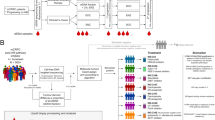
Key Points
-
Large-scale next-generation genetic analyses of prostate cancer emphasize the frequent occurrence and importance of focal genomic deletions inactivating PTEN
-
Phosphatase and tensin homologue (PTEN) loss in radical prostatectomy samples is often concurrent with genomic rearrangements involving the ETS family transcription factors
-
PTEN loss is reproducibly associated with adverse oncological outcomes by itself or in combination with other biomarkers and helps distinguish indolent tumours from those likely to progress
-
PTEN might be a useful prognostic biomarker to distinguish potentially aggressive Grade Group 1 or 2 tumours, which might make patients poor candidates for active surveillance programmes
-
Robust clinical assays using immunohistochemistry and fluorescence in situ hybridization (FISH) have been developed to reproducibly measure PTEN protein and gene loss using diagnostic tissue biopsies and circulating tumour cells from plasma
-
PTEN loss is associated with suppression of androgen receptor (AR) transcriptional output, and phosphoinositide 3-kinase (PI3K) inhibitors activate AR signalling, suggesting potential efficacy of combination therapies targeting the PI3K and AR signalling pathways
-
Emerging studies indicate that PTEN loss is associated with alterations to cellular interferon responses in the tumour microenvironment — tumours with loss of PTEN are more likely to have an immunosuppressive microenvironment, suggesting that advanced prostate cancers with PTEN loss might be amenable to immune-based therapies
Abstract
Genomic aberrations of the PTEN tumour suppressor gene are among the most common in prostate cancer. Inactivation of PTEN by deletion or mutation is identified in ∼20% of primary prostate tumour samples at radical prostatectomy and in as many as 50% of castration-resistant tumours. Loss of phosphatase and tensin homologue (PTEN) function leads to activation of the PI3K–AKT (phosphoinositide 3-kinase–RAC-alpha serine/threonine-protein kinase) pathway and is strongly associated with adverse oncological outcomes, making PTEN a potentially useful genomic marker to distinguish indolent from aggressive disease in patients with clinically localized tumours. At the other end of the disease spectrum, therapeutic compounds targeting nodes in the PI3K–AKT–mTOR (mechanistic target of rapamycin) signalling pathway are being tested in clinical trials for patients with metastatic castration-resistant prostate cancer. Knowledge of PTEN status might be helpful to identify patients who are more likely to benefit from these therapies. To enable the use of PTEN status as a prognostic and predictive biomarker, analytically validated assays have been developed for reliable and reproducible detection of PTEN loss in tumour tissue and in blood liquid biopsies. The use of clinical-grade assays in tumour tissue has shown a robust correlation between loss of PTEN and its protein as well as a strong association between PTEN loss and adverse pathological features and oncological outcomes. In advanced disease, assessing PTEN status in liquid biopsies shows promise in predicting response to targeted therapy. Finally, studies have shown that PTEN might have additional functions that are independent of the PI3K–AKT pathway, including those affecting tumour growth through modulation of the immune response and tumour microenvironment.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 67, 7–30 (2017).
Tosoian, J. J. et al. Intermediate and longer-term outcomes from a prospective active-surveillance program for favorable-risk prostate cancer. J. Clin. Oncol. 33, 3379–3385 (2015).
Cancer Genome Atlas Research, N. The molecular taxonomy of primary prostate cancer. Cell 163, 1011–1025 (2015).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Maehama, T. & Dixon, J. E. The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J. Biol. Chem. 273, 13375–13378 (1998).
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 13, 283–296 (2012).
Li, S. et al. The tumor suppressor PTEN has a critical role in antiviral innate immunity. Nat. Immunol. 17, 241–249 (2016).
Chen, L. & Guo, D. The functions of tumor suppressor PTEN in innate and adaptive immunity. Cell. Mol. Immunol. 14, 581–589 (2017).
Tamura, M. et al. Inhibition of cell migration, spreading, and focal adhesions by tumor suppressor PTEN. Science 280, 1614–1617 (1998).
Zhang, S. et al. Combating trastuzumab resistance by targeting SRC, a common node downstream of multiple resistance pathways. Nat. Med. 17, 461–469 (2011).
