Key Points
-
In addition to being an important member of the normal intestinal microflora of humans and other mammals, the species Escherichia coli contains many pathotypes that cause a variety of diseases. At least six different pathotypes cause enteric disease, such as diarrhoea or dysentery, and other pathotypes cause extra-intestinal infections, including urinary tract infections and meningitis.
-
Virulence factors of E. coli can affect a wide range of eukaryotic cellular processes, including cell signalling, ion secretion, protein synthesis, mitosis, cytoskeletal function and mitochondrial function.
-
Virulence factors of pathogenic E. coli are frequently encoded on genetic elements such as plasmids, bacteriophage, transposons and pathogenicity islands that can be mobilized into different strains to create novel combinations of virulence factors.
-
The genomic structure of the E. coli pathotypes that have been sequenced so far show a striking mosaic pattern, with 2,000 genes present in 247 islands in one pathotype that are not present in K-12. Up to 0.53 MB of DNA present in K-12 can also be absent from pathogenic E. coli.
-
Genes that encode virulence factors of pathogenic E. coli are regulated by both pathotype-specific regulators that are absent from commensal E. coli, and by 'housekeeping' regulators that are present in commensal E. coli.
Abstract
Few microorganisms are as versatile as Escherichia coli. An important member of the normal intestinal microflora of humans and other mammals, E. coli has also been widely exploited as a cloning host in recombinant DNA technology. But E. coli is more than just a laboratory workhorse or harmless intestinal inhabitant; it can also be a highly versatile, and frequently deadly, pathogen. Several different E. coli strains cause diverse intestinal and extraintestinal diseases by means of virulence factors that affect a wide range of cellular processes.
Similar content being viewed by others
Main
Escherichia coli typically colonizes the gastrointestinal tract of human infants within a few hours after birth. Usually, E. coli and its human host coexist in good health and with mutual benefit for decades. These commensal E. coli strains rarely cause disease except in immunocompromised hosts or where the normal gastrointestinal barriers are breached — as in peritonitis, for example. The niche of commensal E. coli is the mucous layer of the mammalian colon. The bacterium is a highly successful competitor at this crowded site, comprising the most abundant facultative anaerobe of the human intestinal microflora. Despite the enormous body of literature on the genetics and physiology of this species, the mechanisms whereby E. coli assures this auspicious symbiosis in the colon are poorly characterized. One interesting hypothesis suggests that E. coli might exploit its ability to utilize gluconate in the colon more efficiently than other resident species, thereby allowing it to occupy a highly specific metabolic niche1.
However, there are several highly adapted E. coli clones that have acquired specific virulence attributes, which confers an increased ability to adapt to new niches and allows them to cause a broad spectrum of disease. These virulence attributes are frequently encoded on genetic elements that can be mobilized into different strains to create novel combinations of virulence factors, or on genetic elements that might once have been mobile, but have now evolved to become 'locked' into the genome. Only the most successful combinations of virulence factors have persisted to become specific 'PATHOTYPES' of E. coli that are capable of causing disease in healthy individuals. Three general clinical syndromes can result from infection with one of these pathotypes: enteric/diarrhoeal disease, urinary tract infections (UTIs) and sepsis/meningitis. Among the intestinal pathogens there are six well-described categories: enteropathogenic E. coli (EPEC), enterohaemorrhagic E. coli (EHEC), enterotoxigenic E. coli (ETEC), enteroaggregative E. coli (EAEC), enteroinvasive E. coli (EIEC) and diffusely adherent E. coli (DAEC)2 (Fig. 1). UTIs are the most common extraintestinal E. coli infections and are caused by uropathogenic E. coli (UPEC). An increasingly common cause of extraintestinal infections is the pathotype responsible for meningitis and sepsis — meningitis-associated E. coli (MNEC). The E. coli pathotypes implicated in extraintestinal infections have recently been called ExPEC3. EPEC, EHEC and ETEC can also cause disease in animals using many of the same virulence factors that are present in human strains and unique colonization factors that are not found in human strains (Table 1). An additional animal pathotype, known as avian pathogenic E. coli (APEC), causes extraintestinal infections — primarily respiratory infections, pericarditis, and septicaemia of poultry. This review will focus on E. coli strains that are pathogenic for humans.
