Abstract
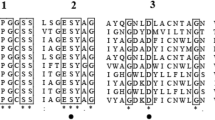
Aspartic proteinase A from yeast is specifically and potently inhibited by a small protein called IA3 from Saccharomyces cerevisiae . Although this inhibitor consists of 68 residues, we show that the inhibitory activity resides within the N-terminal half of the molecule. Structures solved at 2.2 and 1.8 Å, respectively, for complexes of proteinase A with full-length IA3 and with a truncated form consisting only of residues 2–34, reveal an unprecedented mode of inhibitor–enzyme interactions. Neither form of the free inhibitor has detectable intrinsic secondary structure in solution. However, upon contact with the enzyme, residues 2–32 become ordered and adopt a near-perfect α-helical conformation. Thus, the proteinase acts as a folding template, stabilizing the helical conformation in the inhibitor, which results in the potent and specific blockage of the proteolytic activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Handbook of proteolytic enzymes (eds Barrett, A.J., Woessner, J.F. & Rawlings, N.D.) 1–1650 (Academic Press, London; 1998).
Schreuder H.A. et al. Nature Struct. Biol. 1, 48– 54 (1994).
Engh, R.A. et al. J. Mol. Biol. 234, 1060– 1069 (1993).
Gomis-Ruth, F.X. et al. Nature, 389, 77–81 (1997).
Christeller, J.T., Farley, P. E., Ramsay, R.J., Sullivan, P. A. & Laing, W.A. Eur. J. Biochem. 254 , 160–167 (1998).
Dreyer, T., Valler, M. J., Kay, J., Charlton, P. & Dunn, B.M. Biochem. J. 231, 777– 779 (1985).
Craig, C. et al. AIDS 12, 1611–1618 (1998).
Dame, J.B. et al. Mol. Biochem. Parasitol. 64, 177– 190 (1994).
Monod, M., Togni, G., Hube, B. & Sanglard, D. Mol. Microbiol. 13, 357–368 ( 1994).
Wlodawer, A. & Vondrasek, J. Annu. Rev. Biophys. Biomol. Struct. 27, 249–284 ( 1998).
Moon, R.P. et al. Eur. J. Biochem. 244, 552– 560 (1997).
Roberts, N.A., Craig, J.C. & Sheldon, J. AIDS 12, 453– 460 (1998).
Takahashi, S. et al. J. Biochem. 125, 348– 353 (1999) .
Kageyama, T. Eur. J. Biochem. 253, 804–809 (1998).
Kreft, S. et al. Phytochemistry 44, 1001– 1006 (1997).
Saheki, T., Matsuda, Y. & Holzer, H. Eur. J. Biochem. 47, 325– 332 (1974).
Aguilar, C.F. et al. J. Mol. Biol. 267, 899– 915 (1997).
Schu, P. & Wolf, D.H. FEBS Lett. 283, 78–84 (1991).
Biedermann, K., Montali, U., Martin, B., Svendsen, I. B. & Ottesen, M. Carlsberg Res. Commun. 45, 225 –235 (1980).
Kondo, H. et al. J. Biochem. 124, 141– 147 (1998).
Rost, B. & Sander, C. J. Mol. Biol. 232, 584–599 (1993).
Davies, D.R. Annu. Rev. Biophys. Biophys. Chem. 19, 189– 215 (1990).
Richter, C., Tanaka, T., Koseki, T. & Yada, R.Y. Eur. J. Biochem. 261, 746–752 ( 1999).
Khan, A.R. & James, M.N.G. Protein Sci. 7, 815–836 (1998).
Wu, C.Y., Robertson, D.H.L., Hubbard, S.J., Gaskell, S. J. & Beynon, R. J. J. Biol. Chem. 274 , 1108–1115 (1999).
Cooper, J.B. et al. Biochemistry 28, 8596– 8603 (1989).
Tigue, N. J. & Kay, J. J. Biol. Chem. 273, 26441–26446 (1998).
Otwinowski, Z. & Minor, W. Methods Enzymol. 276, 307–326 ( 1997).
Navaza, J. Acta Crystallogr. A 50, 157–163 (1994).
Jones, T. A. & Kieldgaard, M. Methods Enzymol. 277, 173–208 (1997).
Brünger, A. T. et al. Acta Crystallogr. D 54, 905– 921 (1998).
Acknowledgements
This paper is dedicated to the memory of H. Holzer, formerly of the University of Freiburg im Breisgau, Germany. We thank A. Chung at the Protein Chemistry Core, University of Florida, for synthesis of peptides; A. Edison and R.A. Byrd for their valuable contributions with the NMR; V. Dhanaraj and B. Brownsey for their extensive help with modeling and mass spectroscopy analysis, respectively; J. Collins for help with the preparation of Fig. 1; and A. Arthur for editorial comments. This work was supported in part by awards from the BBSRC (ROPA) and from Actelion, Allschwil, Switzerland (J.K.), by an NIH grant (B.M.D.) and by the National Cancer Institute, DHHS, under contract with ABL (A.W.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Li, M., Phylip, L., Lees, W. et al. The aspartic proteinase from Saccharomyces cerevisiae folds its own inhibitor into a helix. Nat Struct Mol Biol 7, 113–117 (2000). https://doi.org/10.1038/72378
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/72378
This article is cited by
-
Homology Modeling and Analysis of Vacuolar Aspartyl Protease from a Novel Yeast Expression Host Meyerozyma guilliermondii Strain SO
Arabian Journal for Science and Engineering (2023)
-
Intrinsically Semi-disordered State and Its Role in Induced Folding and Protein Aggregation
Cell Biochemistry and Biophysics (2013)
-
Predicting functional residues of the Solanum lycopersicum aspartic protease inhibitor (SLAPI) by combining sequence and structural analysis with molecular docking
Journal of Molecular Modeling (2012)
-
Calcium-bound structure of calpain and its mechanism of inhibition by calpastatin
Nature (2008)
-
The Squash Aspartic Proteinase Inhibitor SQAPI Is Widely Present in the Cucurbitales, Comprises a Small Multigene Family, and Is a Member of the Phytocystatin Family
Journal of Molecular Evolution (2006)