Weng, L. P., Brown, J. L. & Eng, C. PTEN coordinates G(1) arrest by down-regulating cyclin D1 via its protein phosphatase activity and up-regulating p27 via its lipid phosphatase activity in a breast cancer model. Hum. Mol. Genet. 10, 599–604 (2001).
Lindsay, Y. et al. Localization of agonist-sensitive PtdIns(3,4,5)P3 reveals a nuclear pool that is insensitive to PTEN expression. J. Cell Sci. 119, 5160–5168 (2006).
Shen, W. H. et al. Essential role for nuclear PTEN in maintaining chromosomal integrity. Cell 128, 157–170 (2007).
Bassi, C. et al. Nuclear PTEN controls DNA repair and sensitivity to genotoxic stress. Science 341, 395–399 (2013).
Cairns, P. et al. Frequent inactivation of PTEN/MMAC1 in primary prostate cancer. Cancer Res. 57, 4997–5000 (1997).
Steck, P. A. et al. Identification of a candidate tumour suppressor gene, MMAC1, at chromosome 10q23.3 that is mutated in multiple advanced cancers. Nat. Genet. 15, 356–362 (1997).
Suzuki, H. et al. Interfocal heterogeneity of PTEN/MMAC1 gene alterations in multiple metastatic prostate cancer tissues. Cancer Res. 58, 204–209 (1998).
Berger, M. F. et al. The genomic complexity of primary human prostate cancer. Nature 470, 214–220 (2011).
Barbieri, C. E. et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nat. Genet. 44, 685–689 (2012).
Yoshimoto, M. et al. FISH analysis of 107 prostate cancers shows that PTEN genomic deletion is associated with poor clinical outcome. Br. J. Cancer 97, 678–685 (2007).
Krohn, A. et al. Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion-positive and fusion-negative prostate cancer. Am. J. Pathol. 181, 401–412 (2012).
Troyer, D. A. et al. A multicenter study shows PTEN deletion is strongly associated with seminal vesicle involvement and extracapsular extension in localized prostate cancer. Prostate 75, 1206–1215 (2015).
Feilotter, H. E., Nagai, M. A., Boag, A. H., Eng, C. & Mulligan, L. M. Analysis of PTEN and the 10q23 region in primary prostate carcinomas. Oncogene 16, 1743–1748 (1998).
Pesche, S. et al. PTEN/MMAC1/TEP1 involvement in primary prostate cancers. Oncogene 16, 2879–2883 (1998).
Wang, S. I., Parsons, R. & Ittmann, M. Homozygous deletion of the PTEN tumor suppressor gene in a subset of prostate adenocarcinomas. Clin. Cancer Res. 4, 811–815 (1998).
Whang, Y. E. et al. Inactivation of the tumor suppressor PTEN/MMAC1 in advanced human prostate cancer through loss of expression. Proc. Natl Acad. Sci. USA 95, 5246–5250 (1998).
Yoshimoto, M. et al. Interphase FISH analysis of PTEN in histologic sections shows genomic deletions in 68% of primary prostate cancer and 23% of high-grade prostatic intra-epithelial neoplasias. Cancer Genet. Cytogenet. 169, 128–137 (2006).
Verhagen, P. C. et al. The PTEN gene in locally progressive prostate cancer is preferentially inactivated by bi-allelic gene deletion. J. Pathol. 208, 699–707 (2006).
McCall, P., Witton, C. J., Grimsley, S., Nielsen, K. V. & Edwards, J. Is PTEN loss associated with clinical outcome measures in human prostate cancer? Br. J. Cancer 99, 1296–1301 (2008).
Yoshimoto, M. et al. Absence of TMPRSS2:ERG fusions and PTEN losses in prostate cancer is associated with a favorable outcome. Mod. Pathol. 21, 1451–1460 (2008).
Attard, G. et al. Characterization of ERG, AR and PTEN gene status in circulating tumor cells from patients with castration-resistant prostate cancer. Cancer Res. 69, 2912–2918 (2009).
Sircar, K. et al. PTEN genomic deletion is associated with p-Akt and AR signalling in poorer outcome, hormone refractory prostate cancer. J. Pathol. 218, 505–513 (2009).
Han, B. et al. Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod. Pathol. 22, 1083–1093 (2009).
Krohn, A. et al. Heterogeneity and chronology of PTEN deletion and ERG fusion in prostate cancer. Mod. Pathol. 27, 1612–1620 (2014).