The six recognized categories of diarrhoeagenic E. coli each have unique features in their interaction with eukaryotic cells. Here, the interaction of each category with a typical target cell is schematically represented. These descriptions are largely the result of in vitro studies and might not completely reflect the phenomena that occurs in infected humans. a | EPEC adhere to small bowel enterocytes, but destroy the normal microvillar architecture, inducing the characteristic attaching and effacing lesion. Cytoskeletal derangements are accompanied by an inflammatory response and diarrhoea. 1. Initial adhesion, 2. Protein translocation by type III secretion, 3. Pedestal formation. b | EHEC also induce the attaching and effacing lesion, but in the colon. The distinguishing feature of EHEC is the elaboration of Shiga toxin (Stx), systemic absorption of which leads to potentially life-threatening complications. c | Similarly, ETEC adhere to small bowel enterocytes and induce watery diarrhoea by the secretion of heat-labile (LT) and/or heat-stable (ST) enterotoxins. d | EAEC adheres to small and large bowel epithelia in a thick biofilm and elaborates secretory enterotoxins and cytotoxins. e | EIEC invades the colonic epithelial cell, lyses the phagosome and moves through the cell by nucleating actin microfilaments. The bacteria might move laterally through the epithelium by direct cell-to-cell spread or might exit and re-enter the baso-lateral plasma membrane. f | DAEC elicits a characteristic signal transduction effect in small bowel enterocytes that manifests as the growth of long finger-like cellular projections, which wrap around the bacteria. AAF, aggregative adherence fimbriae; BFP, bundle-forming pilus; CFA, colonization factor antigen; DAF, decay-accelerating factor; EAST1, enteroaggregative E. coli ST1; LT, heat-labile enterotoxin; ShET1, Shigella enterotoxin 1; ST, heat-stable enterotoxin.
The various pathotypes of E. coli tend to be clonal groups that are characterized by shared O (lipopolysaccharide, LPS) and H (flagellar) antigens that define SEROGROUPS (O antigen only) or SEROTYPES (O and H antigens)2,4. Pathogenic E. coli strains use a multi-step scheme of pathogenesis that is similar to that used by other mucosal pathogens, which consists of colonization of a mucosal site, evasion of host defences, multiplication and host damage. Most of the pathogenic E. coli strains remain extracellular, but EIEC is a true intracellular pathogen that is capable of invading and replicating within epithelial cells and macrophages. Other E. coli strains might be internalized by epithelial cells at low levels, but do not seem to replicate intracellularly.
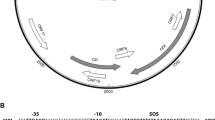
Adhesion/colonization. Pathogenic E. coli strains possess specific adherence factors that allow them to colonize sites that E. coli does not normally inhabit, such as the small intestine and the urethra (Table 1). Most frequently these adhesins form distinct morphological structures called fimbriae (also called pili) or fibrillae, which can belong to one of several different classes (Fig. 2). Fimbriae are rod-like structures of 5–10 nm diameter that are distinct from flagella. Fibrillae are 2–4 nm in diameter, and are either long and wiry or curly and flexible5. The Afa adhesins that are produced by many diarrhoeagenic and uropathogenic E. coli are described as afimbrial adhesins, but in fact seem to have a fine fibrillar structure that is difficult to visualize6. Adhesins of pathogenic E. coli can also include outer-membrane proteins, such as intimin of UPEC and EHEC, or other non-fimbrial proteins. Some surface structures trigger signal transduction pathways or cytoskeletal rearrangements that can lead to disease. For example, the members of the Dr family of adhesins that are expressed by DAEC and UPEC bind to the DECAY-ACCELERATING FACTOR (DAF, also known as CD55), which results in activation of phosphatidylinositol 3-kinase (PI-3-kinase) and cell-surface expression of the major histocompatibility complex (MHC) class I-related molecule MICA7. The IcsA protein of EIEC nucleates actin filaments at one pole of the bacterium, which allows it to move within the cytoplasm and into adjacent epithelial cells on a 'tail' of polymerized actin8. Even surface structures that are present on commensal E. coli strains can induce signalling cascades if the organism encounters the appropriate receptor. The LPS of E. coli and other Gram-negative bacteria binds to Toll-like receptor 4 (TLR4), triggering a potent cytokine cascade that can lead to septic shock and death9. Flagellin, the main component of flagella, can bind to TLR5, thereby activating interleukin (IL)-8 expression and an inflammatory response10.
