Abstract
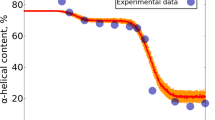
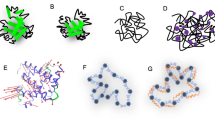
The structure and dynamics of two partially folded states of apomyoglobin have been characterized at equilibrium using multi-dimensional NMR spectroscopy. Residue-specific measurements of chemical shift and internal dynamics in these states and in the native apoprotein and holoprotein indicate progressive accumulation of secondary structure and increasing restriction of backbone dynamics as the chain collapses to form increasingly compact states. Under weakly folding conditions, the polypeptide fluctuates between unfolded states and local elements of structure that become extended and stabilized as the chain becomes more compact. These results provide a detailed model for molecular events that are likely to occur during folding of myoglobin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Matthews, C.R. Pathways of protein folding. Annu. Rev. Biochem. 62, 653–683 (1993).
Baldwin, R.L. The nature of protein folding pathways: the classical versus the new view. J. Biomol. NMR 5, 103–109 (1995).
Fersht, A.R. Characterizing transition states in protein folding: An essential step in the puzzle. Curr. Opin. Struct. Biol. 5, 79–84 (1995).
Dyson, H.J. & Wright, P.E. Insights into protein folding from NMR. Ann. Rev. Phys. Chem. 47, 369–395 (1996).
Bryngelson, J.D, Onuchic, J.N, Socci, N.D. & Wolynes, P.G. Funnels, pathways, and the energy landscape of protein folding: A synthesis. Proteins 21, 167–195 (1995).
Wolynes, P.G., Onuchic, J.N. & Thirumalai, D. Navigating the folding routes. Science 267, 1619–1620 (1995).
Jennings, P.A. & Wright, P.E. Formation of a molten globule intermediate early in the kinetic folding pathway of apomyoglobin. Science 262, 892–896 (1993).
Loh, S.N., kay M.S. & Baldwin, R.L. Structure and stability of a second molten globule intermediate in the apomyoglobin folding pathway. Proc. Natl. Acad. Sci. USA, 92, 5446–896 (1995).
Griko, Y.V., Privalov, P.L, Venyaminov, S.Y. & Kutyshenko, V.P. Thermodynamic study of the apomyoglobin structure. J. Mol. Biol. 202, 127–138 (1988).
Hughson, F.M., Wright, P.E. & Baldwin, R.L. Structural characterization of a partly folded apomyoglobin intermediate. Science 249, 1544–1548 (1990).
Eliezer, D. et al. The radius of gyration of an apomyoglobin folding intermediate. Science 270, 487–488 (1995).
Eliezer, D. & Wright, P.E. Is apomyoglobin a molten globule? Structural characterization by NMR. J. Mol. Biol. 263, 531–538 (1996)
Griko, Y.V. & Privalov, P.L. Thermodynamic puzzle of apomyoglobin unfolding. J. Mol. Biol. 235, 1381–1325 (1994)
Eliezer, D., Jennings, P.A., Dyson, H.J. & Wright, P.E. Populating the equilibrium molten globule state of apomyoglobin under conditions suitable for characterization by NMR. FEBS Lett. 417, 92–96 (1997).
Wright, P.E., Dyson, H.J. & Lerner, R.A. Conformation of peptide fragments of proteins in aqueous solution: implications for initiation of protein folding. Biochemistry 27, 7167–7175 (1988)
Nishii, I., Kataoka, M., Tokunaga, F. & Goto, Y. Cold denaturation of the molten globule states of apomyoglobin and a profile for protein folding. Biochemistry 33, 4903–4909 (1994).
Cocco, M.J. & Lecomte, J.TJ. Characterization of hydrophobic cores in apomyoglobin: a proton NMR spectroscopy study. Biochemistry 29, 11067–11072 (1990).
Lecomte, J.T.J ., Kao, Y.H. & Cocco, M.J. The native state of apomyoglobin described by proton NMR spectroscopy: The A-;B-;G-H interface of wild-type sperm whale apomyoglobin. Proteins 25, 267–285 (1996)
Jennings, P.A., Stone, M.J. & Wright, P.E. Overexpression of myoglobin and assignment of the amide, Cα and Cβ resonances. J. Biomol. NMR 6, 271–276 (1995).
Braun, D., Wider, G., & Wuthrich, K. Sequence-corrected 15N ”random coil“ chemical shifts. J. Am. Chem. Soc. 116, 8466–8469 (1994).