Steurer, S. et al. TMPRSS2-ERG fusions are strongly linked to young patient age in low-grade prostate cancer. Eur. Urol. 66, 978–981 (2014).
Lotan, T. L. et al. Analytic validation of a clinical-grade PTEN immunohistochemistry assay in prostate cancer by comparison with PTEN FISH. Mod. Pathol. 29, 904–914 (2016).
Ahearn, T. U. et al. A prospective investigation of PTEN loss and ERG expression in lethal prostate cancer. J. Natl Cancer Inst. 108, djv34 (2016).
Liu, W. et al. Copy number analysis indicates monoclonal origin of lethal metastatic prostate cancer. Nat. Med. 15, 559–565 (2009).
Grasso, C. S. et al. The mutational landscape of lethal castration-resistant prostate cancer. Nature 487, 239–243 (2012).
Robinson, D. et al. Integrative clinical genomics of advanced prostate cancer. Cell 161, 1215–1228 (2015).
Khani, F. et al. Evidence for molecular differences in prostate cancer between African American and Caucasian men. Clin. Cancer Res. 20, 4925–4934 (2014).
Tosoian, J. J. et al. Prevalence and prognostic significance of PTEN loss in African-American and European-American men undergoing radical prostatectomy. Eur. Urol. 71, 697–700 (2017).
Lindquist, K. J. et al. Mutational landscape of aggressive prostate tumors in African American men. Cancer Res. 76, 1860–1868 (2016).
Huang, F. W. et al. Exome sequencing of African-American prostate cancer reveals loss-of-function ERF mutations. Cancer Discov. 7, 973–983 (2017).
Reid, A. H. et al. Novel, gross chromosomal alterations involving PTEN cooperate with allelic loss in prostate cancer. Mod. Pathol. 25, 902–910 (2012).
Murphy, S. J. et al. Integrated analysis of the genomic instability of PTEN in clinically insignificant and significant prostate cancer. Mod. Pathol. 29, 143–156 (2016).
Ibeawuchi, C. et al. Exploring prostate cancer genome reveals simultaneous losses of PTEN, FAS and PAPSS2 in patients with PSA recurrence after radical prostatectomy. Int. J. Mol. Sci. 16, 3856–3869 (2015).
Whang, Y. E., Wu, X. Y. & Sawyers, C. L. Identification of a pseudogene that can masquerade as a mutant allele of the PTEN/MMAC1 tumor suppressor gene. J. Natl Cancer Inst. 90, 859–861 (1998).
Zysman, M. A., Chapman, W. B. & Bapat, B. Considerations when analyzing the methylation status of PTEN tumor suppressor gene. Am. J. Pathol. 160, 795–800 (2002).
Beltran, H. et al. Targeted next-generation sequencing of advanced prostate cancer identifies potential therapeutic targets and disease heterogeneity. Eur. Urol. 63, 920–926 (2013).
Bermudez Brito, M., Goulielmaki, E. & Papakonstanti, E. A. Focus on PTEN Regulation. Front. Oncol. 5, 166 (2015).
Garcia, J. M. et al. Promoter methylation of the PTEN gene is a common molecular change in breast cancer. Genes Chromosomes Cancer 41, 117–124 (2004).
Konishi, N. et al. Heterogeneous methylation and deletion patterns of the INK4a/ARF locus within prostate carcinomas. Am. J. Pathol. 160, 1207–1214 (2002).
Tay, Y. et al. Coding-independent regulation of the tumor suppressor PTEN by competing endogenous mRNAs. Cell 147, 344–357 (2011).
Poliseno, L. et al. Identification of the miR-106b∼25 microRNA cluster as a proto-oncogenic PTEN-targeting intron that cooperates with its host gene MCM7 in transformation. Sci. Signal. 3, ra29 (2010).
Leslie, N. R. & Foti, M. Non-genomic loss of PTEN function in cancer: not in my genes. Trends Pharmacol. Sci. 32, 131–140 (2011).
Salmena, L., Carracedo, A. & Pandolfi, P. P. Tenets of PTEN tumor suppression. Cell 133, 403–414 (2008).
Gundem, G. et al. The evolutionary history of lethal metastatic prostate cancer. Nature 520, 353–357 (2015).
Haffner, M. C. et al. Tracking the clonal origin of lethal prostate cancer. J. Clin. Invest. 123, 4918–4922 (2013).
