Abstract
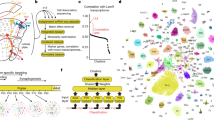
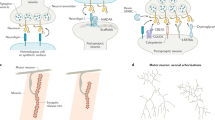
Our understanding of how the enormously complex task of interconnecting millions of nerve cells is accomplished remains rudimentary. What molecular mechanisms control its exquisite specificity? Can we pinpoint single molecular interactions that might help to explain some of the specificity requirements that underlie neuronal wiring? A series of recent studies on the molecular diversity of the Drosophila melanogaster cell-surface receptor Down syndrome cell-adhesion molecule (Dscam) provide one exceptional example of a novel mechanistic model of neuronal-wiring specificity, progressing from structural studies of single protein–protein interactions to biochemical analysis in vitro and to an understanding of complex neuronal differentiation at the single-cell or tissue levels.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Schmucker, D. & Flanagan, J. G. Generation of recognition diversity in the nervous system. Neuron 44, 219–222 (2004).
Zipursky, S. L., Wojtowicz, W. M. & Hattori, D. Got diversity? Wiring the fly brain with Dscam. Trends Biochem. Sci. 31, 581–588 (2006).
Muotri, A. & Gage, F. Generation of neuronal variability and complexity. Nature 441, 1087–1093 (2006).
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000).
Chen, B. E. et al. The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila. Cell 125, 607–620 (2006).
Wojtowicz, W. M., Flanagan, J. J., Millard, S. S., Zipursky, S. L. & Clemens, J. C. Alternative splicing of Drosophila Dscam generates axon guidance receptors that exhibit isoform-specific homophilic binding. Cell 118, 619–633 (2004).
Wojtowicz, W. M. et al. A vast repertoire of Dscam binding specificities arises from modular interactions of variable Ig domains. Cell 130, 1134–1145 (2007).
Hughes, M. et al. Homophilic Dscam interactions control complex dendrite morphogenesis. Neuron 54, 417–427 (2007).
Soba, P. et al. Drosophila sensory neurons require Dscam for dendritic self-avoidance and proper dendritic field organization. Neuron 54, 403–416 (2007).
Matthews, B. et al. Dendrite self-avoidance is controlled by Dscam. Cell 129, 593–604 (2007).
Meijers, R. et al. Structural basis of Dscam isoform specificity. Nature 449, 487–491 (2007).
Yamakawa, K. et al. DSCAM: a novel member of the immunoglobulin superfamily maps in a Down syndrome region and is involved in the development of the nervous system. Hum. Mol. Genet. 7, 227–237 (1998).
Agarwala, K. et al. Cloning and functional characterization of DSCAML1, a novel DSCAM-like cell adhesion molecule that mediates homophilic intercellular adhesion. Biochem. Biophys. Res. Commun. 285, 760–772 (2001).
Graveley, B. R. et al. The organization and evolution of the dipteran and hymenopteran Down syndrome cell adhesion molecule (Dscam) genes. RNA 10, 1499–1506 (2004).
Li, W. & Guan, K. The Down syndrome cell adhesion molecule (DSCAM) interacts with and activates Pak. J. Biol. Chem. 279, 32824–32831 (2004).
Zhan, X. L. et al. Analysis of Dscam diversity in regulating axon guidance in Drosophila mushroom bodies. Neuron 43, 673–686 (2004).
Watson, F. L. et al. Extensive diversity of Ig-superfamily proteins in the immune system of insects. Science 309, 1874–1878 (2005).
Neves, G., Zucker, J., Daly, M. & Chess, A. Stochastic yet biased expression of multiple Dscam splice variants by individual cells. Nature Genet. 36, 240–246 (2004).
Celotto, A. M. & Graveley, B. R. Alternative splicing of the Drosophila Dscam pre-mRNA is both temporally and spatially regulated. Genetics 159, 599–608 (2001).
Hummel, T. et al. Axonal targeting of olfactory receptor neurons in Drosophila is controlled by Dscam. Neuron 37, 221–231 (2003).
Wang, J. et al. Transmembrane/juxtamembrane domain-dependent Dscam distribution and function during mushroom body neuronal morphogenesis. Neuron 43, 663–672 (2004).
Wang, J., Zugates, C. T., Liang, I. H., Lee, C. H. & Lee, T. Drosophila Dscam is required for divergent segregation of sister branches and suppresses ectopic bifurcation of axons. Neuron 33, 559–571 (2002).
Zhu, H. et al. Dendritic patterning by Dscam and synaptic partner matching in the Drosophila antennal lobe. Nature Neurosci. 9, 349–355 (2006).
Hattori, D. et al. Dscam diversity is essential for neuronal wiring and self-recognition. Nature 449, 223–227 (2007).
Dong, Y., Taylor, H. & Dimopoulos, G. AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol. 4, 1137–1146 (2006).
Boehm, T. Two in one: dual function of an invertebrate antigen receptor. Nature Immunol. 8, 1031–1033 (2007).
Grueber, W. B., Jan, L. Y. & Jan, Y. N. Tiling of the Drosophila epidermis by multidendritic sensory neurons. Development 129, 2867–2878 (2002).
Gao, F. B., Brenman, J. E., Jan, L. Y. & Jan, Y. N. Genes regulating dendritic outgrowth, branching, and routing in Drosophila. Genes Dev. 13, 2549–2561 (1999).
Sugimura, K. et al. Distinct developmental modes and lesion-induced reactions of dendrites of two classes of Drosophila sensory neurons. J. Neurosci. 23, 3752–3760 (2003).
Mombaerts, P. et al. Visualizing an olfactory sensory map. Cell 87, 675–686 (1996).
Wang, F., Nemes, A., Mendelsohn, M. & Axel, R. Odorant receptors govern the formation of a precise topographic map. Cell 93, 47–60 (1998).
Bozza, T., Feinstein, P., Zheng, C. & Mombaerts, P. Odorant receptor expression defines functional units in the mouse olfactory system. J. Neurosci. 22, 3033–3043 (2002).
Vassalli, A., Rothman, A., Feinstein, P., Zapotocky, M. & Mombaerts, P. Minigenes impart odorant receptor-specific axon guidance in the olfactory bulb. Neuron 35, 681–696 (2002).
Feinstein, P. & Mombaerts, P. A contextual model for axonal sorting into glomeruli in the mouse olfactory system. Cell 117, 817–831 (2004).
Serizawa, S. et al. A neuronal identity code for the odorant receptor-specific and activity-dependent axon sorting. Cell 127, 1057–1069 (2006).
Author information
Authors and Affiliations
Related links
Rights and permissions
About this article
Cite this article
Schmucker, D. Molecular diversity of Dscam: recognition of molecular identity in neuronal wiring. Nat Rev Neurosci 8, 915–920 (2007). https://doi.org/10.1038/nrn2256
Issue Date:
DOI: https://doi.org/10.1038/nrn2256
This article is cited by
-
Alternative splicing of Drosophila Nmnat functions as a switch to enhance neuroprotection under stress
Nature Communications (2015)
-
Improvement of Dscam homophilic binding affinity throughout Drosophilaevolution
BMC Evolutionary Biology (2014)
-
Gene expression signature of cerebellar hypoplasia in a mouse model of Down syndrome during postnatal development
BMC Genomics (2009)
-
IgCAMs redundantly control axon navigation in Caenorhabditis elegans
Neural Development (2009)
-
The Pak1 Kinase: An Important Regulator of Neuronal Morphology and Function in the Developing Forebrain
Molecular Neurobiology (2008)