E. coli produce a variety of colonization factors, many of which are hair-like structures of various morphologies called fimbriae (also called pili) or fibrillae. a | Long, straight colonization factor antigen (CFA)/III fimbriae of ETEC (5–7 nm in diameter) protruding peritrichously from the bacterial surface. b | Abundant long, straight CFA/I fimbriae (5–7 nm) of ETEC contrasting with thicker, wavy flagella. c | P pili of UPEC showing the thin (∼3 nm) fibrillar adhesive tip at the end of the pilus (∼10 nm). d | Thin (2–3 nm), flexible, wiry CS3 fibrillar structures produced by ETEC that extend several micrometres from the cell surface. e | Bundle-forming pilus (BFP) of EPEC, a member of the type IV pili family, aggregates laterally to form large rope-like structures (>10 μm long) of variable width. f | Thin (2–5 nm), coiled, highly aggregative curli fibres produced by a variety of pathogenic and non-pathogenic E. coli. Additional characteristics of colonization factors of diarrhoeagenic E. coli have been reviewed elsewhere (see Ref. 5). Panels a,b,d–f are courtesy of J. Girón. Panel c is reproduced from Ref. 147 Nature © Macmillan Magazines Ltd (1992).
Toxins. More numerous than surface structures that trigger signal transduction pathways are secreted toxins and other effector proteins that affect an astonishing variety of fundamental eukaryotic processes (Table 2). Concentrations of important intracellular messengers, such as cyclic AMP, cyclic GMP and Ca2+, can be increased, which leads to ion secretion by the actions of the heat-labile enterotoxin (LT), heat-stable enterotoxin a (STa) and heat-stable enterotoxin b (STb), respectively — all of which are produced by different strains of ETEC (reviewed in Ref. 11). The Shiga toxin (Stx) of EHEC cleaves ribosomal RNA, thereby disrupting protein synthesis and killing the intoxicated epithelial or endothelial cells12. The cytolethal distending toxin (CDT) has DNaseI activity that ultimately blocks cell division in the G2/M phase of the cell cycle13. Another toxin that blocks cell division in the same phase, called Cif (cycle-inhibiting factor), does not possess DNaseI activity, but might act by inhibition of Cdk1 kinase activity14. The cytotoxic nectrotizing factors (CNF 1 and CNF 2) deaminate a crucial glutamine residue of RhoA, Cdc42 and Rac, thereby locking these important signalling molecules in the 'on' position and leading to marked cytoskeletal alterations, multinucleation with cellular enlargement, and necrosis15. The Map protein of EPEC and EHEC has at least two independent activities — stimulating Cdc42-dependent filopodia formation and targeting mitochondria to disrupt membrane potential in these organelles16.
The various toxins are transported from the bacterial cytoplasm to the host cells by several mechanisms. LT is a classic A–B subunit toxin that is secreted to the extracellular milieu by a type II secretion system17. Several toxins, such as Sat, Pet and EspC, are called autotransporters because part of these proteins forms a β-barrel pore in the outer membrane that allows the other part of the protein extracellular access18. The SPATEs (serine protease autotransporters of enterobacteriaceae) are a subfamily of serine protease autotransporters that are produced by diarrhoeagenic and uropathogenic E. coli and Shigella strains. EPEC, EHEC and EIEC contain type III secretion systems, which are complex structures of more than 20 proteins forming a 'needle and syringe' apparatus that allows effector proteins, such as Tir and IpaB, to be injected directly into the host cell19. The UPEC haemolysin is the prototype of the type I secretion mechanism that uses TolC for export from the cell20. No type IV secretion systems have been described for pathogenic E. coli, with the exception of the type IV-like systems that are involved in conjugal transfer of some plasmids. By one means or another, pathogenic E. coli have evolved several mechanisms by which they can damage host cells and cause disease.
