Abstract
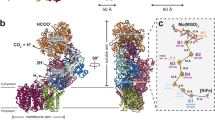
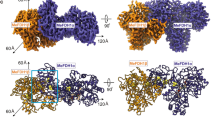
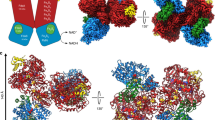
Oxidative decarboxylation of pyruvate to form acetyl–coenzyme A, a crucial step in many metabolic pathways, is carried out in most aerobic organisms by the multienzyme complex pyruvate dehydrogenase. In most anaerobes, the same reaction is usually catalyzed by a single enzyme, pyruvate:ferredoxin oxidoreductase (PFOR). Thus, PFOR is a potential target for drug design against certain anaerobic pathogens. Here, we report the crystal structures of the homodimeric Desulfovibrio africanus PFOR (data to 2.3 Å resolution), and of its complex with pyruvate (3.0 Å resolution). The structures show that each subunit consists of seven domains, one of which affords protection against oxygen. The thiamin pyrophosphate (TPP) cofactor and the three [4Fe–4S] clusters are suitably arranged to provide a plausible electron transfer pathway. In addition, the PFOR–pyruvate complex structure shows the noncovalent fixation of the substrate before the catalytic reaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Patel, M.S. & Roche, T.E. Molecular biology and biochemistry of pyruvate dehydrogenase complexes. FASEB J. 4, 3224–3233 (1990).
Kerscher, L. & Oesterhelt, D. Pyruvate : ferredoxin oxidoreductase — new findings on an ancient enzyme. Trends Biochem. Sci. 7, 371–374 (1982).
Smith, E.T., Blamey, J.M. & Adams, M.W. Pyruvate ferredoxin oxidoreductases of the hyperthermophilic archaeon, Pyrococcus furiosus, and the hyperthermophilic bacterium, Thermatoga maritima, have different catalytic mechanisms. Biochemistry 33, 1008–1016 (1994).
Uyeda, K. & Rabinowitz, J.C. Pyruvate–ferredoxin oxidoreductase. IV. Studies on the reaction mechanism. J Biol. Chem. 246, 3120–3125 (1971).
Wahl, R.C. & Orme–Johnson, W.H. Clostridial pyruvate oxidoreductase and the pyruvate–oxidizing enzyme specific to nitrogen fixation in Klebsiella pneumoniae are similar enzymes. J. Biol. Chem. 262, 10489–10496 (1987).
Hrdy, I. & Müller, M. Primary structure and eubacterial relationships of the pyruvate:ferredoxin oxidoreductase of the amitochondriate eukaryote Trichomonas vaginalis. J. Mol. Evol. 41, 388–396 (1995).
Hatchikian, E.C. & LeGall, J. Study of dicarboxylic acid and pyruvate metabolism in sulfate–reducing bacteria. II. Electron transport; final acceptors. Ann. Inst. Pasteur 118, 288–301 (1970).
Kletzin, A. & Adams, M.W.W. Molecular and phylogenetic characterization of pyruvate and ketoisovalerate ferredoxin oxidoreductases from Pyrococcus furiosus and pyruvate ferredoxin oxidoreductase from Thermotoga maritima. J. Bacteriol. 178, 248–257 (1996).
Bock, A.–K., Schönheit, P. & Teixeira, M. The iron–sulfur centers of the pyruvate : ferredoxin oxidoreductase from Methanosarcina barkeri (Fusaro). FEBS Lett. 414, 209–212 (1997).
Ben Rosenthal, B. et al. Evidence for the bacterial origin of genes encoding fermentation enzymes of the amitochondriate protozoan parasite Entamoeba histolytica. J. Bacteriol. 179, 3736–3745 (1997).
Brostedt, E. & Nordlund, S. Purification and partial characterization of a pyruvate oxidoreductase from the photosynthetic bacterium Rhodospirillum rubrum grown under nitrogen–fixing conditions. Biochem. J. 279, 155–158 (1991).
Menon, S. & Ragsdale, S.W. Unleashing hydrogenase activity in carbon monoxide dehydrogenase/acetyl–CoA synthase and pyruvate:ferredoxin oxidoreductase. Biochemistry 35, 12119–12125 (1996).
