Abstract
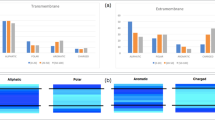
We present a comprehensive view of the tolerance of a membrane protein to sequence substitution. We find that the protein, diacylglycerol kinase from Escherichia coli, is extremely tolerant to sequence changes with three-quarters of the residues tolerating non-conservative changes. The conserved residues are distributed with approximately the same frequency in the soluble and transmembrane portions of the protein, but the most critical active-site residues appear to reside in the second cytoplasmic domain. It is remarkable that a unique structure of the membrane embedded portion of the protein can be encoded by a sequence that is so tolerant to substitution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Bowie, J.U., Reidhaar-Olson, J.F., Lim, W.A. & Sauer, R.T. Decipering the message in protein sequences: tolerance to amino acid substitution. Science 247, 1306–1310 (1990).
Loomis, C., Walsh, J. & Bell, R. sn-1,2-Diacylglycerol kinase of Escherichia coli. J. Biol. Chem. 260, 4091–4097 (1985).
Walsh, J. & Bell, R. sn-1,2-diacylglycerol kinase of Escherichia coli: Mixed micellar analysis of the phospholipid cofactor requirement and divalent cation dependence. J. Biol. Chem. 261, 6239–6247 (1986).
Walsh, J., Bell, R. sn-1,2-diacylglycerol kinase of Escherichia coli: structural and kinetic analysis of the lipid cofactor dependence. J. Biol. Chem. 261, 15062–15069 (1986).
Walsh, J., Fahrner, L. & Bell, R. sn-1,2-diacylglycerol kinase of Escherichia coli: Diacylglycerol analogues define specificity and mechanism. J. Biol. Chem. 265, 4374 (1990).
Smith, R., O'Toole, J., Maguire, M. & Sanders, C. Membrane topology of Escherichia coli diacylglycerol kinase. J. Bact. 176, 5459–5465 (1994).
Bowie, J.U., Sauer, R.T. Identifying determinants of folding and activity for a protein of unknown structure. Proc. Natl. Acad. Sci. USA. 86, 2152–2156 (1989).
Raetz, C. & Newman, K. Neutral lipid accumulation in the membranes of Escherichia coli mutants lacking diglyceride kinase. J. Biol. Chem. 253, 3882–3887 (1978).
Dayhoff, M.O. & Schwartz, R.M. in Atlas of Protein Sequence and Structure vol. 5 (ed. M.O. Dayhoff) 353 (National Biomedical Research Foundation, Washington, D.C., 1979).
Poteete, A., Rennell, D. & Bouvier, S. Functional significance of conserved amino acid residues, Proteins Struct. Funct. Genet. 13, 38–40 (1992).
Eisenberg, D., Weiss, R. & Terwilliger, T. The helical hydrophobic moment: A measure of the amphiphilicity of a helix. Nature 299, 371–374 (1982).
Hinkle, P., Hinkle, P. & Kaback, H. Information content of amino acid residues in putative helix VIII of the lac permease from Escherichia coli. Biochemistry 29, 10989–10994 (1990).
Lemmon, M., Flanagan, J., Treutlein, H., Zhang, J. & Engelman, D. Sequence specificity in the dimerization of transmembrane α-helices. Biochemistry 31, 12719–12725 (1992).
Lemmon, M. et al. Glycophorin A dimerization is driven by specific interaction between transmembrane α-Helices, J. Biol. Chem. 267, 7683–7689 (1992).
Williams, K. et al. Packing of coat protein amphipathic and transmembrane helices in filamentious bacteriphage M13: Role of small residues in protein oligomerization. J. Mol. Biol. 252, 6–14 (1995).
Rennell, D., Bouvier, S., Hardy, L. & Poteete, A. Systematic mutation of bacteriophage T4 lysozyme. J. Mol. Biol. 222, 67–87 (1991).
Kleina, L., Miller, J. Genetic studies of the lac represser XIII. Extensive amino acid replacements generated by the use of natural and synthetic nonsense suppressors. J. Mol. Biol. 212, 295–318 (1990).
