Abstract
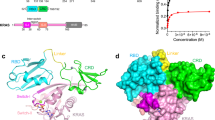
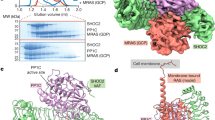
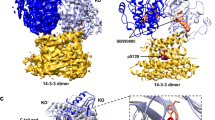
The Ras protein signals to a number of distinct pathways by interacting with diverse downstream effectors. Among the effectors of Ras are the Raf kinase and RalGDS, a guanine nucleotide dissociation stimulator specific for Ral. Despite the absence of significant sequence similarities, both effectors bind directly to Ras, but with different specificities. We report here the 2.1 Å crystal structure of the complex between Ras and the Ras-interacting domain (RID) of RalGDS. This structure reveals that the β-sheet of the RID joins the switch I region of Ras to form an extended β-sheet with a topology similar to that found in the Rap–Raf complex. However, the side chain interactions at the joining junctions of the two interacting systems and the relative orientation of the two binding domains are distinctly different. Furthermore, in the case of the Ras–RID complex a second RID molecule also interacts with a different part of the Ras molecule, the switch II region. These findings account for the cross-talk between the Ras and Ral pathways and the specificity with which Ras distinguishes the two effectors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Hall, A. Curr. Opin. Cell Biol. 5, 265–268 (1993).
Milburn, M.V. et al. Science 247, 939–945 (1990).
Vojtek, A.B., Hollenberg, S.M. & Cooper, J.A. Cell 74, 205–214 (1993).
Nassar, N. et al. Nature 375, 554–560 (1995).
Brtva, T.R. et al. J. Biol. Chem. 270, 9809–9812 (1995).
Hu, Q., Klippel, A., Muslin, A.J., Fantl, W.J. & Williams, L.T. Science 268, 100–102 (1995).
Hofer, F., Fields, S., Schneider, C. & Martin, G.S. Proc. Watl. Acad. Sci. USA 91, 11089–11093 (1994).
Spaargaren, M. & Bischoff, J.R. Proc. Natl. Acad. Sci. USA 91, 12609–12613 (1994).
Kikuchi, A., Demo, S.D., Ye, Z.H., Chen, Y.W. & Williams, L.T. Mol. Cell. Biol. 14, 7483–7491 (1994).
Ponting, C.P. & Benjamin, D.R. Trends Biochem. Sci. 21, 422–425 (1996).
Herrmann, C., Horn, G., Spaargaren, M. & Wittenghofer, A. J. Biol. Chem. 271, 6794–6800 (1996).
Huang, L., Weng, X., Hofer, F., Martin, G.S. & Kim, S.-H. Nature Struct. Biol. 4, 609–615 (1997).
Geyer, M., Herrmann, C., Wohlgemuth, S., Wittinghofer, A. & Kalbitzer, H.R. Nature Struct. Biol. 4, 694–699 (1997).
White, M.A. et al. Cell 80, 533–541 (1995).
White, M.A., Vale, T., Camonis, J.H., Schaefer, E. & Wigler, M.H. J. Biol. Chem. 271, 16439–16442 (1996).
Khosravi-Far, R. et al. Mol. Cell. Biol. 16, 3923–3933 (1996).
Joneson, T., White, M., Wigler, M.H. & Bar-Sagi, D. Science 271, 810–812 (1996).
Okazaki, M. et al. Oncogene 14, 515–521 (1997).
Katz, M.E. & McCormick, F. Curr. Opin. Genet. Dev. 7, 75–79 (1997).
Feig, L.A., Urano, T. & Cantor, S. Trends Biochem. Sci 21, 438–441 (1996).
Urano, T., Emkey, R. & Feig, L.A. EMBO. J. 15, 810–816 (1996).
Hwang, M.C., Sung, Y.J. & Hwang, Y.W. J. Biol. Chem. 271, 8196–8202 (1996).
Nassar, N. et al. Nature Struct. Biol. 3, 723–729 (1996).
Miura, K. et al. Jap. J. Cancer Res. 77, 45–51 (1986).
Pai, E.F. et al. EMBO J. 9, 2351–2359 (1990).
Huang, L., Lancarik, J., Kim, S.-H., Hofer, F. & Martin, G.S. Acta Crystallogr. D 52, 1033–1035 (1996).
Brunger, A.T. X-PLOR, version 3.85 (Yale University, New Haven, Connecticut; 1996).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta Crystallogr. A47, 110–119 (1991).
Kraulis, P.J. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M.E.P. Acta Crystallogr D. 50, 869–873 (1994).
Kabsch, W. & Sander, C. Biopolymers 22, 2577–2637 (1983).
Nicholls, A., Sharp, K.A. & Honig, B. Proteins Struct Funct Genet. 11, 281–296 (1991).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huang, L., Hofer, F., Martin, G. et al. Structural basis for the interaction of Ras with RaIGDS. Nat Struct Mol Biol 5, 422–426 (1998). https://doi.org/10.1038/nsb0698-422
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0698-422
This article is cited by
-
The energetic and allosteric landscape for KRAS inhibition
Nature (2024)
-
Molecular insight into isoform specific inhibition of PI3K-α and PKC-η with dietary agents through an ensemble pharmacophore and docking studies
Scientific Reports (2021)
-
Engineering subtilisin proteases that specifically degrade active RAS
Communications Biology (2021)
-
Functional characterisation of a novel class of in-frame insertion variants of KRAS and HRAS
Scientific Reports (2019)
-
The Structural Basis of Oncogenic Mutations G12, G13 and Q61 in Small GTPase K-Ras4B
Scientific Reports (2016)