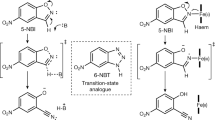
Abstract
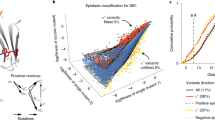
A random library of single amino acid mutants of myoglobin was generated using a highly efficient, single–base–misincorporation random mutagenesis method to discover new ligand–binding pathways in myoglobin. A surprisingly large fraction of the library exhibits ligand–binding kinetics that are substantially different from the wild–type protein. In addition to residues 45, 64 and 68, which comprise the best studied ligand–binding pathway single mutations of several other clusters of residues far away from that pathway are discovered which profoundly affect the ligand–binding kinetics. These results provide a new approach to explore the relationship between the fluctuations in protein structure and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kendrew, J.C. et al. Structure of myoglobin. A three-dimensional fourier synthesis at 2 Å resolution Nature 185, 422–427 (1960).
Perutz, M.F. & Mathews, F.S. An x-ray study of azide methaemoglobin. J. molec. Biol. 21, 199–202 (1966).
Nobbs, C.L. in Heme and Hemoproteins (eds Chance, B., Estabrook, R. W. & Yonetani, T.) 143–147 (Academic Press, New York, 1966).
Takano, T. Structure of myoglobin refined at 2.0 Å resolution. I. Crystallographic refinement of metmyoglobin from sperm whale J. molec. Biol. 110, 537–568 (1977).
Takano, T. Structure of myoglobin refined at 2.0 Å resolution. II. Structure of deoxymyoglobin from sperm whale J. molec. Biol. 110, 569–584(1977).
Phillips, S.E.V. Structure and refinement of oxymyoglobin at 1. 6 Å resolution J. molec. Biol. 142, 531–554 (1980)
Kuriyan, J., Wilz, S., Karplus, M. & Petsko, G.A. X-ray structure and refinement of carbon-monoxy (Fell)-myoglobin at 1.5 Å resolution J. molec. Biol. 192, 133–154 (1986).
Bolognesi, M. et al. Reactivity of ferric aplysia and sperm whale myoglobins towards imidazole X-ray and binding study. J. molec. Biol. 158, 305–315 (1982).
Ringe, D., Petsko, G.A., Kerr, D.E. & Ortiz de Montellano, P.R. Reaction of myoglobin with phenylhydrazine: A molecular doorstop. Biochemistry 23, 2–4 (1984).
Johnson, K.A., Olson, J.S. & Phillips, G.N., Jr. Structure of myoglobin-ethyl isocyanide Histidine as a swinging door for ligand entry. J. molec. Biol. 207, 459–463 (1989).
Frauenfelder, H., Petsko, G.A. & Tsernoglou, D. Temperature-dependent x-ray diffraction as a probe of protein structural dynamics Nature 280, 558–563 (1979).
Case, D.A. & Karplus, M. Dynamics of ligand binding to heme proteins J. molec. Biol. 132, 343–368 (1979).
Tilton, R.F., Jr et al. Computational studies of the interaction of myoglobin and xenon J. molec. Biol. 192, 443–456 (1986).
Kottalam, J. & Case, D.A. Dynamics of ligand escape from the heme pocket of myoglobin J. Am. chem. Soc. 110, 7690–7697 (1988).
Tilton, R.F., Jr, Singh, U.C., Kuntz, I.D., Jr & Kollman, P.A. Protein-ligand dynamics. A 96 picosecond simulation of a myoglobin-xenon complex J. molec. Biol. 199, 195–211 (1988).
Lambright, D.G., Balasubramanian, S. & Boxer, S.G. Ligand and proton exchange dynamics in recombinant human myoglobin mutants J. molec. Biol. 207, 289–299 (1989).
Balasubramanian, S., Lambright, D.G., Marden, M.C. & Boxer, S.G. CO recombination to human myoglobin mutants in glycerol-water solutions Biochemistry 32, 2202–2212 (1993).
