Abstract
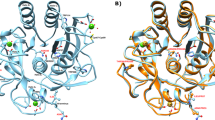
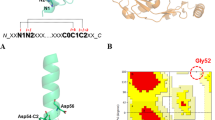
Small proteins generally fold cooperatively: disruption of significant parts of the folded structure leads to unfolding of the rest of the protein. We show here, using proline scanning mutagenesis, that the native-like tertiary fold of the α-lactalbumin (α-LA) molten globule is formed by the non-cooperative assembly of its constituent helices. In contrast to the drastic destabilizing effects of proline substitutions in cooperatively folded proteins, proline mutations in the molten globule appear to cause only individual helices to unfold, without significantly influencing the other helices or the overall topology. Thus, the key determinants of a protein's overall fold may not be of the all-or-none type.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kuwajima, K. The molten globule state as a clue for understanding the folding and cooperativity of globular-protein structure. Proteins 6, 87–103 (1989).
Ptitsyn, O.B. The molten globule state. in Protein Folding (ed. Creighton, T.E.) 243–300 (W. H. Freeman and Co., New York, 1992).
Dobson, C.M. Solid evidence for molten globules. Curr. Biol. 4, 636–40 (1994).
Eliezer, D., Jennings, P.A., Wright, P.E., Doniach, S., Hodgson, K.O. & Tsuruta, H. The radius of gyration of an apomyoglobin folding intermediate. Science 270, 487–8 (1995).
Peng, Z.-y. & Kim, P.S. A protein dissection study of a molten globule. Biochemistry 33, 2136–41 (1994).
Wu, L.C., Peng, Z.-y. & Kim, P.S. Bipartite structure of the α-lactalbumin molten globule. Nature Struct. Biol. 2, 281–86 (1995).
Peng, Z.-y., Wu, L.C. & Kim, P.S. Local structural preferences in the α-lactalbumin molten globule. Biochemistry 34, 3248–52 (1995).
Polverino de Laureto, P., De Filippis, V., Di Bello, M., Zambonin, M. & Fontana, A. Probing the molten globule state of alpha-lactalbumin by limited proteolysis. Biochemistry 34, 12596–604 (1995).
Wilson, G. et al. Vibrational Raman optical activity of alpha-lactalbumin: comparison with lysozyme, and evidence for native tertiary folds in molten globule states. J. Mol. Biol. 254, 747–60 (1995).
Alexandrescu, A.T., Evans, P.A., Pitkeathly, M., Baum, J. & Dobson, C.M. Structure and dynamics of the acid-denatured molten globule state of alpha-lactalbumin: a two-dimensional NMR study. Biochemistry 32, 1707–1718 (1993).
Cunningham, B.C. & Wells, J.A. High-resolution epitope mapping of hGH-receptor interactions by alanine-scanning mutagenesis. Science 244, 1081–1085 (1989).
Richardson, J.S. & Richardson, D.C. Amino acid preferences for specific locations at the ends of alpha helices. Science 240, 1648–52 (1988).
Strehlow, K.G., Robertson, A.D. & Baldwin, R.L. Proline for alanine substitutions in the C-peptide helix of ribonuclease A. Biochemistry 30, 5810–4 (1991).
Minor, D., Jr. & Kim, P.S. Measurement of the beta-sheet-forming propensities of amino acids. Nature 367, 660–663 (1994).
Smith, C.K., Withka, J.M. & Regan, L. Athermodynamicscaleforthe beta-sheet forming tendencies of the amino acids. Biochemistry 33, 5510–5517 (1994).
Minor, D., Jr. & Kim, P.S. Context is a major determinant of beta-sheet propensity. Nature 371, 264–267 (1994).
Wood, S.J., Wetzel, R., Martin, J.D. & Hurle, M.R. Prolines and amyloidogenicity in fragments of the Alzheimer's peptide beta/A4. Biochemistry 34, 724–30 (1995).
O'Neil, K.T. & De Grado, W.F. A thermodynamic scale for the helix-forming tendencies of the commonly occurring amino acids. Science 250, 646–651 (1990).
Horovitz, A., Matthews, J.M. & Fersht, A.R. Alpha-helix stability in proteins. II. Factors that influence stability at an internal position. J. Mol. Biol. 227, 560–568 (1992).
Sauer, U.H., San, D.P. & Matthews, B.W. Tolerance of T4 lysozyme to proline substitutions within the long interdomain alpha-helix illustrates the adaptability of proteins to potentially destabilizing lesions. J. Biol. Chem. 267, 2393–2399 (1992).