Bismar, T. A. et al. PTEN genomic deletion is an early event associated with ERG gene rearrangements in prostate cancer. BJU Int. 107, 477–485 (2011).
Gumuskaya, B. et al. Assessing the order of critical alterations in prostate cancer development and progression by IHC: further evidence that PTEN loss occurs subsequent to ERG gene fusion. Prostate Cancer Prostat. Dis. 16, 209–215 (2013).
Lotan, T. L. et al. PTEN loss as determined by clinical-grade immunohistochemistry assay is associated with worse recurrence-free survival in prostate cancer. Eur. Urol. Focus 2, 180–188 (2016).
Lotan, T. L. et al. Cytoplasmic PTEN protein loss distinguishes intraductal carcinoma of the prostate from high-grade prostatic intraepithelial neoplasia. Mod. Pathol. 26, 587–603 (2013).
Morais, C. L. et al. Utility of PTEN and ERG immunostaining for distinguishing high-grade PIN from intraductal carcinoma of the prostate on needle biopsy. Am. J. Surg. Pathol. 39, 169–178 (2015).
Morais, C. L. et al. ERG and PTEN status of isolated high-grade PIN occurring in cystoprostatectomy specimens without invasive prostatic adenocarcinoma. Hum. Pathol. 55, 117–125 (2016).
Trotman, L. C. et al. Pten dose dictates cancer progression in the prostate. PLOS Biol. 1, E59 (2003).
Di Cristofano, A., Pesce, B., Cordon-Cardo, C. & Pandolfi, P. P. Pten is essential for embryonic development and tumour suppression. Nat. Genet. 19, 348–355 (1998).
Stambolic, V. et al. High incidence of breast and endometrial neoplasia resembling human Cowden syndrome in pten+/− mice. Cancer Res. 60, 3605–3611 (2000).
Podsypanina, K. et al. Mutation of Pten/Mmac1 in mice causes neoplasia in multiple organ systems. Proc. Natl Acad. Sci. USA 96, 1563–1568 (1999).
Wang, S. et al. Prostate-specific deletion of the murine Pten tumor suppressor gene leads to metastatic prostate cancer. Cancer Cell 4, 209–221 (2003).
Chen, Z. et al. Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 436, 725–730 (2005).
Shen, M. M. & Abate-Shen, C. Pten inactivation and the emergence of androgen-independent prostate cancer. Cancer Res. 67, 6535–6538 (2007).
Jiao, J. et al. Murine cell lines derived from Pten null prostate cancer show the critical role of PTEN in hormone refractory prostate cancer development. Cancer Res. 67, 6083–6091 (2007).
Mulholland, D. J. et al. Cell autonomous role of PTEN in regulating castration-resistant prostate cancer growth. Cancer Cell 19, 792–804 (2011).
Carver, B. S. et al. Reciprocal feedback regulation of PI3K and androgen receptor signaling in PTEN-deficient prostate cancer. Cancer Cell 19, 575–586 (2011).
Choucair, K. et al. PTEN genomic deletion predicts prostate cancer recurrence and is associated with low AR expression and transcriptional activity. BMC Cancer 12, 543 (2012).
Bismar, T. A. et al. Interactions and relationships of PTEN, ERG, SPINK1 and AR in castration-resistant prostate cancer. Histopathology 60, 645–652 (2012).
Grabowska, M. M. et al. Mouse models of prostate cancer: picking the best model for the question. Cancer Metastasis Rev. 33, 377–397 (2014).
Carver, B. S. et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat. Genet. 41, 619–624 (2009).
King, J. C. et al. Cooperativity of TMPRSS2-ERG with PI3-kinase pathway activation in prostate oncogenesis. Nat. Genet. 41, 524–526 (2009).
Chen, Y. et al. ETS factors reprogram the androgen receptor cistrome and prime prostate tumorigenesis in response to PTEN loss. Nat. Med. 19, 1023–1029 (2013).
Kim, J. et al. A mouse model of heterogeneous, c-MYC-initiated prostate cancer with loss of Pten and p53. Oncogene 31, 322–332 (2012).
Couto, S. S. et al. Simultaneous haploinsufficiency of Pten and Trp53 tumor suppressor genes accelerates tumorigenesis in a mouse model of prostate cancer. Differentiation 77, 103–111 (2009).
