Abstract
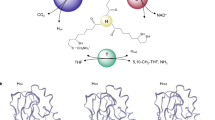
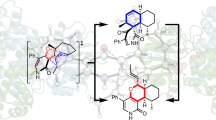
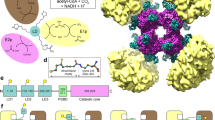
Glycine decarboxylase consists of four protein components. Its structural and mechanistic heart is provided by the lipoic acid-containing H-protein which undergoes a cycle of reductive methylamination, methylamine transfer and electron transfer. Lipoic acid attached to a specific lysine side chain is assumed to act as a ‘swinging arm’ conveying the reactive dithiolane ring from one catalytic centre to another. The X-ray crystal structures of two forms of the H-protein have been determined. The lipoate cofactor is located in the loop of a hairpin configuration but following methylamine transfer it is pivoted to bind into a cleft at the surface of the H-protein. The lipoamide-methylamine arm is, therefore, not free to move in aqueous solvent.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Douce, R., Bourguignon, J., Macherel, D. & Neuburger, M. The glycine decarboxylase system in higher plant mitochondria: Structure, function and biogenesis. Biochem. Soc. Trans. 22, 184–188 (1994).
Mattevi, A., de Kok, A. & Perham, R.N. The pyruvate dehydrogenase multienzyme complex. Curr. Opin. struct. Biol. 2, 877–887 (1992).
Reed, L.J. Multienzyme complexes. Acct. chem. Res. 7, 40–46 (1974).
Sieker, L., Cohen-Addad, C., Neuburger, M. & Douce, R. Crystallographic data for H-protein from the glycine decarboxylase complex. J. molec. Biol. 220, 223–224 (1991).
Pares, S., Cohen-Addad, C., Sieker, L., Neuburger, M. & Douce, R. X-ray structure determination at 2. 6 Å-resolution of a lipoate containing protein: The H-protein of the glycine decarboxylase complex from pea leaves. Proc. natn. Acad. Sci. U.S.A. 91, 4850–4853 (1994).
Dardel, F., Davis, A.L., Laue, E.D. & Perham, R.N. The three-dimensional structure of the lipoyl domain from Bacillus stearothermophilus pyruvate dehydrogenase multienzyme complex J. molec. Biol. 229, 1037–1048 (1993).
Brocklehurst, S.M. & Perham, R.N. Prediction of the three-dimensional structures of the biotinylated domain from yeast pyruvate carboxylase and of the lipoylated H-protein from the pea leaf glycine cleavage system: A new automated method for the prediction of protein tertiary sructure. Prot. Sci. 2, 626–639 (1993).
Neuburger, M., Jourdain, A. & Douce, R. Isolation of H-protein loaded with methylamine as a transient species in glycine decarboxylase reactions. Biochem. J. 278, 765–769 (1991).
Chothia, C. & Murzin, A.G. New folds for all-β proteins. Structure 1, 217–222 (1993).
Douce, R., Bourguignon, J., Brouquisse, R. & Neuburger, M. Isolation of plant mitochondria: general principles and criteria of integrity. Meths Enzymol. 148, 403–415 (1987).
Bourguignon, J., Neuburger, M. & Douce, R. Resolution and characterization of the glycine-cleavage reaction in pea leaf mitochondria. Biochem. J. 255, 169–178 (1988).
Navaza, J. AMoRe: an automated package for molecular replacement. Acta crystallogr. A50, 157–163 (1994).
Brunger, A.T., Kuriyan, J. and Karplus, M., Crystallographics R-factor refinement by molecular dynamics. Science 35, 458–460 (1987).
Kikuchi, G. and Hiraga, K. The mitochondrial glycine cleavage system. Molec. Cell. Biochem., 45, 137–149 (1982).
Carson, M. Ribbon 2.0 . J.appl.Crystallogr. 24, 958–961 (1991).
Jones, T.A., Zou, J.Y., Couran, S.W. and Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta crystallogr. A47, 110–119 (1991).
Mérand, V. et al. Characterization of the primary structure of H-protein from Pisum sativum and localisation of a lipoic acid residue by combined LC-MS and LC-MS-MS. Biol Mass Spectrometry 22, 447–456 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Cohen-Addad, C., Pares, S., Sieker, L. et al. The lipoamide arm in the glycine decarboxylase complex is not freely swinging. Nat Struct Mol Biol 2, 63–68 (1995). https://doi.org/10.1038/nsb0195-63
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nsb0195-63
This article is cited by
-
Structure-based dynamic analysis of the glycine cleavage system suggests key residues for control of a key reaction step
Communications Biology (2020)
-
Tribute Roland Douce, 1939–2018
Photosynthesis Research (2019)
-
Classification of ligand molecules in PDB with graph match-based structural superposition
Journal of Structural and Functional Genomics (2016)
-
The role of plant mitochondria in the biosynthesis of coenzymes
Photosynthesis Research (2007)
-
Structure of P-protein of the glycine cleavage system: implications for nonketotic hyperglycinemia
The EMBO Journal (2005)