Blaber, M. et al. Determination of alpha-helix propensity within the context of a folded protein. Sites 44 and 131 in bacteriophage T4 lysozyme. J. Mol. Biol. 235, 600–624 (1994).
Baum, J., Dobson, C.M., Evans, P.A. & Hanley, C. Characterization of a partly folded protein by NMR methods: studies on the molten globule state of guinea pig alpha-lactalbumin. Biochemistry 28, 7–13 (1989).
Chyan, C.L., Wormald, C., Dobson, C.M., Evans, P.A. & Baum, J. Structure and stability of the molten globule state of guinea-pig alpha-lactalbumin: a hydrogen exchange study. Biochemistry 32, 5681–5691 (1993).
Schulman, B.A., Peng, Z.-y., Redfield, C., Dobson, C.M. & Kim, P.S. Different subdomains are most protected from hydrogen exchange in the molten globule and native states of α-lactalbumin. J. Mol. Biol. 253, 651–657 (1995).
Nozaka, M., Kuwajima, K., Nitta, K. & Sugai, S. Detection and characterization of the intermediate on the folding pathway of human alpha-lactalbumin. Biochemistry 17, 3753–3758 (1978).
Acharya, K.R., Ren, J.S., Stuart, D.I., Phillips, D.C. & Fenna, R.E. Crystal structure of human alpha-lactalbumin at 1.7 Å resolution. J. Mol. Biol. 221, 571–581 (1991).
Siddiqui, A.S. & Barton, G.J. Continuous and discontinuous domains: an algorithm for the automated generation of reliable protein domain definitions. Prot Sci. 4, 872–884 (1995).
Xie, D., Bhakuni, V. & Freire, E. Calorimetric determination of the energetics of the molten globule intermediate in protein folding: apo-α-lactalbumin. Biochemistry 30, 10673–10678 (1991).
Yutani, K., Ogasahara, K. & Kuwajima, K. Absence of the thermal transition in apo-alpha-lactalbumin in the molten globule state. A study by differential scanning microcalorimetry. J. Mol. Biol. 228, 347–350 (1992).
Jennings, P.A. & Wright, P.E. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science 262, 892–8926 (1993).
Kreimer, D.I., Szosenfogel, R., Goldfarb, D., Silman, I. & Weiner, L. Two-state transition between molten globule and unfolded states of acetylcholinesterase as monitored by electron paramagnetic resonance spectroscopy. Proc. Natl. Acad. Sci. USA 91, 12145–12149 (1994).
Chan, H.S., Bromberg, S. & Dill, K.A. Models of cooperativity in protein folding. Philos. Trans. R. Soc. Lond. B Biol. Sci. 348, 61–70 (1995).
Ptitsyn, O.B., Bychkova, V.E. & Uversky, V.N. Kinetic and equilibrium folding intermediates. Philos. Trans. R. Soc. Lond. BB iol. Sci. 348, 35–41 (1995).
Kiefhaber, T. & Baldwin, R.L. Intrinsic stability of individual alpha helices modulates structure and stability of the apomyoglobin molten globule form. J. Mol. Biol. 252, 122–132 (1995).
Loh, S.N., Kay, M.S. & Baldwin, R.L. Structure and stability of a second molten globule intermediate in the apomyoglobin folding pathway. Proc. Natl. Acad. Sci. USA 92, 5446–5450 (1995).
Marmorino, J.L. & Pielak, G.J. A native tertiary interaction stabilizes the A state of cytochromec. Biochemistry 34, 3140–3143 (1995).
Shimizu, A., Ikeguchi, M. & Sugai, S. Unfolding of the molten globule state of alpha-lactalbumin studied by 1H NMR. Biochemistry 32, 13198–203 (1993).
Dill, K.A. Dominant forces in protein folding. Biochemistry 29, 7133–55 (1990).
Kay, M.S. & Baldwin, R.L. Packing interactions in the apomyoglobin folding intermediate. Nature Struct. Biol. 3, 439–45 (1996).
Bai, Y., Sosnick, T.R., Mayne, L. & Englander, S.W. Protein folding intermediates: native-state hydrogen exchange. Science 269, 192–197 (1995).
Dill, K.A. & Shortle, D. Denatured states of proteins. Annu. Rev. Biochem. 60, 795–825 (1991).
Dobson, C.M. Unfolded proteins, compact states and molten globules. Curr. Opin. Struct. Biol. 2, 6–12 (1992).
Shortle, D. Denatured states of proteins and their roles in folding and stability. Curr. Opin. Struct. Biol. 3, 66–74 (1993).