Pathotypes and pathogenesis
Enteropathogenic E. coli (EPEC). EPEC was the first pathotype of E. coli to be described. Large outbreaks of infant diarrhoea in the United Kingdom led Bray, in 1945, to describe a group of serologically distinct E. coli strains that were isolated from children with diarrhoea but not from healthy children. Although large outbreaks of infant diarrhoea due to EPEC have largely disappeared from industrialized countries, EPEC remains an important cause of potentially fatal infant diarrhoea in developing countries2. For decades, the mechanisms by which EPEC caused diarrhoea were unknown and this pathotype could only be identified on the basis of O:H serotyping. However, since 1979, numerous advances in our understanding of the pathogenesis of EPEC diarrhoea have been made, such that EPEC is now among the best understood of all the pathogenic E. coli.
A characteristic intestinal histopathology is associated with EPEC infections; known as 'attaching and effacing' (A/E), the bacteria intimately attach to intestinal epithelial cells and cause striking cytoskeletal changes, including the accumulation of polymerized actin directly beneath the adherent bacteria. The microvilli of the intestine are effaced and pedestal-like structures on which the bacteria perch frequently rise up from the epithelial cell (Fig. 3). The ability to induce this A/E histopathology is encoded by genes on a 35-kb pathogenicity island (PAI; see below) called the locus of enterocyte effacement (LEE)21. Homologues of LEE are also found in other human and animal pathogens that produce the A/E histopathology, including EHEC, rabbit EPEC (REPEC) and Citrobacter rodentium, which induces colonic hyperplasia in mice. The LEE encodes a 94-kDa outer-membrane protein called intimin, which mediates the intimate attachment of EPEC to epithelial cells22. Intimin not only functions as a ligand for epithelial cell adhesion, but also stimulates mucosal TH1 IMMUNE RESPONSES and intestinal crypt hyperplasia23. Most of the 41 open reading frames of the core LEE PAI encode a type III secretion system and the associated chaperones and effector proteins. One of these effector proteins, known as Tir (translocated intimin receptor), is inserted into the host-cell membrane, where it functions as a receptor for the intimin outer-membrane protein24. This is a fascinating example of a pathogen that provides its own receptor for binding to eukaryotic cells, although additional eukaryotic proteins have also been reported to act as receptors for intimin. A recent study showed that EPEC can disrupt cell polarity, causing basolateral membrane proteins, in particular β1-integrins, to migrate to the apical cell surface where they can bind to intimin25. In addition to β1-integrin, Tir has also been shown to bind to NUCLEOLIN26. In addition to its role as a receptor for intimin, Tir has important signalling functions in epithelial cells. The portion of Tir that is exposed to the cytosol nucleates cytoskeletal proteins, initially binding directly to the adaptor protein Nck, which recruits the amino terminus of Wiskott–Aldrich syndrome protein (N-WASP) and the actin-related protein 2/3 (Arp2/3) complex; recruitment of Arp2/3 results in actin filament nucleation and initiation of the characteristic pedestal complex27 (Fig. 1). Interestingly, the Tir protein of EHEC O157:H7 is not functionally identical to the Tir protein of EPEC O127:H6 because pedestals are formed independently of Nck, which indicates that additional bacterial factors are translocated to trigger actin signalling28. Other cytoskeletal proteins, such as vinculin, cortactin, talin and α-actinin, are also recruited to the pedestal complex29. Formation of the pedestal is a dynamic process whereby the force of actin polymerization can propel the pedestal across the surface of ptK2 epithelial cells30 (see movement of EPEC on ptK2 cells in the Online links). Tir also has a GAP (GTPase-activating protein) motif that has been implicated in the ability of Tir to downregulate filopodia formation16. Another secreted effector protein is EspF, which causes apoptosis31 and induces redistribution of the tight-junction-associated protein occludin, which leads to loss of trans-epithelial electrical resistance32. As noted above, the Map protein affects mitochondrial function and filopodia formation, and additional effectors — for example, EspG and EspH — have recently been described.