Ku, S. Y. et al. Rb1 and Trp53 cooperate to suppress prostate cancer lineage plasticity, metastasis, and antiandrogen resistance. Science 355, 78–83 (2017).
Tan, H. L. et al. Rb loss is characteristic of prostatic small cell neuroendocrine carcinoma. Clin. Cancer Res. 20, 890–903 (2014).
Aparicio, A. M. et al. Combined tumor suppressor defects characterize clinically defined aggressive variant prostate cancers. Clin. Cancer Res. 22, 1520–1530 (2016).
Hubbard, G. K. et al. Combined MYC activation and Pten loss are sufficient to create genomic instability and lethal metastatic prostate cancer. Cancer Res. 76, 283–292 (2016).
Blattner, M. et al. SPOP mutation drives prostate tumorigenesis in vivo through coordinate regulation of PI3K/mTOR and AR signaling. Cancer Cell 31, 436–451 (2017).
Zhao, D. et al. Synthetic essentiality of chromatin remodelling factor CHD1 in PTEN-deficient cancer. Nature 542, 484–488 (2017).
Moschini, M. et al. Low-risk prostate cancer: identification, management, and outcomes. Eur. Urol. 72, 238–249 (2017).
Bruinsma, S. M. et al. Active surveillance for prostate cancer: a narrative review of clinical guidelines. Nat. Rev. Urol. 13, 151–167 (2016).
Barrett, T. & Haider, M. A. The emerging role of MRI in prostate cancer active surveillance and ongoing challenges. AJR Am. J. Roentgenol. 208, 131–139 (2017).
Ma, T. M. et al. The role of multiparametric magnetic resonance imaging/ultrasound fusion biopsy in active surveillance. Eur. Urol. 71, 174–180 (2017).
Epstein, J. I. et al. The 2014 International Society of Urological Pathology (ISUP) Consensus Conference on Gleason Grading of Prostatic Carcinoma: definition of grading patterns and proposal for a new grading system. Am. J. Surg. Pathol. 40, 244–252 (2016).
Epstein, J. I. et al. A contemporary prostate cancer grading system: a validated alternative to the Gleason score. Eur. Urol. 69, 428–435 (2016).
Ross, A. E., D'Amico, A. V. & Freedland, S. J. Which, when and why? Rational use of tissue-based molecular testing in localized prostate cancer. Prostate Cancer Prostat. Dis. 19, 1–6 (2016).
Lotan, T. L. et al. PTEN protein loss by immunostaining: analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin. Cancer Res. 17, 6563–6573 (2011).
Wang, Y. & Dai, B. PTEN genomic deletion defines favorable prognostic biomarkers in localized prostate cancer: a systematic review and meta-analysis. Int. J. Clin. Exp. Med. 8, 5430–5437 (2015).
Reid, A. H. et al. Molecular characterisation of ERG, ETV1 and PTEN gene loci identifies patients at low and high risk of death from prostate cancer. Br. J. Cancer 102, 678–684 (2010).
Trock, B. J. et al. PTEN loss and chromosome 8 alterations in Gleason grade 3 prostate cancer cores predicts the presence of un-sampled grade 4 tumor: implications for active surveillance. Mod. Pathol. 29, 764–771 (2016).
Lotan, T. L. et al. PTEN loss is associated with upgrading of prostate cancer from biopsy to radical prostatectomy. Mod. Pathol. 28, 128–137 (2015).
Picanco-Albuquerque, C. G. et al. In prostate cancer needle biopsies, detections of PTEN loss by fluorescence in situ hybridization (FISH) and by immunohistochemistry (IHC) are concordant and show consistent association with upgrading. Virchows Arch. 468, 607–617 (2016).
Guedes, L. B., Tosoian, J. J., Hicks, J., Ross, A. E. & Lotan, T. L. PTEN loss in Gleason score 3 + 4 = 7 prostate biopsies is associated with nonorgan confined disease at radical prostatectomy. J. Urol. 197, 1054–1059 (2017).
Lokman, U., Erickson, A. M., Vasarainen, H., Rannikko, A. S. & Mirtti, T. PTEN loss but not ERG expression in diagnostic biopsies is associated with increased risk of progression and adverse surgical findings in men with prostate cancer on active surveillance. Eur. Urol. Focus https://doi.org/10.1016/j.euf.2017.03.004 (2017).