Lambright, D.G., Balasubramanian, S. & Boxer, S.G. Dynamics of protein relaxation in site-specific mutants of human myoglobin Biochemistry 32, 10116–10124 (1993).
Rohlfs, R.J. et al. The effects of amino acid substitution at position E7 (residue 64) on the kinetics of ligand binding to sperm whale myoglobin J. biol. Chem. 265, 3168–3176 (1990).
Egeberg, K.D. et al. The role of Val68(E11) in ligand binding to sperm whale myoglobin Site-directed mutagenesis of a synthetic gene. J. biol. Chem. 265, 11788–11795 (1990).
Carver, T.E. et al. Analysis of the kinetics barriers for the ligand binding to sperm whale myoglobin using site-directed mutagenesis and laser photolysis techniques J. biol. Chem. 265, 20007–20020 (1990).
Smerdon, S.J. et al. Distal pocket polarity in ligand binding to myoglobin: structural and functional characterization of a threonine (E11) mutant Biochemistry 30, 6252–6260(1991).
Olson, J.S. et al. The role of the distal histidine in myoglobin and haemoglobin Nature 336, 265–266 (1988).
Rizzi, M. et al. Crystal structure of a distal site double mutant of sperm whale myoglobin at 1.6 Å resolution FEBS Letters 320, 13–16 (1993).
Elber, R. & Karplus, M. Enhanced sampling in molecular dynamics: use of the time-dependent Hartree approximation for a simulation of carbon monoxide diffusion through myoglogin J. Am. chem. Soc. 112, 9161–9175(1990).
Chatfield, M.D., Walda, K.N. & Magde, D. Activation parameters for ligand escape from myoglobin proteins at room temperature J. Am. chem. Soc. 112, 4680–4687 (1990).
Shortle, D. & Lin, B. Genetic analysis of staphylococcal nuclease: identification of three intragenic “global” suppressors of nuclease-minus mutations Genetics 110, 539–555 (1985).
Oliphant, A.R. & Struhl, K. An efficient method for generating proteins with altered enzymatic properties: application to β-lactamase Proc. natn. Acad. Sci. U.S.A. 86, 9094–9098 (1989).
Rennell, D., Bouvier, S.E., Hardy, L. & Poteete, A.R. Systematic mutation of bacteriophage T4 lysozyme J. molec. Biol. 222, 67–87 (1991).
Springer, B.A. & Sligar, S.G. High-level expression of sperm whale myoglobin in Escherichia Coli. Proc. natn. Acad. Sci. U.S.A. 84, 8961–8965(1987).
Lambright, D.G., Balasubramanian, S. & Boxer, S.G. Protein relaxation dynamics in human myoglogin Chem. Phys. 158, 249–260 (1991).
Tilton, R.F., Jr., Kuntz, I.D., Jr. & Petsko, G.A. Cavities in proteins: structure of a metmyoglobin-xenon complex solved to 1.9 Å. Biochemistry 23, 2849–2857 (1984).
Dickerson, R.E. & Geis, I. in Hemoglobin (The Benjamin/Cummings Publishing Company, Inc., Menlo Park, California, U.S.A., 1983).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Acta crystallogr. A47, 110–119 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Huang, X., Boxer, S. Discovery of new ligand binding pathways in myoglobin by random mutagenesis. Nat Struct Mol Biol 1, 226–229 (1994). https://doi.org/10.1038/nsb0494-226
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0494-226
This article is cited by
-
Protein tertiary structure and the myoglobin phase diagram
Scientific Reports (2019)
-
Myoglobin ligand gate mechanism analysis by a novel 3D visualization technique
Journal of Mathematical Chemistry (2019)
-
Molecular evolution of myoglobin in the Tibetan Plateau endemic schizothoracine fish (Cyprinidae, Teleostei) and tissue-specific expression changes under hypoxia
Fish Physiology and Biochemistry (2018)
-
Carbon monoxide binding properties of domain-swapped dimeric myoglobin
JBIC Journal of Biological Inorganic Chemistry (2015)
-
Dynamic void distribution in myoglobin and five mutants
Scientific Reports (2014)