Additional EPEC virulence factors that are encoded outside the LEE have also been described. One very large protein of ∼385 kDa called lymphostatin (LifA) inhibits lymphocyte activation33. This protein is also present in strains of EHEC, where it is known as Efa1, and an adhesive property has been attributed to it34. Typical EPEC strains possess a plasmid of 70–100 kb called the EAF (EPEC adherence factor) plasmid35. This plasmid encodes a type IV pilus called the bundle-forming pilus (BFP)36, which mediates interbacterial adherence and possibly adherence to epithelial cells (Fig. 2). It also contains the per locus (plasmid-encoded regulator), the products of which regulate the bfp operon and most of the genes in the LEE by the LEE-encoded regulator (Ler). So-called atypical EPEC contain the LEE but do not contain the EAF plasmid. In industrialized countries, atypical EPEC are more frequently isolated from diarrhoeal cases than are typical EPEC that contain the EAF plasmid, although typical EPEC dominate in developing countries37. Atypical EPEC have also caused large outbreaks of diarrhoeal disease involving both children and adults in industrialized countries.
The model of EPEC pathogenesis is considerably more complex than simple binding to epithelial cells by a single adhesin and secretion of an enterotoxin that induces diarrhoea. The emerging model, several aspects of which are reviewed elsewhere2,38,39,40, indicates that EPEC initially adhere to epithelial cells by an adhesin, the identity of which is not yet clearly established; potential candidates include BFP, the EspA filament, flagella, LifA/Efa1 and intimin (by host-cell receptors). The type III secretion system is then activated and various effector proteins — including Tir, EspF, EspG, EspH and Map — are translocated into the host cell. EPEC binds through the interaction of intimin with Tir inserted in the membrane and numerous cytoskeletal proteins accumulate underneath the attached bacteria. Protein kinase C (PKC), phospholipase Cγ, myosin light-chain kinase and mitogen-activated protein (MAP) kinases are activated, which leads to several downstream effects, including increased permeability due to loosened tight junctions. Nuclear factor (NF)-κB is activated, leading to production of IL-8 and an inflammatory response that involves transmigration of polymorphonuclear leukocytes (PMNs) to the lumenal surface and activation of the adenosine receptor. The galanin-1 receptor is upregulated41, thereby increasing the response of the epithelial cells to the neuropeptide GALANIN, which is an important mediator of intestinal secretion. Some, but not all, typical EPEC strains produce an enterotoxin, EspC, that increases short circuit current in USSING CHAMBERS157. Diarrhoea probably results from multiple mechanisms, including active ion secretion, increased intestinal permeability, intestinal inflammation and loss of absorptive surface area resulting from microvillus effacement.
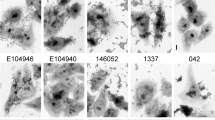
Enterohaemorrhagic E. coli (EHEC). First recognized as a cause of human disease in 1982, EHEC causes bloody diarrhoea (haemorrhagic colitis), non-bloody diarrhoea and haemolytic uremic syndrome (HUS). The principal reservoir of EHEC is the bovine intestinal tract and initial outbreaks were associated with consumption of undercooked hamburgers. Subsequently, a wide variety of food items have been associated with disease, including sausages, unpasteurized milk, lettuce, cantaloupe melon, apple juice and radish sprouts — the latter were responsible for an outbreak of 8,000 cases in Japan. Facilitated by the extremely low infectious dose required for infection (estimated to be <100 cells), EHEC has also caused numerous outbreaks associated with recreational and municipal drinking water, person-to-person transmission and petting zoo and farm visitations. A recent report indicates potential airborne transmission after exposure to a contaminated building42. EHEC strains of the O157:H7 serotype are the most important EHEC pathogens in North America, the United Kingdom and Japan, but several other serotypes, particularly those of the O26 and O111 serogroups, can also cause disease and are more prominent than O157:H7 in many countries.
