Abstract
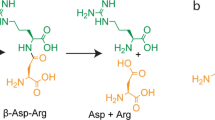
Almost half of the entire set of predicted genomic products from Methanococcus jannaschii are classified as functionally unknown hypothetical proteins. We present a structure-based identification of the biochemical function of a protein with an as yet unknown function from a M. jannaschii gene, Mj0226. The crystal structure of Mj0226 protein determined at 2.2 Å resolution reveals that the protein is a homodimer and each monomer folds into an elongated α/β structure of a new fold family. Comparisons of Mj0226 protein with protein structures in the database, however, indicate that one part of the protein is homologous to some of the nucleotide-binding proteins. Biochemical analysis shows that Mj0226 protein is a novel nucleotide triphosphatase that can efficiently hydrolyze nonstandard nucleotides such as XTP to XMP or ITP to IMP, but not the standard nucleotides, in the presence of Mg2+ or Mn2+ ions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kerlavage, A.R. TIGR Microbial genome database. http://www.tigr.org/tdb/mdb/mdb.html (1999).
Rost, B. Marrying structure and genomics. Structure 6, 259 –263 (1998).
Gaasterland, T. http://www.mcs.anl.gov/home/gaasterl/genomes.html ( 1999).
Altschul, S.F. et al, Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Kerlavage, A.R. http://www.tigr.org/softlab (1999).
Deckert, G. et al. The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature 392, 353– 358 (1998).
Klenk, H.P. et al. The complete genome sequence of the hyperthermophilic, sulfite reducing archeon Archaeoglobus fulgidus. Nature 390, 364–370 (1997).
Fraser, C.M. et al. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science 270, 397– 403 (1995).
Fleischmann, R.D. et al. Whole genome random sequencing and assembly of Haemophilus influenzae Rd. Science 269, 496– 512 (1995).
Tomb, J.F. et al. The complete genome sequence of thegastric pathogen Helicobacter pylori. Nature 388, 539– 547 (1997).
Bult, C.J. et al. Complete genome sequence of the methanogenic archeon, Methanococcus janaschii. Science 273, 1058–1073 (1996).
Cole, S.T. et al. Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393. 537–544 (1998).
Kim, S.-H. Shining a light on structural genomics. Nature Struct. Biol. 5, 643–645 (1998).
Shapiro, L. & Lima, C.D. The Argonne structural genomics workshop: Lamaze class for the birth of new science. Structure 6, 265–267 (1998).
Sali, A. 100,000 protein structures for the biologist. Nature Struct. Biol. 5, 1029–1032 ( 1998).
Montelione, G.T. & Anderson, S. Structural genomics: keystone for a human proteome project. Nature Struct. Biol. 6, 11–12 (1999).
Murzin, A., Brenner, S.E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995).
Orengo, C.A., Jones, D.T. & Thornton, J.M. CATH––a hierarchic classification of protein domain structures. Nature 372, 631– 634 (1994).
Gibrat, J.F., Madej, T. & Bryant, S.H. Surprising similarities in structure comparison. Curr. Opin. Struct. Biol. 6, 377–385 (1996).
Dalal, S., Balasubramanian, S., & Regan, L. Protein alchemy: changing beta-sheet into alpha-helix. Nature Struct. Biol. 4, 548– 552 (1997).
Zarembinsky, T.I. et al. Structure-based assignment of the biochemical function of a hypothetical protein: a test case of structural genomics. Proc. Natl. Acad. Sci. USA 95, 15189–15193 (1998).
Sander, C. & Schneider, R. Data base of homology derived protein structures and the structural meaning of sequence alignment. Proteins Struct. Funct. Genet. 9, 56– 68 (1991).
Aberg, A., Yaremchuk, A., Tukalo, M., Rasmussen, B. & Cusack, S. Crystal structure analysis of the activation of histidine by Thermus thermophilus histidyl t-RNA synthetase. Biochemistry, 36, 3084– 3090 (1997).
Logan, D.T., Mazauric, M.-H, Kern, D. & Moras, D. Crystal structure of glycyl-tRNA synthetase from T. thermophilus. EMBO J. 14, 4156–4167 (1995).
Usher, K.C. et al. Crystal structures of CheY from Thermatoga maritima do not support conventional explanations for the structural basis of enhanced stability. Protein Sci. 2, 403– 412 (1998).
Teplyakov, A. et al. Crystal structure of bacteriophage T4 deoxynucleotide kinase with its substrates dGMP and ATP. EMBO J. 15, 3487–3491 (1996).
Lee, J.O., Rieu, P., Arnaout, M.A. & Liddington, R. Crystal structure of the A domain from the alpha subunit of integrin CR3 (CD11b/ CD18). Cell 80, 631–638 ( 1995).
