Abstract
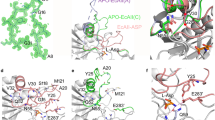
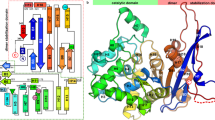
The structure of L-aspartate-α-decarboxylase from E. coli has been determined at 2.2 Å resolution. The enzyme is a tetramer with pseudofour-fold rotational symmetry. The subunits are six-stranded β-barrels capped by small α-helices at each end. The active sites are located between adjacent subunits. The electron density provides evidence for catalytic pyruvoyl groups at three active sites and an ester at the fourth. The ester is an intermediate in the autocatalytic self-processing leading to formation of the pyruvoyl group. This unprecedented structure provides novel insights into the general phenomenon of protein processing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Chong, S. et al Protein splicing involving the Saccharomyces cerevisiae VMA1 intein J. Biol. Chem. 271, 22159–22168 (1996).
Shao, Y. & Kent, S.B.H. Protein splicing: occurrence, mechanisms and related phenomena. Chem. Biol. 4, 187–194 (1997).
van Poelje, P.D. & Snell, E.E. Pyruvoyl-dependent enzymes. Annu. Rev. Biochem. 59, 29–59 (1990).
Shao, Y., Xu, M.-Q. & Paulus, H. Protein splicing: evidence for an N-O acyl rearrangement as the initial step in the splicing process. Biochemistry 35, 3810–3815 (1996).
Xu, M.-Q. & Perler, F.B. The mechanism of protein splicing and its modulation by mutation. EMBOJ. 15, 5146–5153 (1996).
Recsei, P.A., Huynh, Q.K. & Snell, E.E. Conversion of prohistidine decarboxylase to histidine-decarboxylase peptide-chain cleavage by non-hydrolytic serinolysis. Proc. Natl. Acad. Sci. USA 80, 973–977 (1983).
Cronan, J.E. Beta-Alanine synthesis in Escherichia coli. J. Bacteriol. 141, 1291–1297 (1980).
Ramjee, M.K., Genschel, U., Abell, C. & Smith, A.G. Escherichia coli L-aspartate-α-decarboxylase: preprotein processing and observation of reaction intermediates by electrospray mass spectrometry. Biochem. J. 323, 661–669 (1997).
Gallagher, T., Snell, E.E. & Hackert, M.L. Pyruvoyl-dependent histidine-decarboxylase - active-site structure and mechanistic analysis. J. Biol. Chem. 264, 12737–12743 (1989).
Williamson, J.M. & Brown, G.M. J. Biol. Chem. 254, 8074–8082 (1979).
Song, L.Z. et al. Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 274, 1859–1866 (1996).
Sutcliffe, M.J., Haneef, I., Carney, D. & Blundell, T.L. Knowledge-based modeling of homologous proteins. 1. 3-dimensional frameworks derived from the simultaneous superposition of multiple structures. Protein Engng. 1, 377–384 (1987).
Hodel, A., Kim, S.H. & Brunger, A.T. Model bias in macromolecular crystal structures. Acta Crystallogr. A. 48, 851–859 (1992).
Read, R.J. Improved Fourier coefficients for maps using phases from partial structures with errors. Acta Crystallogr. A42, pp. 140–149 (1986).
Iwai, K. & Ando, T. N-O acyl shifts. Meth. Enz. 11, 262–282 (1987).
Donate, L.E., Rufino, S.D., Canard, L.H.J. & Blundell, T.L. Conformational analysis and clustering of short and medium size loops connecting regular secondary structures: A database for modeling and prediction. Protein Sci. 5, 2600–2616 (1996).
Rufino, S.D., Donate, L.E., Canard, L.H.J & Blundell T.L Predicting the conformational class of short and medium size loops connecting regular secondary structures: Application to comparative modelling. J. Mol. Biol. 267, 352–367 (1997).
Vanopdenbosch, N., Cramer, R. & Giarrusso, F.F. SYBYL, The integrated molecular modeling system. J. Mol. Graph. 3, 110–111 (1985).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Meth Enz. 276, pp.307–326 (1997).
Sheldrick, G.M. The SHELXS system. In Crystallographic computing 5. (Podjarny, A.D. & Thierry, J.C., eds) 145–157 (IUCR, Oxford University Press, Oxford, UK; 1991).
Bailey, S. The CCP4 suite - programs for protein crystallography. Acta Crystallogr. D50, 760–763 (1994).
de la Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Meth. Enz. 276, 472–494 (1997).
Jones, T.A., Zou, J.-Y, Cowan, S.W. & Kjelgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. Crystallographic refinement by simulated annealing application to a 2.8-Å resolution structure of aspartate-aminotransferase. J. Mol. Biol. 203, 803–816 (1988)
Kleywegt, G.J. Dictionaries for Heteros. ESF/CCP4 Newsletter 31, 45–50 (1995).
Alien, F.H. et. al. The development of Version 3 and Version 4 of the Cambridge Structural Database System. J. Chem. Info. Comp. Sci. 31, 187–204 (1991).
Laskowski, R.A., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK - A program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Kraulis, P.J. MOLSCRIPT - A program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E.A. & Murphy, M. Version 2.0, a program for photorealistic molecular graphics. Acta Crystallogr. D50, 869–873 (1994).
Nicholls, A., Bharadwaj, R. & Honig, B. GRASP - graphical representation and analysis of surface-properties. Biophys. J. 64, A 166 (1993).
Luzzati, V. Traitement statistique des erreurs dans la determination des structures cristallines. Acta Crystallogr. 5, 802–810 (1952).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Albert, A., Dhanaraj, V., Genschel, U. et al. Crystal structure of aspartate decarboxylase at 2.2 Å resolution provides evidence for an ester in protein self–processing. Nat Struct Mol Biol 5, 289–293 (1998). https://doi.org/10.1038/nsb0498-289
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0498-289
This article is cited by
-
A High-Specific-Activity L-aspartate-α-Decarboxylase from Bacillus aryabhattai Gel-09 and Site-Directed Mutation to Improve Its Substrate Tolerance
Applied Biochemistry and Biotechnology (2023)
-
Research progress of l-aspartate-α-decarboxylase and its isoenzyme in the β-alanine synthesis
World Journal of Microbiology and Biotechnology (2023)
-
Structural insights into phosphatidylethanolamine formation in bacterial membrane biogenesis
Scientific Reports (2021)
-
Identification of mutations restricting autocatalytic activation of bacterial l-aspartate α-decarboxylase
Amino Acids (2018)
-
Significance of Arg3, Arg54, and Tyr58 of l-aspartate α-decarboxylase from Corynebacterium glutamicum in the process of self-cleavage
Biotechnology Letters (2014)