The key virulence factor for EHEC is Stx, which is also known as verocytotoxin (VT). Stx consists of five identical B subunits that are responsible for binding the holotoxin to the glycolipid globotriaosylceramide (Gb3) on the target cell surface, and a single A subunit that cleaves ribosomal RNA, causing protein synthesis to cease12. The Stx family contains two subgroups — Stx1 and Stx2 — that share approximately 55% amino acid homology. Stx is produced in the colon and travels by the bloodstream to the kidney, where it damages renal endothelial cells and occludes the microvasculature through a combination of direct toxicity and induction of local cytokine and chemokine production, resulting in renal inflammation (reviewed in Ref. 43). This damage can lead to HUS, which is characterized by haemolytic anaemia, thrombocytopoenia and potentially fatal acute renal failure. Stx also induces apoptosis in intestinal epithelial cells — a process that is regulated by the Bcl-2 family44. Stx was first purified from Shigella dysenteriae, and HUS can also result from infection with this species, although not with other Shigella species or EIEC, which do not produce Stx. Stx also mediates local damage in the colon, which results in bloody diarrhoea, haemorrhagic colitis, necrosis and intestinal perforation.
In addition to Stx, most EHEC strains also contain the LEE pathogenicity island that encodes a type III secretion system and effector proteins that are homologous to those that are produced by EPEC. Animal models have shown the importance of the intimin adhesin in intestinal colonization, and HUS patients develop a strong antibody response to intimin and other LEE-encoded proteins. EHEC O157:H7 is believed to have evolved from LEE-containing O55 EPEC strains that acquired bacteriophage encoding Stx45. Although more than 200 serotypes of E. coli can produce Stx, most of these serotypes do not contain the LEE pathogenicity island and are not associated with human disease. This has led to the use of Shiga toxin-producing E. coli (STEC) or verotoxin-producing E. coli (VTEC) as general terms for any E. coli strain that produces Stx, and the term EHEC is used to denote only the subset of Stx-positive strains that also contain the LEE. However, there are LEE-negative STEC strains that are associated with disease — for example, O103:H21 strains — thereby demonstrating that there are additional virulence factors yet to be characterized. Several other potential adherence factors have been described for O157:H7 and/or non-O157:H7 strains, although the significance of these factors in human disease is not as well established as intimin. One potential adhesin is a large 362-kDa protein (ToxB) encoded on the 93-kb plasmid that is present in O157:H7 and other EHEC strains46. This protein shares sequence similarity with the large Clostridium toxin family, and to the EPEC LifA protein33 and the Efa-1 protein that has been implicated as an adhesin in non-O157:H7 EHEC strains34. This plasmid (pO157)47, also encodes an RTX (repeats in toxin) toxin that is similar to the UPEC haemolysin, a serine protease (EspP), a catalase and the StcE protein. StcE cleaves the C1 esterase inhibitor (C1-INH) of the complement pathway and could potentially contribute to the tissue damage, intestinal oedema and thrombotic abnormalities that are seen in EHEC infections48. The genome sequence of O157:H7 revealed numerous chromosomal islands (see below) that encode additional potential virulence factors. Included among these potential factors are novel fimbriae, iron uptake and utilization systems49, and a urease that is similar to those produced by Klebsiella and other urinary tract pathogens50.
Enterotoxigenic E. coli (ETEC). ETEC causes watery diarrhoea, which can range from mild, self-limiting disease to severe purging disease. The organism is an important cause of childhood diarrhoea in the developing world and is the main cause of diarrhoea in travellers to developing countries2.