Louie, G.V. et al. Structure of porphobilinogen deaminase reveals a flexible multidomain polymerase with a single catalytic core. Nature 359, 33–39 (1992).
Chen, P. et al. Crystal structure of glycinamide ribonucleotide transformylase from Escherichia coli at 3.0 Å resolution. A target enzyme for chemotherapy. J. Mol. Biol. 227, 283– 292 (1992).
Friedman, A.M., Fischmann, T.O. & Steitz, T.A. Crystal structure of lac repressor core tetramer and its implications for DNA looping. Science 268, 1721–1727 (1995).
Baikalov, I. et al. Structure of the Escherichia coli response regulator NarL. Biochemistry 35, 11053– 11061 (1996).
Rao, F. et al. Structure of the oxidized long chain flavodoxin from Anabena 7120 at 2 Å resolution. Protein Sci. 1, 1413–1427 (1992).
Huang, M.E., Manus, S., Chuat, J.C. & Galibert, N. Analysis of 62 kb DNA sequence of chromosome X reveals 36 open reading frames and a gene cluster with a counterpart on chromosome XI. Yeast 12, 869–875 (1996).
Noskov, V.N. et al. HAM1, a gene controlling 6-N-hydroxylaminopurine sensitivity and mutagenesis in the yeast Saccharomyces cerevisiae. Yeast. 12. 17–29 ( 1996).
Pavlov, Y. I. et al .Base analog N6-hydroxyaminopurine mutagensis in Escherichia coli: genetic control and molecular specificity. Mutat. Res . 253, 33–46 ( 1991).
Kozmin, S. G. et al. Multiple antimutagenesis mechanisms affect mutagenic activity and specificity of the base analogue 6-N-hydroxylaminopurine in bacteria and yeast. Mutat. Res. 402, 41– 50 (1998).
Klinker, J.F. & Seifert, R. Functionally nonequivalent interactions of guanosine 5´-triphosphate, inosine 5´-triphosphate, and xanthine 5´-triphosphate with the retinol G-protein, transducin, and with Gi-proteins in HL-60 leukemia cell membranes. Biochem. Pharmacol 54, 551 –562 (1997).
Jancarik, J. & Kim, S.-H. Sparse matrix sampling: a screening method for crystallization of proteins. J. Appl. Crystallogr. 24, 409–411 (1991).
Collaborative Computational Project Number 4. The CCP4 suite: program for protein crystallography. Acta Crystallogr. D 50, 760–763 ( 1994).
Sack, J.S. CHAIN: a crystallographic modelling program. J. Mol. Graph. 6, 224–225 (1988).
Brünger, A.T. X-PLOR, a system for crystallography and NMR, Version 3.1. (Yale University Press, New Haven, Connecticut; 1992).
Laskowski, R.A., Macarther, M.W., Moss, P.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283– 291 (1993).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insight from the interfacial and thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281–296 (1991).
Acknowledgements
We are grateful to C.A. Caperelli (University of Cincinnati) for providing glycinamide ribonucleotide and 10-formyl-5,8-dideazafolate, and helpful comments on the GAR assay. We also thank B.K. Lee and J.S. Kim for the program SHEBA. This work was supported by funds from the Korea Institute of Science and Technology (KIST 2000 program), Korean Ministry of Science and Technology (Biotech 2000 program), Korean Ministry of Health and Welfare, Office of Biological and Environmental Research, Office of Energy Research, US Department of Energy, and Korean Academy of Science and Technology (a Young Scientist Award to Y.C.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hwang, K., Chung, J., Kim, SH. et al. Structure-based identification of a novel NTPase from Methanococcus jannaschii. Nat Struct Mol Biol 6, 691–696 (1999). https://doi.org/10.1038/10745
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/10745
This article is cited by
-
Inosine triphosphate pyrophosphatase from Trypanosoma brucei cleanses cytosolic pools from deaminated nucleotides
Scientific Reports (2022)
-
A new method to investigate the catalytic mechanism of YhdE pyrophosphatase by using a pyrophosphate fluorescence probe
Scientific Reports (2017)
-
A disease spectrum for ITPA variation: advances in biochemical and clinical research
Journal of Biomedical Science (2016)
-
Crystal structure of the MazG-related nucleoside triphosphate pyrophosphohydrolase from Thermotoga maritima MSB8
Journal of Structural and Functional Genomics (2015)
-
A nuclear magnetic resonance based approach to accurate functional annotation of putative enzymes in the methanogen Methanosarcina acetivorans
BMC Genomics (2011)