Abstract
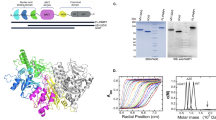
The status of the poly(A) tail at the 3′-end of mRNAs controls the expression of numerous genes in response to developmental and extracellular signals. Poly(A) tail regulation requires cooperative binding of two human U1A proteins to an RNA regulatory region called the polyadenylation inhibition element (PIE). When bound to PIE RNA, U1A proteins also bind to the enzyme responsible for formation of the mature 3′-end of most eukaryotic mRNAs, poly(A) polymerase (PAP). The NMR structure of the 38 kDa complex formed between two U1A molecules and PIE RNA shows that binding cooperativity depends on helix C located at the end of the RNA-binding domain and just adjacent to the PAP-interacting domain of U1A. Since helix C undergoes a conformational change upon RNA binding, the structure shows that binding cooperativity and interactions with PAP occur only when U1A is bound to its cognate RNA. This mechanism ensures that the activity of PAP enzyme, which is essential to the cell, is only down regulated when U1A is bound to the U1A mRNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Wickens, M., Anderson, P. & Jackson, R.J. Life and death in the cytoplasm: Messages from the 3′ end. Curr. Op. Genet. Develop. 7, 220–232 (1997).
Boelens, W.C. et al. The human U1 snRNP-specific U1A protein inhibits polyadenylation of its own pre-mRNA. Cell 72, 881– 892 (1993).
van Gelder, C.W.G. et al. A complex secondary structure in U1A pre-mRNA that binds two molecules of U1A protein is required for regulation of polyadenylation. EMBO J. 12, 5191–5200 ( 1993).
Gunderson, S.I. et al. The Human U1A snRNP Protein Regulates Polyadenylation via a Direct Interaction with Poly(A) Polymerase. Cell 76, 531–541 (1994).
Gunderson, S.I., Vagner, S., Polycarpou-Schwarz, M. & Mattaj, I.W. Involvement of the carboxy terminus of vertebrate poly A polymerase in U1A autoregulation and in the coupling of splicing and polyadenylation. Genes & Dev. 11, 761–773 (1997).
Barabino, S.M.L. & Keller, W. Last but not least: regulated poly(A) tail formation. Cell 99, 9–11 (1999).
Varani, G. & Nagai, K. RNA Recognition by RNP proteins during RNA processing and maturation. Ann. Rev. Biophys. Biomol. Struct. 27, 407–445 ( 1998).
Allain, F.-H.T. et al. Specificity of ribonucleoprotein interaction determined by RNA folding during complex formation. Nature 380, 646–650 (1996).
Allain, F.H.-T., Howe, P.W.A., Neuhaus, D. & Varani, G. Structural Basis of the RNA binding specificity of human U1A protein. EMBO J. 16, 5764–5774 ( 1997).
Folkers, P.J.M., Folmer, R.H.A., Konings, R.N.H. & Hilbers, C.W. Overcoming the ambiguity problem encountered in the analysis of nuclear Overhauser magnetic resonance spectra of symmetric dimer proteins. J. Am. Chem. Soc. 115, 3798–3799 (1993).
Mittermaier, A., Varani, L., Muhandiram, D.R., Kay, L.E. & Varani, G. Changes in sidechain and backbone dynamics identify determinants of specificity in RNA recognition by human U1A protein. J. Mol. Biol. 294, 967– 979 (1999).
Pervushin, K., Riek, R., Wider, G. & Wüthrich, K. Attenuation of T2 relaxation by mutual cancellation by dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl. Acad. Sci. USA 94, 12366–12371 (1997).
Zwhalen, C. et al. Methods for Measurement of intermolecular NOEs by multinuclear NMR spectroscopy: application to a bacteriophage λ N-peptide/boxb RNA complex. J. Am. Chem. Soc. 119, 6711– 6721 (1997).
Gillespie, J.R. & Shortle, D. Characterization of long-range structure in the denatured state of Staphyloccocal nuclease. I. Paramagnetic relaxation enhancement by nitroxide spin labels. J. Mol. Biol. 268, 158–169 (1997).
Ramos, A. & Varani, G. A new method to detect long-range protein–RNA contacts: NMR detection of electron-proton relaxation induced by nitroxide spin-labeled RNA. J. Am. Chem. Soc. 120 , 10992–10993 (1998).
Howe, P.W.A., Allain, F.H.-T., Varani, G. & Neuhaus, D. Determination of the NMR structure of the complex between U1A protein and its RNA polyadenylation inhibition element. J. Biomol. NMR 11, 59–84 (1998).
Allain, F.H.-T. & Varani, G. Structure of the P1 helix from group I self splicing introns. J. Mol. Biol. 250, 333–353 (1995).
Bayer, P., Varani, L. & Varani, G. Refinement of the structure of protein–RNA complexes by residual dipolar coupling analysis. J. Biomol. NMR 14, 149–155 (1999).
