Abstract
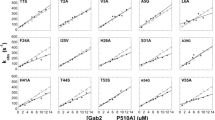
A critical challenge of structural genomics is to extract functional information from protein structures. We present an example of how this may be accomplished using the Evolutionary Trace (ET) method in the context of the regulators of G protein signaling (RGS) family. We have previously applied ET to the RGS family and identified a novel, evolutionarily privileged site on the RGS domain as important for regulating RGS activity. Here we confirm through targeted mutagenesis of RGS7 that these ET-identified residues are critical for RGS domain regulation and are likely to function as global determinants of RGS function. We also discuss how the recent structure of the complex of RGS9, Gt/i1α–GDP–AlF4− and the effector subunit PDEγ confirms their contact with the effector–G protein interface, forming a structural pathway that communicates from the effector-contacting surface of the G protein and RGS catalytic core domain to the catalytic interface between Gα and RGS. These results demonstrate the effectiveness of ET for identifying binding sites and efficiently focusing mutational studies on their key residues, thereby linking raw sequence and structure data to functional information.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cowan, C.W., He, W. & Wensel, T.G. Prog. Nucl. Acid Res. Mol. Biol. 65, 341–359 (2000).
Benzing, T. et al. J. Biol. Chem. 275, 28167–28172 (2000).
Zheng, B., Chen, D. & Farquhar, M. Proc. Natl. Acad. Sci. USA 97, 4040–4045 (2000).
Kovoor, A. et al. J. Biol. Chem. 275, 3397–3402 (2000).
Posner, B.A., Gilman, A.G. & Harris, B.A. J. Biol. Chem. 274, 31087–31093 (1999).
McEntaffer, R.L., Natochin, M. & Artemyev, N.O. Biochemistry 38, 4931–4937 (1999).
Nekrasova, E.R. et al. Biochemistry 36, 7638–7643 (1997).
Natochin, M., Granovsky, A.E. & Artemyev, N.O. J. Biol. Chem. 272, 17444–17449 (1997).
Popov, S., Krishna, U., Falck, J. & Wilkie, T. J. Biol. Chem. 27, 18962–18968 (2000).
Womack, K. et al. J. Neurosci. 20, 2792–2799 (2000).
Tu, Y., Popov, S., Slaughter, C. & Ross, E. J. Biol. Chem. 274, 38260–38267 (1999).
Chen, C., Seow, K.T., Guo, K., Yaw, L.P. & Lin, S.C. J. Biol. Chem. 274, 19799–197806 (1999).
Druey, K.M. et al. J. Biol. Chem. 274, 18836–18842 (1999).
Lichtarge, O., Bourne, H.R. & Cohen, F.E. J. Mol. Biol. 257, 342–358 (1996).
Sowa, M.E., He, W., Wensel, T.G. & Lichtarge, O. Proc. Natl. Acad. Sci. USA 97, 1483–1488 (2000).
He, W., Cowan, C.W. & Wensel, T.G. Neuron 20, 95–102 (1998).
Makino, E.R., Handy, J.W., Li, T. & Arshavsky, V.Y. Proc. Natl. Acad. Sci. USA 96, 1947–1952 (1999).
Chen, C.K. et al. Nature 403, 557–560 (2000).
Arshavsky, V. & Bownds, M.D. Nature 357, 416–417 (1992).
Lichtarge, O., Bourne, H.R. & Cohen, F.E. Proc. Natl. Acad. Sci. USA 93, 7507–7511 (1996).
Lichtarge, O., Yamamoto, K.R. & Cohen, F.E. J. Mol. Biol. 274, 325–337 (1997).
Onrust, R. et al. Science 275, 381–384 (1997).
Slep, K.C. et al. Nature, in the press (2001).
Tesmer, J.J., Berman, D.M., Gilman, A.G. & Sprang, S.R. Cell 89, 251–261 (1997).
Cowan, C.W., Wensel, T.G. & Arshavsky, V.Y. Methods Enzymol. 315, 524–538 (2000).
Nicholls, A., Sharp, K. & Honig, B. Proteins 11, 281–296 (1991).
Kraulis, P. J. Appl. Crystallogr. 24, 946–950 (1991).
Merritt, E. & Bacon, D. Methods Enzymol. 277, 505–524 (1997).
Acknowledgements
This work was supported by NIH, NLM, the Welch Foundation, and the Keck Center for Computational Biology. M.A.K. is a Howard Hughes Medical Institute Fellow of the Life Sciences Research Foundation. O.L gratefully acknowledges the support of the American Heart Association and NHGRI.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sowa, M., He, W., Slep, K. et al. Prediction and confirmation of a site critical for effector regulation of RGS domain activity. Nat Struct Mol Biol 8, 234–237 (2001). https://doi.org/10.1038/84974
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/84974
This article is cited by
-
CRF-based models of protein surfaces improve protein-protein interaction site predictions
BMC Bioinformatics (2014)
-
Integrating energy calculations with functional assays to decipher the specificity of G protein–RGS protein interactions
Nature Structural & Molecular Biology (2011)
-
Prediction of enzyme function based on 3D templates of evolutionarily important amino acids
BMC Bioinformatics (2008)
-
Compensatory Evolution of a WW Domain Variant Lacking the Strictly Conserved Trp Residue
Journal of Molecular Evolution (2008)
-
Evolutionary proteomics identifies amino acids essential for ligand-binding of the cytokinin receptor CHASE domain
BMC Evolutionary Biology (2007)