Yao, J., Dyson, H.J., & Wright, P.E. Chemical shift dispersion and secondary structure prediction in unfolded and partly-folded proteins. FEBS Lett. 419, 285–289 (1997).
Zhang, O., Kay, L.E., Shortle, D. & Forman-Kay, J.D. Comprehensive NOE characterization of a partly folded large fragment of staphylococcal nuclease D131D, using NMR methods with improved resolution. J. Mol. Biol. 272, 9–20 (1997).
Clore, G.M. & Gronenborn, A.M . Multidimensional heteronuclear nuclear magnetic resonance of proteins. Meth. Enz. 239, 349–363 (1994).
Baum, J., Dobson, C.M., Evans, P.A. & Hanley, C. Characterization of a partly folded protein by NMR methods: studies on the molten globule state of guinea pig a-lactalbumin. Biochemistry 28, 7–13 1998).
Alexandrescu, A.T., Evans, P.A., Pitkeathly, M., Baum, J. & Dobson, C.M. Structure and dynamics of the acid-denatured molten globule state of alpha-lactalbumin: A two-dimensional NMR study. Biochemistry 32, 1707–1718 (1993).
de Dios, A.C., Pearson, J.G. & Oldfield, E. Secondary and tertiary structural effects on protein NMR chemical shifts: an ab initio approach. Science 260, 1491–1496 (1993).
Spera, S. & Bax, A. Empirical correlation between protein backbone conformation and Ca and Cfi 13C nuclear magnetic resonance chemical shifts. J. Am. Chem. Soc. 113, 5490–5492 (1991).
Wishart, D.S. & Sykes, B.D. The 13C chemical-shift index: A simple method for the identification of protein secondary structure using 13C chemical-shift data. J. Biomol. NMR4, 4, 171–180 (1994).
Wishart, D.S., Bigam, C.G., Holm, A., Hodges, R.S. & Sykes, B.D. 1H, 13C and 15N random coil NMR chemical shifts of the common amino acids: I. Investigations of nearest neighbor effects. J. Biomol. NMR 5, 67–81 (1995).
Wuthrich, K., Billeter, M. & Braun, W. Polypeptide secondary structure determination by nuclear magnetic resonance observation of short proton-proton distances. J. Mol. Biol. 180, 715–740 (1984).
Waltho, J.P., Feher, V.A., Merutka, G., Dyson, H.J. & Wright, P.E. Peptide models of protein folding initiation sites. 1. Secondary structure formation by peptides corresponding to the G- and H-helices of myoglobin. Biochemistry 32, 6337–6347 (1993).
Kuriyan, J., Wilz, S., Karplus, M. & Petsko, G.A. X-ray structure and refinement of carbon-monoxy (Fe ll)-myoglobin at 1.5 A resolution. J. Mol. Biol. 192, 133–154 (1986).
Reymond, M.T., Merutka, G., Dyson, H.J. & Wright, P.E. Folding propensities of peptide fragments of myoglobin. Prot. So. 6, 706–716 (1997).
Kyte, J., Doolittle, R.F. A simple method for displaying the hydropathic character of a protein. J. Mol. Biol. 157, 105–132 (1982)
Alexandrescu, A.T. & Shortle, D. Backbone dynamics of a highly disordered 131 residue fragment of staphylococcal nuclease. J. Mol. Biol. 242, 527–546 (1994).
Muchmore, S.W. et al. X-ray and NMR structure of human Bcl-xL, an inhibitor of programmed cell death. Nature 381, 335–341 (1996).
Lin, L., Pinker, R.J., Forde, K., Rose, G.D. & Kallenbach, N.R. Molten globular characteristics of the native state of apomyoglobin. Nature Struct. Biol. 1, 447–452 (1994).
Fontana, A. et al. Probing the conformational state of apomyoglobin by limited proteolysis. J. Mol. Biol. 266, 223–230 (1997).
Ballew, R.M., Sabelko, J. & Gruebele, M. Direct observation of fast protein folding: The initial collapse of apomyoglobin. Proc. Natl. Acad. Sci. USA 93, 5759–5764 (1996).
Ballew, R.M., Sabelko, J. & Gruebele, M. Observation of distinct nanosecond and microsecond protein folding events. Nature Struct. Biol. 3, 923–926 (1996).