Evans, M.C.W., Buchanan, B.B. & Arnon, D.I. A new ferredoxin–dependent carbon reduction cycle in a photosynthetic bacterium. Proc. Natl. Acad. Sci. USA 55(4), 928–934 (1966).
Tersteegen, A., Linder, D., Thauer, R.K. & Hedderich, R. Structures and functions of four anabolic 2–oxoacid oxidoreductases in Methanobacterium thermoautotrophicum. Eur. J. Biochem. 242, 862–868 (1997).
Yoon, K.S., Ishii, M., Kodamam, T. & Igarashi, Y. Purification and characterization of pyruvate : ferredoxin oxidoreductase from Hydrogenobacter thermophilus TK–6. Arch. Microbiol. 167, 275–279 (1997).
Ma, K., Hutchins, A., Sung, S.J. & Adams, M.W.W. Pyruvate ferredoxin oxidoreductase from the hyperthermophilic archeon, Pyrococcus furiosus, functions as a CoA–dependent pyruvate decarboxylase. Proc. Natl. Acad. Sci. USA. 94, 9608–9613 (1997).
Zhang, Q., Iwasaki, T., Wakagi, T. & Oshima, T. 2–Oxoacid : ferredoxin oxidoreductase from the thermoacidophilic archeon, Sulfolobus sp. Strain 7. J. Biochem. (Tokyo) 120, 587–599 (1996).
Pieulle, L. et al. Isolation and characterization of the pyruvate – ferredoxin oxidoreductase from the sulfate–reducing bacterium Desulfovibrio africanus. Biochim. Biophys. Acta 1250, 49–59 (1995).
Pieulle, L., Magro, V. & Hatchikian, E.C. Isolation and analysis of the gene encoding the pyruvate–ferredoxin oxidoreductase of Desulfovibrio africanus, production of the recombinant enzyme in Escherichia coli, and effect of carboxy–terminal deletions on its stability. J. Bacteriol. 179, 5684–5692 (1997).
Matsubara, H. & Saeki, K. Structural and functional diversity of ferredoxins and related proteins. Adv. Inorg. Chem. 38, 223–280 (1992).
Moulis, J.M., Davasse, V., Meyer, J. & Gaillard, J. Molecular mechanism of pyruvate–ferredoxin oxidoreductases based on data obtained with the Clostridium pasteurianum enzyme. FEBS Lett. 380, 287–290 (1996).
Lindqvist, Y., Schneider, G., Ermler, U. & Sundström, M. Three–dimensional structure of transketolase, a thiamine diphosphate dependent enzyme, at 2.5 Å resolution. EMBO J. 11, 2373–2379 (1992).
Muller, Y.A. et al. A thiamin diphosphate binding fold revealed by comparison of the crystal structures of transketolase, pyruvate oxidase and pyruvate decarboxylase. Structure 1, 95–103 (1993).
Moser, C.C., Keske, J.M., Farid, R.S. & Dutton, P.L. Nature of biological electron transfer. Nature 355, 796–802 (1992).
Séry, A. et al. Crystal structure of the ferredoxin I from Desulfovibrio africanus at 2.3 Å resolution. Biochemistry 33, 15408–15417 (1994).
Menon, S. & Ragsdale, S.W. Mechanism of the Clostridium thermoaceticum pyruvate : ferredoxin oxidoreductase : Evidence for the common catalytic intermediacy of the hydroxyethylthiamine pyrophosphate radical. Biochemistry 36, 8484–8494 (1997).
Pianzzola, M.J., Soubes, M. & Touati, D. Overproduction of the rbo gene product from Desulfovibrio species suppresses all deleterious effects of lack of superoxide dismutase in Escherichia coli. J. Bacteriol. 178, 6736–6742 (1996).
Johnson, M.S., Zhulin, I.B., Gapuzan, M.E. R. & Taylor, B.L. Oxygen–dependent growth of the obligate anaerobe Desulfovibrio vulgaris Hildenborough. J. Bacteriol. 179, 5598–5601 (1997).
Kern, D. et al. How thiamine diphosphate is activated in enzymes. Science 275, 67–70 (1997).