ETEC colonizes the surface of the small bowel mucosa and elaborates enterotoxins, which give rise to intestinal secretion. Colonization is mediated by one or more proteinaceous fimbrial or fibrillar colonization factors (CFs), which are designated by CFA (colonization factor antigen), CS (coli surface antigen) or PCF (putative colonization factor) followed by a number. More than 20 antigenically diverse CFs have been characterized, yet epidemiological studies indicate that approximately 75% of human ETEC express either CFA/I, CFA/II or CFA/IV51. Antibodies to CFAs might ameliorate ETEC colonization and disease. ETEC are also an important cause of diarrhoeal disease in animals and these animal strains express fimbrial intestinal colonization factors, such as K88 and K99, which are not found in human ETEC strains.
ETEC enterotoxins belong to one of two groups: the heat-labile enterotoxins (LTs) and the heat-stable enterotoxins (STs). ETEC strains might express only an LT, only an ST, or both LTs and STs. LTs are a class of enterotoxins that are closely related in structure and function to cholera enterotoxin (CT), which is expressed by Vibrio cholerae52. The LT that is found predominantly in human isolates (LT-I; a related protein called LT-II is found in some animal ETEC isolates) has ∼80% amino acid identity with CT and, like CT, consists of a single A subunit and five identical B subunits. The B subunits mediate binding of the holotoxin to the cell surface gangliosides GM1 and GD1b, and the A subunit is responsible for the enzymatic activity of the toxin. LT has ADP-ribosyl transferase activity and transfers an ADP-ribosyl moiety from NAD to the α-subunit of the stimulatory G protein — a regulatory protein of the basolateral membrane that regulates adenylate cyclase. The resulting permanent activation of adenylate cyclase leads to increased levels of intracellular cAMP, activation of cAMP-dependent kinases and the eventual activation of the main chloride channel of epithelial cells — the cystic fibrosis transmembrane conductance regulator (CTFR). The net result of CFTR phosphorylation is increased Cl− secretion from secretory crypt cells, which leads to diarrhoea (reviewed in Ref. 11). LT can also stimulate prostaglandin synthesis and stimulate the enteric nervous system; both of these activities can also lead to stimulation of secretion and inhibition of absorption11. LT is also a potent mucosal adjuvant independent of its toxic activity53 and has been incorporated into numerous vaccine candidates containing a variety of antigens, resulting in increased antibody responses to these antigens when they are delivered orally, nasally or even transdermally.
STs are small, single-peptide toxins that include two unrelated classes — STa and STb — which differ in both structure and mechanism of action. Only toxins of the STa class have been associated with human disease2. The mature STa toxin is a ∼2-kDa peptide, which contains 18 or 19 amino acid residues, six of which are cysteines that form three intramolecular disulphide bridges (reviewed in Ref. 11). The main receptor for STa is a membrane-spanning guanylate cyclase; binding of STa to guanylate cyclase stimulates guanylate cyclase activity, leading to increased intracellular cGMP, which, in turn, activates cGMP-dependent and/or cAMP-dependent kinases and, ultimately, increases secretion. Interestingly, intestinal guanylate cyclase is the receptor for an endogenous ligand called guanylin54, which has a similar structure to that of STa. So the ST family seems to represent a case of molecular mimicry. The STb toxin is associated with animal disease and is a 48-amino-acid peptide containing two disulphide bonds (reviewed in Ref. 55). STb can elevate cytosolic Ca2+ concentrations, stimulate the release of prostaglandin E2 and stimulate the release of serotonin, all of which are mechanisms that could lead to increased ion secretion.
ETEC is largely a pathogen of developing countries, and it is well known that these countries typically have a low rate of colon cancer. Pitari et al.56 have reported that STa suppresses colon cancer cell proliferation through a guanylyl cyclase C-mediated signalling cascade. So the high prevalence of ETEC in developing countries might have a protective effect against this important disease, and indicates that infectious diseases might exist in a complex evolutionary balance with their human populations.