Mithal, P. et al. PTEN loss in biopsy tissue predicts poor clinical outcomes in prostate cancer. Int. J. Urol. 21, 1209–1214 (2014).
Shah, R. B., Bentley, J., Jeffery, Z. & DeMarzo, A. M. Heterogeneity of PTEN and ERG expression in prostate cancer on core needle biopsies: implications for cancer risk stratification and biomarker sampling. Hum. Pathol. 46, 698–706 (2015).
Wobker, S. E. & Epstein, J. I. Differential diagnosis of intraductal lesions of the prostate. Am. J. Surg. Pathol. 40, e67–e82 (2016).
Epstein, J. I. & Herawi, M. Prostate needle biopsies containing prostatic intraepithelial neoplasia or atypical foci suspicious for carcinoma: implications for patient care. J. Urol. 175, 820–834 (2006).
Tosoian, J. J., Alam, R., Ball, M. W., Carter, H. B. & Epstein, J. I. Managing high-grade prostatic intraepithelial neoplasia (HGPIN) and atypical glands on prostate biopsy. Nat. Rev. Urol. 15, 55–66 (2018).
Guo, C. C. & Epstein, J. I. Intraductal carcinoma of the prostate on needle biopsy: histologic features and clinical significance. Mod. Pathol. 19, 1528–1535 (2006).
Robinson, B. D. & Epstein, J. I. Intraductal carcinoma of the prostate without invasive carcinoma on needle biopsy: emphasis on radical prostatectomy findings. J. Urol. 184, 1328–1333 (2010).
Hickman, R. A. et al. Atypical intraductal cribriform proliferations of the prostate exhibit similar molecular and clinicopathologic characteristics as intraductal carcinoma of the prostate. Am. J. Surg. Pathol. 41, 550–556 (2017).
De Marzo, A. M., Haffner, M. C., Lotan, T. L., Yegnasubramanian, S. & Nelson, W. G. Premalignancy in prostate cancer: rethinking what we know. Cancer Prev. Res. 9, 648–656 (2016).
Pettersson, A. et al. The TMPRSS2:ERG rearrangement, ERG expression, and prostate cancer outcomes: a cohort study and meta-analysis. Cancer Epidemiol. Biomarkers Prev. 21, 1497–1509 EPI-12-0042 (2012).
Leinonen, K. A. et al. Loss of PTEN is associated with aggressive behavior in ERG-positive prostate cancer. Cancer Epidemiol. Biomarkers Prev. 22, 2333–2344 (2013).
Fontugne, J. et al. Recurrent prostate cancer genomic alterations predict response to brachytherapy treatment. Cancer Epidemiol. Biomarkers Prev. 23, 594–600 (2014).
Lahdensuo, K. et al. Loss of PTEN expression in ERG-negative prostate cancer predicts secondary therapies and leads to shorter disease-specific survival time after radical prostatectomy. Mod. Pathol. 29, 1565–1574 (2016).
Yoshimoto, M. et al. PTEN genomic deletions that characterize aggressive prostate cancer originate close to segmental duplications. Genes Chromosomes Cancer 51, 149–160 (2012).
Yoshimoto, M. et al. Incorporation of flanking probes reduces truncation losses for fluorescence in situ hybridization analysis of recurrent genomic deletions in tumor sections [abstract]. Cancer Res. 73, 63 (2014).
Sangale, Z. et al. A robust immunohistochemical assay for detecting PTEN expression in human tumors. Appl. Immunohistochem. Mol. Morphol. 19, 173–183 (2011).
Chaux, A. et al. Loss of PTEN expression is associated with increased risk of recurrence after prostatectomy for clinically localized prostate cancer. Mod. Pathol. 25, 1543–1549 (2012).
Punnoose, E. A. et al. PTEN loss in circulating tumour cells correlates with PTEN loss in fresh tumour tissue from castration-resistant prostate cancer patients. Br. J. Cancer 113, 1225–1233 (2015).
Wyatt, A. W. et al. Genomic alterations in cell-free DNA and enzalutamide resistance in castration-resistant prostate cancer. JAMA Oncol. 2, 1598–1606 (2016).
Xia, Y. et al. Copy number variations in urine cell free DNA as biomarkers in advanced prostate cancer. Oncotarget 7, 35818–35831 (2016).