Shin, W., Oh, D.G., Chae, C.H. & Yoon, T.S. Conformational analysis of thiamin–related compounds. A stereochemical model for thiamin catalysis. J. Am. Chem. Soc. 115, 12238–12250 (1993).
Breslow, R. The mechanism of thiamine action: Predictions from model experiments. Ann. NY Acad. Sci. 98, 445–452 (1962).
Lobell, M. & Crout, H.G. Pyruvate decarboxylase: A molecular modeling study of pyruvate decarboxylation and acyloin formation. J. Am. Chem. Soc. 118, 1867–1873 (1996).
Cammack, R., Kerscher, L. & Oesterhelt, D. A stable free radical intermediate in the reaction of 2–oxoacid : ferredoxin oxidoreductase of Halobium halobacterium. FEBS Lett. 118, 271–273 (1980).
Pieulle, L., Chabriere, E., Hatchikian, C., Fontecilla–Camps, J.C. & Charon, M.H. Crystallization and preliminary crystallographic analysis of the pyruvate:ferredoxin oxidoreductase from Desulfovibrio africanus. Acta Crystallogr. D, in the press.
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Collaborative computational project, number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994).
Hendrickson, W.A. Determination of macromolecular structures from anomalous diffraction of synchrotron radiation. Science 254, 51–58 (1991).
Sheldrick, G.M. Patterson superposition and ab initio phasing. Meth. Enz. 276, 628–641 (1997).
Otwinowski, Z. Isomorphous replacement and anomalous scattering. Proc. CCP4 Study Weekend 25–26 January 1991 (compiled by Wolf, W., Evans, P.R. & Leslie, A.G.W.), pp. 60–68 (1991).
Ramakrishnan, V. & Biou, V. Treatment of MAD data as a special case of MIR. Meth. Enz. 276, 538–557 (1997).
Wang, B.–C. Resolution of phase ambiguity in macromolecular crystallography. Meth. Enz. 115, 90–112 (1985).
Vellieux, F.M.D.A.P., Hunt, J.F., Roy, S. & Read, R.J. DEMON/ANGEL: A suite of programs to carry out density modification. J. Appl. Crystallogr. 28, 347–351 (1995).
Navaza, J. AMoRe: An automated package for molecular replacement. Acta Crystallogr. A 50, 157–163 (1994).
Jones, T.A., Zou, J.Y., Cowan, S.W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brünger, A.T. X–PLOR version 3.1. A system for X–ray crystallography and NMR. (Yale Univ. Press, New Haven, Connecticut,1992).
Brünger, A.T. Free R value: A novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992).
Jones, T.A. A graphic model building and refinement system for macromolecules. J. Appl. Crystallogr. 11, 268–272 (1978).
Kraulis, P.J. MOLSCRIPT: A program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Merrit, E.A. & Bacon, D.J. Raster3D: Photorealistic molecular graphics. Meth. Enz. 277, 505–524 (1997).
Acknowledgements
We are indebted to N. Forget for her assistance in the purification of the enzyme and to R. Toci for growing the bacteria. We thank J.L. Ferrer (BM02) and A. Thompson (BM14) for help with data collection at European Synchrotron Radiation Facilities. The assistance of M. Roth in the collection and treatment of the MAD data is gratefully acknowledged. This study was supported by the CEA and the CNRS.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Chabrière, E., Charon, M., Volbeda, A. et al. Crystal structures of the key anaerobic enzyme pyruvate:ferredoxin oxidoreductase, free and in complex with pyruvate. Nat Struct Mol Biol 6, 182–190 (1999). https://doi.org/10.1038/5870
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/5870
This article is cited by
-
Metagenomic characterization of a novel non-ammonia-oxidizing Thaumarchaeota from hadal sediment
Microbiome (2024)
-
H2 generated by fermentation in the human gut microbiome influences metabolism and competitive fitness of gut butyrate producers
Microbiome (2023)
-
A cell-free self-replenishing CO2-fixing system
Nature Catalysis (2022)
-
Improvement of anaerobic co-digestion of plant waste and excess sludge using calcium peroxide
Environmental Science and Pollution Research (2021)
-
Adaptations of Escherichia coli strains to oxidative stress are reflected in properties of their structural proteomes
BMC Bioinformatics (2020)