Markiewicz, P., Kleina, L., Cruz, C., Ehret, S. & Miller, J. Analysis of 4000 altered Escherichia coli lac repressers resulting from suppression of nonsense mutations at 328 positions in the lacl gene. J. Mol. Biol., 240 421–433 (1994).
Normanly, J., Masson, J., Kleina, L., Abelson, J. & Miller, J. Construction of two Escherichia coli amber suppressor genes: tRNAPhe and tRNACys. Proc. Natl. Acad. Sci. USA. 83, 6548–52 (1986).
Kleina, L., Masson, J., Normanly, J., Abelson, J. & Miller, J. Construction of Escherichia coli amber suppressor tRNA genes II.Synthesis of additional tRNA genes and improvement of suppressor efficiency., J. Mol. Biol. 213, 705–717 (1990).
Bowie, J.U., Lüthy, R. & Eisenberg, D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 253, 164–170 (1991).
Lim, W.A. & Sauer, R.T. Alternative packing arrangements in the hydrophobic core of λ represser. Nature 339, 31–36 (1989).
Kamtekar, S., Schiffer, J., Xiong, H., Babik, J. & Hecht, M. Protein design by binary patterning of polar and nonpolar amino acids. Science 262, 1680–1685 (1993).
Lemmon, M. & Engelman, D. Helix-helix interactions inside lipid bilayers. Curr. Opin. Struct. Biol. 2, 511–18 (1992).
Sahin-Toth, M., Dunten, R., Gonzalez, A. & Kaback, H. Functional interactions between putative intramembrane charge residues in the lactose permease of Escherichia coli. Proc. Natl. Acad. Sci. USA. 89, 10547–10551 (1992).
Cosson, P., Lankford, S., Bonifacino, J. & Klausner, R. Membrane protein association by potential intramembrane charge pairs. Nature 351, 414–416 (1991).
Lim, W., Hodel, A., Sauer, R. & Richards, F. The crystal structure of a mutant protein with altered but improved hydrophobic core packing. Proc. Natl. Acad. Sci. USA. 91, 423–427 (1994).
Baldwin, E. & Matthews, B. Core-packing constraints, hydrophobicity and protein design. Curr. Opin. Biotechnol. 5, 396–402 (1994).
Heinz, D. et al. Accommodation of amino acid insertions in an alpha-helix of T4 lysozyme.Structural and thermodynamic analysis. J. Mol. Biol. 236, 869–86 (1994).
Weitzman, C. & Kaback, H. Cysteine scanning mutagenesis of helix V in the lactose permease of Escherichia coli. Biochemistry 34, 9374–9379 (1995).
Dunten, R., Sahin-Toth, M. & Kaback, H. Cysteine scanning mutagenesis of putative helix XI in the lactose permease of Escherichia coli. Biochemistry 32, 12644–12650 (1993).
Sahin-Toth, M. & Kaback, H. Cysteine scanning mutagenesis of putative transmembrane helices IX and X in the lactose permease of Escherichia coli. Prot. Sci. 2, 1024–1033 (1993).
Sahin-Toth, M., Persson, B., Schwieger, J., Cohan, P. & Kaback, H. Cysteine scanning mutagenesis of the N-terminal 32 amino acid residues in the lactose permease of Escherichia coli. Prot. Sci. 3, 240–247 (1994).
Frillingos, S., Sahin-Toth, M., Persson, B. & Kaback, H. Cysteine-scanning mutagenesis of putative helix VII in the lactose permease of Escherichia coli. Biochemistry 33, 8074–8081 (1994).
McDermott, G. et al., Crystal structure of an integral membrane light-harvesting complex from photosynthetic bacteria. Nature 374, 517–521 (1995).
Miller, K., McKinstry, M., Hunt, W. & Nixon, B. Identification of the diacylglycerol kinase structural gene of Rhizobium meliloti 1021. Mol. Plant Microbe Int. 5, 363–371 (1992).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wen, J., Chen, X. & Bowie, J. Exploring the allowed sequence space of a membrane protein. Nat Struct Mol Biol 3, 141–148 (1996). https://doi.org/10.1038/nsb0296-141
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0296-141