Jovine, L., Oubridge, C., Avis, J.M. & Nagai, K. Two structurally different RNA molecules are bound by the spliceosomal protein U1A using the same recognition strategy. Structure 4, 621–631 (1996).
Wolberger, C. Multiprotein–DNA complexes in transcriptional regulation. Ann. Rev. Biophys. Biomol. Struct. 28, 29– 56 (1999).
Li, T., Stark, M.R., Johnson, A.D. & Wolberger, C. Crystal structure of the MATa1/MATα2 homeodomain heterodimer bound to DNA. Science 270, 262–269 (1995).
Phillips, C.L., Stark, M.R., Johnson, A.D. & Dahlquist, F.W. Heterodimerization of the yeast homeodomain transcriptional regulator α2 and a1 induces an interfacial helix in α2. Biochemistry 33, 9294–9302 (1994).
Grainger, R.J., Norman, D.G. & Lilley, D.M.J. Conformational consequences of binding of U1A protein to the 3′ untranslated region of its pre-mRNA. J. Mol. Biol. 288, 585–594 ( 1999).
Stark, M.R. & Johnson, A.D. Interaction between two homeodomain proteins is specified by a short C-terminal tail. Nature 371, 429–432 (1994).
Klein Gunnewiek, J.M.T. et al. 14 Residues of the U1 snRNP-specific U1A protein is involved in homodimerization, cooperative RNA binding and inhibition of polyadenylation . Mol. Cell Biol. in the press (2000 ).
Biamonti, G. & Riva, S. New insights into the auxiliary domains of eukaryotic RNA binding proteins. FEBS Lett. 340, 1–8 (1994).
Birney, E., Kumar, S. & Krainer, A.R. Analysis of the RNA-recognition motif and RS and RGG domains: conservation in metazoan pre-mRNA splicing factors. Nucleic Acids Res. 21, 5803–5816 (1993).
Price, S.R., Oubridge, C., Varani, G. & Nagai, K. Preparation of RNA–protein complexes for X-ray crystallography and NMR. In RNA–Protein Interaction: Practical Approach (ed. Smith, C.) (Oxford University Press, 1998), p. 48–72.
Mori, S., Abeygunawardana, C., O'Neil Johnson, M. & van Zijl, P.C.M. Improved sensitivity of HSQC spectra of exchanging protons at short interscan delays using a new fast HSQC (FHSQC) detection scheme that avoids water saturation . J. Mag. Res. B 108, 94– 98 (1995).
Bax, A., Ikura, M., Kay, L.E., Torchia, D.A. & Tschudin, R. Comparison of different modes of two-dimensional reverse-correlation NMR for the study of proteins. J. Mag. Res. 86, 304–318 (1990).
Talluri, S. & Wagner, G. An optimized 3D NOESY-HSQC. J. Mag. Res. B 112, 200–205 (1996).
Gillespie, J.R. & Shortle, D. Characterization of long-range structure in the denatured state of Staphyloccocal nuclease. II. Distance restraints from paramagnetic relaxation and calculation of an ensemble of structures. J. Mol. Biol. 268, 170–184 (1997).
Ramos, A. et al. RNA Recognition by a Staufen double-stranded RNA binding domain . EMBO J. in the press (2000).
Fletcher, C.M., Jones, D.N.M., Diamond, R. & Neuhaus, D. Treatment of NOE constraints involving equivalent or nonstereoassigned protons in calculations of biomacromolecular structures. J. Biomol. NMR 8, 292–310 ( 1996).
Varani, G., Aboul-ela, F. & Allain, F.H.-T. NMR investigations of RNA structure. Prog. NMR Spectr. 29, 51–127 (1996).
Acknowledgements
We thank F. Allain and C. Gubser for their help at the initial stages of the project; M. Kelly (IMP Berlin) and K. Gardner (University of Toronto) for suggestions on the preparation of deuterated protein. L.V. acknowledges the support of an EU studentship and the MRC. We would also like to thank the Florence Large Scale facility for Biomolecular NMR, funded in part by the EU, for access to the 800 Mhz spectrometer.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Varani, L., Gunderson, S., Mattaj, I. et al. The NMR structure of the 38 kDa U1A protein – PIE RNA complex reveals the basis of cooperativity in regulation of polyadenylation by human U1A protein . Nat Struct Mol Biol 7, 329–335 (2000). https://doi.org/10.1038/74101
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/74101
This article is cited by
-
An annual cycle of gene regulation in the red-legged salamander mental gland: from hypertrophy to expression of rapidly evolving pheromones
BMC Developmental Biology (2019)
-
Interactions between RNA-binding proteins and P32 homologues in trypanosomes and human cells
Current Genetics (2016)
-
Cooperative structure of the heterotrimeric pre-mRNA retention and splicing complex
Nature Structural & Molecular Biology (2014)
-
Modular protein-RNA interactions regulating mRNA metabolism: a role for NMR
European Biophysics Journal (2011)
-
Structural insights into RNA recognition by the alternative-splicing regulator muscleblind-like MBNL1
Nature Structural & Molecular Biology (2008)