Abstract
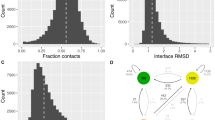
The fundamental event in biological assembly is association of two biological macromolecules. Here we present a successful, accurate ab initio prediction of the binding of uncomplexed lysozyme to the HyHel5 antibody. The prediction combines pseudo Brownian Monte Carlo minimization with a biased–probability global side–chain placement procedure. It was effected in an all–atom representation, with ECEPP/2 potentials complemented with the surface energy, side–chain entropy and electrostatic polarization free energy. The near–native solution found was surprisingly close to the crystallographic structure (root–mean–square deviation of 1.57 Å for all backbone atoms of lysozyme) and had a considerably lower energy (by 20 kcal mol−1) than any other solution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wodak, S.J. & Janin, J. Computer analysis of protein-protein interactions. J. molec. Biol. 124, 323–342 (1978).
Kuntz, I.D., Blaney, J.M., Oatley, S.J., Langridge, R. & Ferrin, T.E. A geometric approach to macromolecule-ligand interactions. J. molec. Biol. 161, 269–288 (1982).
Connolly, M.L. Shape complementarity at the hemoglobin α1\β1 Subunit interface. Biopolymers 25, 1229–1247 (1986).
Warwicker, J. Investigating protein-protein interaction surfaces using a reduced stereochemical and electrostatic model. J. molec. Biol. 206, 381–395 (1989).
Goodsell, A.S. ; Olson, A.J. Automated docking of substrates to proteins by simulated annealing. Proteins Struct. Funct. Genet. 8, 195–202 (1990).
Cherfils, J., Duquerroy, S. & Janin, J. Protein-protein recognition analyzed by docking simulation. Proteins 11, 271–280 (1991).
Jiang, F. & Kim, S.-H. “Soft docking”: matching of molecular surface cubes. J. molec. Biol. 219, 79–102 (1991).
Shoichet, B.K. & Kuntz, I.D. Protein Docking and Complementarity. J. molec. Biol. 221, 327–346 (1991).
Shoichet, B.K., Bodian, D.L. & Kuntz, I.D. Molecular docking using shape descriptors. J. comp. Chem. 13, 380–397 (1992).
Bacon, D.J. & Moult, J. Docking by least-squares fitting of molecular surface patterns. J. molec. Biol. 225, 849–858 (1992).
Walls, P.H. & Sternberg, M.J.E. New algorithm to model protein-protein recognition based on surface complementarity. J. molec. Biol. 228, 277–297 (1992).
Pellegrini, M. & Doniah, S. Computer simulation of antibody binding specificity. Proteins 15, 436–444 (1993).
Sheriff, S. et al. Three-dimensional structure of an antibody-antigen complex. Proc. natn. Acad. Sci. U.S.A. 84, 8075– (1987).
Padlan, E.A. & Kabat, E.A. Modeling of antibody complex. Meth. Enzym. 203, 3–21 (1991).
Rini, J.M., Schulze-Gahmen, U., Wilson, I.A. Structural evidence for induced fit as a mechanism for antibody-antigen recognition. Science 255, 959–965 (1992).
Padlan, E.A., Silverton, E.W., Sheriff, S., Cohen, G.H., Smith-Gill, S.J., Davies, D.R. Structure of an antibody-antigen complex: crystal structure of the HyHEL-10Fab-lysozyme complex. Proc. natn. Acad. Sci. U.S.A. 86, 5938–5942 (1989).
Fischmann, T.O. et al. Crystallographic refinement of the three-dimensional structure of theFabD1.3-lysozyme complex at 2.5 A resolution. J. biol. Chem. 266, 12915–12920 (1991).
Diamond, R. Refinement of the Structure of Hen Egg-white Lysozyme. J. molec. Biol. 82, 371–391 (1974).
Abagyan, R.A., Totrov, M.M. & Kuznetsov, D.A. ICM: a new method for structure modelling and design:Applications to docking and structure prediction from the distorted native conformation. J. comp. Chem. (in the press).
Abagyan, R.A. & Totrov, M.M. Biased probability monte carlo conformational searches and electrostatic calculations for peptides and proteins. J. molec. Biol. 235, 983–1002 (1994).
Totrov, M.M. & Abagyan, R.A. Efficient parallelization of the energy, surface and derivative calculations for internal coordinate mechanics. J. comp. Chem. (in the press).
Bernstein, F.C. et al. The protein data bank: a computer-based archival file for macromolecularstructures. J. molec. Biol. 112, 535–542 (1977).
Momany, F.A., McGuire, R.F., Burgess, A.W. & Scheraga, H.A. Energy parameters in polypeptides VII. Geometric parameters, partial atomic charges, nonbonded interactions, hydrogen bond interactions, and intrinsic torsional potentials for the naturally occurring amino acids. J. phys. Chem. 79, 2361–2381 (1975).
Nemethy, G., Pottle, M.S. & Scheraga, H.A. Energy parameters in polypeptides. 9. updating of geometric parameters, nonbonded interactions and hydrogen bond interactions for the naturally occurring amino acids. J. phys. Chem. 87, 1883–1887 (1983).
Pickersgill, R.W. A rapid method of calculating charge-charge interaction energies in proteins. Protein Eng. 2, 247–248 (1988).
Abagyan, R.A. & Argos, P. Optimal protocol and trajectory visualization for conformational searches of peptides and proteins. J. molec. Biol. 225, 519–532 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Totrov, M., Abagyan, R. Detailed ab initio prediction of lysozyme–antibody complex with 1.6 Å accuracy. Nat Struct Mol Biol 1, 259–263 (1994). https://doi.org/10.1038/nsb0494-259
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0494-259