Lumb, K.J. & Kim, P.S. Formation of a hydrophobic cluster in denatured bovine pancreatic trypsin inhibitor. J. Mol. Biol. 236, 412–420 (1994).
Agashe, V.R., Shastry, M.C.R. & Udgaonkar, J.B. Initial hydrophobic collapse in the folding of barstar. Nature 377, 754–757 (1995).
Oliveberg, M. & Fersht, A. Thermodynamics of transient conformations in the folding pathway of barnase: reorganization of the folding intermediate at low pH. Biochemistry 35, 2738–2749 (1996).
Hughson, F.M., Barrick, D. & Baldwin, R.L. Probing the stability of a partly folded apomyoglobin intermediate by site-directed mutagenesis. Biochemistry 30, 4113–4118 (1991).
Minor, D.L., Jr. & Kim, P.S. Context-dependent secondary structure formation of a designed protein sequence. Nature 380, 730–734 (1996).
Lau, K.F. & Dill, K.A. Theory for protein mutability and biogenesis. Proc. Natl. Acad. Sci. USA 87, 638–642 (1990).
Behe, M.J., Lattman, E.E. & Rose, G.D. The protein-folding problem: the native fold determines packing, but does packing determine the native fold? Proc. Natl. Acad. Sci. USA 88, 4195–4199 (1991).
Bowie, J.U., Luthy, R. & Eisenberg, D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 253, 164–170 (1991).
Xiong, H., Buckwalter, B.L., Shieh, H.M. & Hecht, M.H. Periodicity of polar and nonpolar amino acids is the major determinant of secondary structure in self-assembling oligomeric peptides. Proc. Natl. Acad. Sci. USA 92, 6349–6353 (1995).
Ptitsyn, O.B. How does protein synthesis give rise to the 3D-structure? FEBS Letts 285, 176–181 (1991).
Edelhoch, H. Spectroscopic determination of tryptophan and tyrosine in proteins. Biochemistry 6, 1948–1954 (1967).
Laue, T.M., Shah, B.D., Ridgeway, T.M. & Pelletier, S.L. Computer-aided interpretation of analytical sedimentation data for proteins. in Analytical Ultracentrifugaton in Biochemistry and Polymer Science (eds S.E. Harding, A.J. Rowe & J.C. Horton) 90–125 (The Royal Society of Chemistry, Cambridge, 1992).
Chen, Y.H., Yang, J.T. & Chau, K.H. Determination of the helix and beta form of proteins in aqueous solution by circular dichroism. Biochemistry 13, 3350–3359 (1974).
Ellman, G.L. Tissue sulphydryl groups. Arch. Biochem. Biophys. 82, 70–77 (1959).
Priestle, J.P. RIBBON: A stereo cartoon drawing program for proteins. J. Appl. Crystallogr. 21, 572–576 (1988).
Woody, R.W. Circular Dichroism of Peptides. in The Peptides: Analysis, Synthesis, Structure (ed. V.J. Hruby) 15–114 (Academic Press, Orlando, 1985).
Manning, M.C. & Woody, R.W. Theoretical study of the contribution of aromatic side chains to the circular dichroism of basic bovine pancreatic trypsin inhibitor. Biochemistry 28, 8609–8613 (1989).
Chakrabartty, A., Kortemme, T., Padmanabhan, S. & Baldwin, R.L. Aromatic side-chain contribution to far-ultraviolet circular dichroism of helical peptides and its effect on measurement of helix propensities. Biochemistry 32, 5560–5565 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Schulman, B., Kim, P. Proline scanning mutagenesis of a molten globule reveals non-cooperative formation of a protein's overall topology. Nat Struct Mol Biol 3, 682–687 (1996). https://doi.org/10.1038/nsb0896-682
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0896-682
This article is cited by
-
Using chirality to probe the conformational dynamics and assembly of intrinsically disordered amyloid proteins
Scientific Reports (2017)
-
The region from phenylalanine-28 to lysine-50 of a yeast mitochondrial ATPase inhibitor (IF1) forms an α-helix in solution
Journal of Bioenergetics and Biomembranes (2015)
-
Transient α-helices in the disordered RPEL motifs of the serum response factor coactivator MKL1
Scientific Reports (2014)
-
Cooperativity of folding of the apomyoglobin pH 4 intermediate studied by glycine and proline mutations
Nature Structural & Molecular Biology (1997)
-
A residue-specific NMR view of the non-cooperative unfolding of a molten globule
Natural Structural Biology (1997)