Abstract
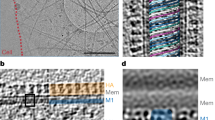
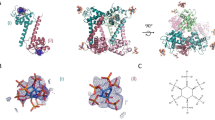
Matrix protein (M1) of influenza virus is a bifunctional protein that mediates the encapsidation of RNA-nucleoprotein cores into the membrane envelope. It is therefore required that M1 binds both membrane and RNA simultaneously. The X-ray crystal structure of the N-terminal portion of type A influenza virus M1—amino acid residues 2–158—has been determined at 2.08 Å resolution at pH 4.0. The protein forms a dimer. A highly positively charged region on the dimer surface is suitably positioned to bind RNA while the hydrophobic surface opposite the RNA binding region may be involved in interactions with the membrane. The membrane-binding hydrophobic surface could be buried or exposed after a conformational change.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Krug, R.M. The influenza viruses 89–152 (Plenum Press, New York).
Pons, M.W., Schulze, I.T., Hirst, G.K. & Hauser, R. Solvation and characterization of the ribonucleoprotein of influenza virus. Virology 39, 250–259 (1969).
Helenius, A. Unpacking the incoming influenza virus. Cell 69, 577–578 (1992).
Lamb, R.A. & Horvath, C.M. Diversity of coding strategies in influenza viruses. Trends Genet. 7(8), 261–266 (1991).
Fujiyoshi, Y., Kume, N.P., Sakata, K. & Sato, S.B. Fine structure of influenza A virus observed by electron cryo-microscopy. EMBO J. 13, 318–326 (1994).
Gregoriades, A. & Frangione, B. Insertion of influenza M protein into the viral lipid bilayer and localization of site of insertion. J. Virol. 40, 323–328 (1981).
Bucher, D.J. et al. Incorporation of influenza virus M-protein into liposomes. J. Virol. 36, 586–590.
Wakefield, L. & Brownlee, G.G. RNA-binding properties of influenza A virus matrix protein Ml. Nucleic Acids Res. 17, 8569–8580 (1989).
Winter, G. & Fields, S. The structure of the gene encoding the nucleoprotein of human influenza virus A/PR/8/34. Virology 114, 423–428 (1981).
Martin, K. & Helenius, A. Nuclear transport of influenza virus ribonucleoproteins: the viral matrix protein (M1) promotes export and inhibits import. Cell 67, 117–130 (1991).
llto, T., Gorman, O.T., Kawaoka, Y., Bean, W.J. & Webster, R.G. Evolutionary analysis of the influenza A virus M gene with comparison of the M1 and M2 proteins. J. Virology 65, 5491–5498 (1991).
Zhirnov, O.P. Isolation of matrix protein M1 from influenza viruses by acid-dependent extraction with nonionic detergent. Virology 186, 324–330 (1992).
Sha, B.D. & Luo, M. Crystallization and preliminary X-ray crystallographic studies of type A influenza virus matrix protein M1. (in the press).
Holm, L. & Sander, C. Searching protein structure databases has come of age. Proteins 19, 165–173 (1995).
Ye, Z., Pal, R., Fox, W. & Wagner, R.R. Functional and antigenic domains of the matrix (M1) protein of influenza A virus. J. Virol. 61, 239–246 (1987).
Ye, Z., Baylor, N.W. & Wagner, R.R. Transcription-inhibition and RNA-binding domains of influenza A virus matrix protein mapped with anti-idiotypic antibodies and synthetic J. Virol. 63, 3586–3594 (1989).
Watanabe, K., Handa, H., Mizumoto, K. & Nagata, K. Mechanism for inhibition of influenza virus RNA polymerase activity by matrix protein. J. Virol. 70, 241–247 (1996).
Nicholls, A. GRASP: Graphical Representation and Analysis Surface Properties (Columbia University, New York, 1992).
Picot, D., Loll, P.J. & Garavito, R.M. The X-ray crystal structure of the membrane protein prostaglandin H2 synthase-1. Nature, 367, 243–249 (1994).
Otwinowski, Z. Proceedings of the CCP4 Study Weekend: Data Collections and Processing, (compiled by Sawyer, L., Isaacs, N. & Bailey, S.) 56–62 (Daresbury Laboratory, UK 1993).
Minor, W. XdisplayF Program. Purdue University, West Lafayette, USA (1993).
McRee, D.E. A visual protein crystallographic software system for X11/XView. J. Mol. Graphics 10, 44–47 (1992).
CCP4 The CCP4 Suite: Programs for Protein Crystallography Acta Crystallogr. D50, 760–763 (1994).
Kleywegt, G.J. & Jones, T.A. From First map Final Model(CCP4) (ed. S. Bailey, R. Hubbard and D. Waller) 59–66 (1994).
Jones, T.A., Zhou, J.Y., Cowan, S.W. & Kjeldgard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A47, 110–119 (1991).
Brunger, A.T. X-PLOR, Version 3.1. A System for X-Ray Crystallography & NMR (Yale University Press, New Haven, 1992).
Carson, M. Ribbons 2.0 J. Appl. Crystallogr. 24, 958–961 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sha, B., Luo, M. Structure of a bifunctional membrane-RNA binding protein, influenza virus matrix protein M1. Nat Struct Mol Biol 4, 239–244 (1997). https://doi.org/10.1038/nsb0397-239
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0397-239
This article is cited by
-
Peptide Models of the Cytoplasmic Tail of Influenza A/H1N1 Virus Hemagglutinin Expand Understanding its pH-Dependent Modes of Interaction with Matrix Protein M1
The Protein Journal (2023)
-
The native structure of the assembled matrix protein 1 of influenza A virus
Nature (2020)
-
Structure of African swine fever virus p15 reveals its dual role for membrane-association and DNA binding
Protein & Cell (2020)
-
Crystal structure of the African swine fever virus structural protein p35 reveals its role for core shell assembly
Protein & Cell (2020)
-
CypA Regulates AIP4-Mediated M1 Ubiquitination of Influenza A Virus
Virologica Sinica (2018)