Wyatt, A. W. et al. Concordance of circulating tumor DNA and matched metastatic tissue biopsy in prostate cancer. J. Natl Cancer Inst. 110, 78–86 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03236688 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02438007 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT03050866 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02853097 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02269982 (2018).
Ferraldeschi, R. et al. PTEN protein loss and clinical outcome from castration-resistant prostate cancer treated with abiraterone acetate. Eur. Urol. 67, 795–802 (2015).
Edlind, M. P. & Hsieh, A. C. PI3K-AKT-mTOR signaling in prostate cancer progression and androgen deprivation therapy resistance. Asian J. Androl. 16, 378–386 (2014).
Dillon, L. M. & Miller, T. W. Therapeutic targeting of cancers with loss of PTEN function. Curr. Drug Targets 15, 65–79 (2014).
Thomas, C. et al. Synergistic targeting of PI3K/AKT pathway and androgen receptor axis significantly delays castration-resistant prostate cancer progression in vivo. Mol. Cancer Ther. 12, 2342–2355 (2013).
Marques, R. B. et al. High efficacy of combination therapy using PI3K/AKT inhibitors with androgen deprivation in prostate cancer preclinical models. Eur. Urol. 67, 1177–1185 (2015).
Schwartz, S. et al. Feedback suppression of PI3Kalpha signaling in PTEN-mutated tumors is relieved by selective inhibition of PI3Kbeta. Cancer Cell 27, 109–122 (2015).
Armstrong, A. J. et al. Phase II trial of the PI3 kinase inhibitor buparlisib (BKM-120) with or without enzalutamide in men with metastatic castration resistant prostate cancer. Eur. J. Cancer 81, 228–236 (2017).
Wei, X. X. et al. A phase I study of abiraterone acetate combined with BEZ235, a dual PI3K/mTOR inhibitor, in metastatic castration resistant prostate cancer. Oncol. 22, e503–e543 (2017).
Massard, C. et al. Phase Ib dose-finding study of abiraterone acetate plus buparlisib (BKM120) or dactolisib (BEZ235) in patients with castration-resistant prostate cancer. Eur. J. Cancer 76, 36–44 (2017).
Chow, H. et al. A phase 2 clinical trial of everolimus plus bicalutamide for castration-resistant prostate cancer. Cancer 122, 1897–1904 (2016).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02106507 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02125084 (2017).
Baselga, J. et al. Everolimus in postmenopausal hormone-receptor-positive advanced breast cancer. N. Engl. J. Med. 366, 520–529 (2012).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02833883 (2017).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01485861 (2018).
de Bono, J. S. et al. PTEN loss as a predictive biomarker for the Akt inhibitor ipatasertib combined with abiraterone acetate in patients with metastatic castration-resistant prostate cancer (mCRPC). Ann. Oncol. 27, 718O (2016).
Fruman, D. A. et al. The PI3K pathway in human disease. Cell 170, 605–635 (2017).
Jia, S. et al. Essential roles of PI(3)K-p110beta in cell growth, metabolism and tumorigenesis. Nature 454, 776–779 (2008).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT01884285 (2018).
US National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/show/NCT02215096 (2018).
Mateo, J. et al. DNA repair in prostate cancer: biology and clinical implications. Eur. Urol. 71, 417–425 (2017).
Mateo, J. et al. DNA-repair defects and olaparib in metastatic prostate cancer. N. Engl. J. Med. 373, 1697–1708 (2015).
Gonzalez-Billalabeitia, E. et al. Vulnerabilities of PTEN-TP53-deficient prostate cancers to compound PARP-PI3K inhibition. Cancer Discov. 4, 896–904 (2014).
van de Ven, A. L. et al. Nanoformulation of olaparib amplifies PARP inhibition and sensitizes PTEN/TP53-deficient prostate cancer to radiation. Mol. Cancer Ther. 16, 1279–1289 (2017).
Hopkins, B. D. et al. A secreted PTEN phosphatase that enters cells to alter signaling and survival. Science 341, 399–402 (2013).
Wang, H. et al. Relevance and therapeutic possibility of PTEN-long in renal cell carcinoma. PLOS One 10, e114250 (2015).
Drake, C. G. Immunotherapy for prostate cancer: an emerging treatment modality. Urol. Clin. North Am. 37, 121–129 (2010).