Chan, H.S. & Dill, K.A. Origins of structure in globular proteins. Proc. Natl. Acad. Sci. USA 87, 6388–6392 (1990).
Kataoka, M. et al. Structural characterization of the molten globule and native states of apomyoglobin by solution X-ray scattering. J. Mol. Biol. 249, 215–228 (1995).
Adler, A.J., Greenfield, N.J. & Fasman, G.D. Circular dichroism and optical rotatory dispersion of proteins and polypeptides. Meth. Enz. 27, 675–735 (1973).
Dyson, H.J., Ranee, M., Houghten, R.A., Wright, P.E. & Lerner, R.A. Folding of immunogenic peptide fragments of proteins in water solution. II The nascent helix. J. Mol. Biol. 201, 201–217 (1988).
Barrick, D. & Baldwin, R.L . Three-state analysis of sperm whale apomyoglobin folding. Biochemistry 32, 3790–3796 (1993)
Gast, K. et al. Compactness of protein molten globules: Temperature-induced structural changes of the apomyoglobin folding intermediate, Eur. Biophys. J. 23, 297–305 (1994)
Hughson, F.M., Barrick, D. & Baldwin, R.L. Probing the stability of a partly folded apomyoglobin intermediate by site-directed mutagenesis. Biochemistry 30, 4113–4118 (1991)
Kay, M.S. & Baldwin, R.L. Packing interactions in the apomyoglobin folding intermediate. Nature Struct. Biol. 3, 439–445 (1996).
Balbach, J. et al. Detection of residue contacts in a protein folding intermediate. Proc. Natal. Acad. Sci. USA 94, 7182–7185 (1997)
Kay, L.E., Ikura, M., Tschudin, R. & Bax, A. Three-dimensional triple-resonance NMR spectroscopy of isotopically enriched proteins. J. Magn. Reson. 89, 496–514 (1990)
Bax & Ikura, M An efficient 3D NMR technique for correlating the proton and 15N backbone amide resonances with the a-carbon of the preceding residue in uniformly 15N/13C enriched proteins. J. Biomol. NMR 1, 99–104 (1991).
Wittekind, M. & Mueller, L, HNCACB, a high-sensitivity 3D NMR experiment to correlate amide-proton and nitrogen resonances with the alpha- and beta-carbon resonances in proteins. J. Magn. Reson. 101, 201–205 (1993).
Grzesiek, S. & Bax, A Correlating backbone amide and side chain resonances in larger proteins by multiple relayed triple resonance NMR. J. Am. Chem. Soc. 114, 6291–6293 (1992).
Grzesiek, S. & Bax, A. Amino acid type determination in the sequential assignment procedure of uniformly 13C/15N-enriched proteins. J. Biomol. NMR 3, 185–204 (1993).
Lohr, F. & Ruterjans, H. A new triple-resonance experiment for the sequential assignment of backbone resonances in proteins. J. Biomol. NMR 6, 189–197 (1995).
Farrow, N.A. et al. Backbone dynamics of a free and a phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry 33, 5984–6003 (1994).
Brucker, E.A., Olson, J.S., Phillips, G.N., Jr., Dou, Y. & Ikeda-Saito, M. High resolution crystal structures of the deoxy, oxy and aquomet forms of cobalt myoglobin. J. Biol. Chem. 271, 25419–25422 (1996).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Eliezer, D., Yao, J., Dyson, H. et al. Structural and dynamic characterization of partially folded states of apomyoglobin and implications for protein folding. Nat Struct Mol Biol 5, 148–155 (1998). https://doi.org/10.1038/nsb0298-148
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsb0298-148
This article is cited by
-
Exploring the folding energy landscapes of heme proteins using a hybrid AWSEM-heme model
Journal of Biological Physics (2022)
-
Perspective: the essential role of NMR in the discovery and characterization of intrinsically disordered proteins
Journal of Biomolecular NMR (2019)
-
Initial Protein Unfolding Events in Ubiquitin, Cytochrome c and Myoglobin Are Revealed with the Use of 213 nm UVPD Coupled to IM-MS
Journal of the American Society for Mass Spectrometry (2019)
-
Probing the Time Scale of FPOP (Fast Photochemical Oxidation of Proteins): Radical Reactions Extend Over Tens of Milliseconds
Journal of the American Society for Mass Spectrometry (2016)
-
Spider wrapping silk fibre architecture arising from its modular soluble protein precursor
Scientific Reports (2015)