., Overton, M.C. & Blumer, K.J. J. Biol. Chem. 273, 1529–1533 (1998).
Natochin, M., McEntaffer, R.L. & Artemyev, N.O. J. Biol. Chem. 273, 6731–6735 (1998).
Posner, B.A., Mukhopadhyay, S., Tesmer, J.J., Gilman, A.G. & Ross, E.M. Biochemistry 38, 7773–7779 (1999).
Yang, W., Hendrickson, W.A., Kalman, E.T. & Crouch, R.J. J. Biol. Chem. 265, 13553–13559 (1990).
Otwinowski, Z. In Data collection and processing (eds Sawyer, N.I.L. & Bailey, S.W.) 56–62 (Science and Engineering Council Daresbury Laboratory, Daresbury, U.K.; 1993).
Terwilliger, T.C. & Berendzen, J. Acta Crystallogr. D 55, 849–861 (1999).
Collaborative computational project, N. Acta Crystallogr. D 50, 760–763 (1994).
Jones, T.A. & Kjeldgaard, M. O Version 5.9 (Uppsala University, Uppsala; 1995).
Brunger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thornton, J.M. J. Appl. Crystallogr. 26, 283–291 (1993).
Berman, H.M. et al. Nucleic Acids Res. 28, 235–242 (2000).
Guex, N., Diemand, A. & Peitsch, M.C. Trends Biochem. Sci. 24, 364–367 (1999).
Van Gunsteren, W.F. http://igc.ethz.ch/gromos (1996).
Singer, W.D., Miller, R.T. & Sternweis, P.C. J. Biol. Chem. 269, 19796–19802 (1994).
Wells, C.W., Jiang, X. & Sternweis, P.C. Methods Enzymol. 345, in the press (2001).
Esser, L. http://www.hhmi.swmed.edu/external/Doc/glr/ (2000).
Esnouf, R. J. Mol. Graph. 15, 133–138 (1997).
The POV-ray Team. http://www.povray.org (1998).
Thompson, J.D., Higgins, D.G. & Gibson, T.J. Nucleic Acids Res. 22, 4673–4680 (1994).
Acknowledgements
We thank A. Loew and T. Xiao for their assistance with data collection and MAD data analysis in addition to their guidance in structure refinement; S. Raghunathan, X. Du and K. Ihara for useful discussions; S. Gutowski for excellent technical assistance with GAP assays; and the staff of MacCHESS and ALS. This work is supported by NIH grants to S.R.S, P.C.S and C.D.W, and Welch Foundation grants to S.R.S and P.C.S.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Chen, Z., Wells, C., Sternweis, P. et al. Structure of the rgRGS domain of p115RhoGEF. Nat Struct Mol Biol 8, 805–809 (2001). https://doi.org/10.1038/nsb0901-805
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0901-805
This article is cited by
-
Interaction kinetics between p115-RhoGEF and Gα13 are determined by unique molecular interactions affecting agonist sensitivity
Communications Biology (2022)
-
Mechanistic insight into GPCR-mediated activation of the microtubule-associated RhoA exchange factor GEF-H1
Nature Communications (2014)
-
p115RhoGEF
AfCS-Nature Molecule Pages (2011)
-
On the mechanism of autoinhibition of the RhoA-specific nucleotide exchange factor PDZRhoGEF
BMC Structural Biology (2009)
-
The R7 RGS Protein Family: Multi-Subunit Regulators of Neuronal G Protein Signaling
Cell Biochemistry and Biophysics (2009)