De Marzo, A. M. et al. Inflammation in prostate carcinogenesis. Nat. Rev. Cancer 7, 256–269 (2007).
Strasner, A. & Karin, M. Immune infiltration and prostate cancer. Front. Oncol. 5, 128 (2015).
Gannon, P. O. et al. Characterization of the intra-prostatic immune cell infiltration in androgen-deprived prostate cancer patients. J. Immunol. Methods 348, 9–17 (2009).
Si, T. G., Wang, J. P. & Guo, Z. Analysis of circulating regulatory T cells (CD4+CD25+CD127-) after cryosurgery in prostate cancer. Asian J. Androl. 15, 461–465 (2013).
Peng, W. et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 6, 202–216 (2016).
Moussavi, M. et al. Oncolysis of prostate cancers induced by vesicular stomatitis virus in PTEN knockout mice. Cancer Res. 70, 1367–1376 (2010).
Champion, B. R., Fisher, K. & Seymour, L. A. PTENtial cause for the selectivity of oncolytic viruses? Nat. Immunol. 17, 225–226 (2016).
Pencik, J. et al. STAT3 regulated ARF expression suppresses prostate cancer metastasis. Nat. Commun. 6, 7736 (2015).
Yu, H., Pardoll, D. & Jove, R. STATs in cancer inflammation and immunity: a leading role for STAT3. Nat. Rev. Cancer 9, 798–809 (2009).
Toso, A. et al. Enhancing chemotherapy efficacy in Pten-deficient prostate tumors by activating the senescence-associated antitumor immunity. Cell Rep. 9, 75–89 (2014).
Cuzick, J. et al. Prognostic value of PTEN loss in men with conservatively managed localised prostate cancer. Br. J. Cancer 108, 2582–2589 (2013).
Lotan, T. L. et al. PTEN loss detection in prostate cancer: comparison of PTEN immunohistochemistry and PTEN FISH in a large retrospective prostatectomy cohort. Oncotarget 8, 65566–65576 (2017).
Acknowledgements
Funding for this research was provided in part by a Transformative Impact Award from the Congressionally Directed Medical Research Program–Prostate Cancer Research Program (CDMRP-PCRP) (W81XWH-13-2-0070, H.I.S. and T.L.L.). T.L.L. was additionally supported by the NIH and National Cancer Institute (NCI) P30 Cancer Center Support Grant CA006973 and the Patrick Walsh Prostate Cancer Research Fund. H.I.S. was additionally supported by NIH and NCI Prostate SPORE Grant P50-CA92629, NIH and NCI P30 Cancer Center Support Grant CA008748, and the Prostate Cancer Foundation. T.J. and D.M.B. were funded by Prostate Cancer Canada and the Movember Foundation (Grant #T2014-01-PRONTO). T.J. was supported by a Transformative Pathology Fellowship funded by the Ontario Institute for Cancer Research through funding provided by the Government of Ontario.
Author information
Authors and Affiliations
Contributions
All authors researched data for the article, took part in discussions of the content, and wrote the manuscript. T.J., D.M.B., H.I.S., A.M.D.M., J.A.S., and T.L.L. reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
T.L.L. has received research support from Ventana Medical Systems. D.M.B. has received financial support from Myriad Genetics and Metamark Genetics.
Rights and permissions
About this article
Cite this article
Jamaspishvili, T., Berman, D., Ross, A. et al. Clinical implications of PTEN loss in prostate cancer. Nat Rev Urol 15, 222–234 (2018). https://doi.org/10.1038/nrurol.2018.9
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrurol.2018.9
This article is cited by
-
PTEN-regulated PI3K-p110 and AKT isoform plasticity controls metastatic prostate cancer progression
Oncogene (2024)
-
Bridging Health Disparities: a Genomics and Transcriptomics Analysis by Race in Prostate Cancer
Journal of Racial and Ethnic Health Disparities (2024)
-
Current Systemic Therapy in Men with Metastatic Castration-Sensitive Prostate Cancer
Current Oncology Reports (2024)
-
Phase 1b study of enzalutamide plus CC-115, a dual mTORC1/2 and DNA-PK inhibitor, in men with metastatic castration-resistant prostate cancer (mCRPC)
British Journal of Cancer (2024)
-
In silico analysis of prognostic and diagnostic significance of target genes from prostate cancer cell lines derived exomicroRNAs
Cancer Cell International (2023)