Abstract
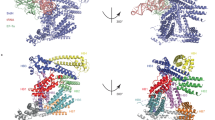
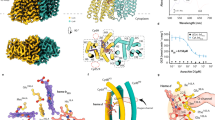
The human pathogen Legionella pneumophila, the etiological agent of the severe and often fatal Legionnaires' disease, produces a major virulence factor, termed 'macrophage infectivity potentiator protein' (Mip), that is necessary for optimal multiplication of the bacteria within human alveolar macrophages. Mip exhibits a peptidyl prolyl cis-trans isomerase (PPIase) activity, which appears to be important for infection. Here we report the 2.4 Å crystal structure of the Mip protein from L. pneumophila Philadelphia 1 and the 3.2 Å crystal structure of its complex with the drug FK506. Each monomer of the homodimeric protein consists of an N-terminal dimerization module, a long (65 Å) connecting α-helix and a C-terminal PPIase domain exhibiting similarity to human FK506-binding protein. In view of the recent significant increase in the number of reported cases of Legionnaires' disease and other intracellular infections, these structural results are of prime interest for the design of new drugs directed against Mip proteins of intracellular pathogens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Schmid, F.X. Annu. Rev. Biophys. Biomol. Struct. 22, 123–142 (1993).
Galat, A. & Metcalfe, S.M. Prog. Biophys. Mol. Biol. 63, 67–118 (1995).
Kallen, J. et al. Nature 353, 276–279 (1991).
Ranganathan, R., Lu, K.P., Hunter, T. & Noel, J.P. Cell 89, 875–886 (1997).
Wilson, K.P. et al. Acta Crystallogr. D 51, 511–521 (1995).
Fischer, G., Bang, H., Ludwig, B., Mann, K. & Hacker, J. Mol. Microbiol. 6, 1375–1383 (1992).
Engleberg, N.C., Carter, C., Weber, D.R., Cianciotto, N.P. & Eisenstein, B.I. Infect. Immun. 57, 1263–1270 (1989).
Cianciotto, N.P. & Fields, B.S. Proc. Natl. Acad. Sci. USA. 89, 5188–5191 (1992).
Hentschel, U., Steinert, M. & Hacker, J. Trends Microbiol. 8, 226–231 (2000).
Hendrickson, W.A. Science 254, 51–58 (1991).
Michnick, S.W., Rosen, M.K., Wandless, T.J., Karplus, M. & Schreiber, S.L. Science 252, 836–839 (1991).
Van Duyne, G.D., Standaert, R.F., Karplus, P.A., Schreiber, S.L. & Clardy, J. Science 252, 839–842 (1991).
Craescu, C.T. et al. Biochemistry 35, 11045–11052 (1996).
Schmidt, B. et al. FEBS Lett. 352, 185–190 (1994).
Schmidt, B. et al. FEBS Lett. 372, 169–172 (1995).
Kuo, C.C. et al. J. Infect. Dis. 167, 841–849 (1993).
Fischer, G., Bang, H. & Mech, C. Biomed. Biochim. Acta 43, 1101–1111 (1984).
Janowski, B., Wollner, S., Schutkowski, M. & Fischer, G. Anal. Biochem. 252, 299–307 (1997).
Riboldi-Tunnicliffe, A. & Hilgenfeld, R. J. Appl. Crystallogr. 32, 1003–1005 (1999).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computing Project, Number 4. Acta Crystallogr. D 50, 760–763 (1994).
Sheldrick, G.M., Dauter, Z., Wilson, K.S., Hope, H. & Sieker, L.C. Acta Crystallogr. D 49, 18–23 (1993).
de La Fortelle, E. & Bricogne, G. Methods Enzymol. 276, 472–494 (1997).
Abrahams, J.P. & Leslie, A.G.W. Acta Crystallogr. D 52, 30–42 (1996).
Brünger, A.T. Nature 355, 472–475 (1992).
Brünger, A.T. et al. Acta Crystallogr. D 54, 905–921 (1998).
Jones, T.A., Zou, J.-Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A 47, 110 (1991).
Read, R.J. Acta Crystallogr. A 42, 140–149 (1986).
Kissinger, C.R., Gehlhaar, D.K. & Fogel, D.B. Acta Crystallogr. D 55, 484–491 (1999).
Esnouf, R.M. J. Mol. Graph. 15, 132–134 (1997).
Merritt, A.E. & Bacon, D.J. Methods Enzymol. 277, 505–524 (1997).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Weiss, M.S. & Hilgenfeld, R. J. Appl. Crystallogr. 30, 203–205 (1997).
Acknowledgements
We thank A. Savoia and the staff of XRD beamline 5.2 at Elettra (Sincrotrone Trieste), Trieste, Italy, for help with data collection. FK506 was a gift from Fujisawa Pharmaceutical Co. This work was supported in part by the Deutsche Forschungsgemeinschaft. R.H., G.F. and J.H. thank the Fonds der Chemischen Industrie. This work is dedicated to the memory of L. Fonda and P.M. Fasella.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Riboldi-Tunnicliffe, A., König, B., Jessen, S. et al. Crystal structure of Mip, a prolylisomerase from Legionella pneumophila. Nat Struct Mol Biol 8, 779–783 (2001). https://doi.org/10.1038/nsb0901-779
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0901-779
This article is cited by
-
Characterization of two putative Dichelobacter nodosus footrot vaccine antigens identifies the first lysozyme inhibitor in the genus
Scientific Reports (2019)
-
Structural basis of interaction between dimeric cyclophilin 1 and Myb1 transcription factor in Trichomonas vaginalis
Scientific Reports (2018)
-
In Silico Analysis of Conformational Changes Induced by Normal and Mutation of Macrophage Infectivity Potentiator Catalytic Residues and its Interactions with Rapamycin
Interdisciplinary Sciences: Computational Life Sciences (2015)
-
Backbone and side-chain 1H, 13C and 15N assignments of the PPIase domain of macrophage infectivity potentiator (Mip) protein from Coxiella burnetii
Biomolecular NMR Assignments (2014)
-
Solution structure of the Legionella pneumophila Mip-rapamycin complex
BMC Structural Biology (2008)