Enteroaggregative E. coli (EAEC). EAEC are increasingly recognized as a cause of often persistent diarrhoea in children and adults in both developing and developed countries, and have been identified as the cause of several outbreaks worldwide. At present, EAEC are defined as E. coli that do not secrete LT or ST and that adhere to HEp-2 cells in a pattern known as auto-aggregative, in which bacteria adhere to each other in a 'stacked-brick' configuration2. It is likely that this definition encompasses both pathogenic and non-pathogenic clones, and it remains controversial as to whether all the EAEC have any common factors that contribute to their shared adherence phenotype. Nevertheless, at least a subset of EAEC are proven human pathogens.
The basic strategy of EAEC infection seems to comprise colonization of the intestinal mucosa, probably predominantly that of the colon, followed by secretion of enterotoxins and cytotoxins57. Studies on human intestinal explants indicate that EAEC induces mild, but significant, mucosal damage58 — these effects are most severe in colonic sections. Mild inflammatory changes are observed in animal models59 and evidence indicates that at least some EAEC strains might be capable of limited invasion of the mucosal surface60,61. The most dramatic histopathological finding in infected animal models is the presence of a thick layer of auto-aggregating bacteria adhering loosely to the mucosal surface. EAEC prototype strains adhere to HEp-2 cells and intestinal mucosa by virtue of fimbrial structures known as aggregative adherence fimbriae (AAFs)62,63,64, which are related to the Dr family of adhesins. At least four allelic variants of AAFs exist, but importantly, each is present in only a minority of strains. It should be noted, however, that not all EAEC strains adhere by virtue of AAFs. A recently described protein called dispersin65 forms a loosely associated layer on the surface of EAEC strains and seems to counter the strong aggregating effects of the AAF adhesin, perhaps facilitating spread across the mucosal surface or penetration of the mucous layer. An additional surface structure that is potentially involved in causing inflammation is a novel EAEC flagellin protein that induces IL-8 release66. Release of this cytokine can stimulate neutrophil transmigration across the epithelium, which can itself lead to tissue disruption and fluid secretion.
Several toxins have been described for EAEC. Two such toxins are encoded by the same chromosomal locus on opposite strands. The larger gene encodes an autotransporter protease with mucinase activity called Pic; the opposite strand encodes the oligomeric enterotoxin that is known as Shigella enterotoxin 1 (ShET1), owing to its presence in most strains of Shigella flexneri 2a 67,68. The mode of action of ShET1 is not yet understood, but it might contribute to the secretory diarrhoea that accompanies EAEC and Shigella infection. A second enterotoxin that is present in many EAEC strains is enteroaggregative E. coli ST (EAST1), a 38-amino-acid homologue of the ETEC STa toxin69. It is conceivable that EAST1 could contribute to watery diarrhoea in EAST1-positive strains; however, the EAST1 gene (astA) can also be found in many commensal E. coli isolates, and therefore the role of EAST1 in diarrhoea remains an open question70. Many EAEC strains secrete an autotransporter toxin called Pet, which is encoded on the large virulence plasmid in close proximity to the gene encoding the AAF. Pet has enterotoxic activity and can also potentially lead to cytoskeletal changes and epithelial-cell rounding by cleavage of the cytoskeletal protein spectrin71.
Although no single virulence factor has been irrefutably associated with EAEC virulence, epidemiological studies implicate a 'package' of plasmid-borne and chromosomal virulence factors, similar to the virulence factors of other enteric pathogens. Several EAEC virulence factors are regulated by a single transcriptional activator called AggR, which is a member of the AraC family of transcriptional activators64 (J.P.N., unpublished data). One consistent observation from studies involving EAEC epidemiology is the association of the AggR regulon with diarrhoeal disease. Jiang et al. have recently shown that the presence of genes associated with the AggR regulon is predictive of significantly increased concentrations of faecal IL-8 and IL-1 in patients with diarrhoea caused by EAEC72. We suggest that the term 'typical EAEC' should be reserved for strains carrying AggR and at least a subset of AggR-regulated genes (for which the traditional EAEC probe is an adequate marker), and that the term 'atypical EAEC' be used for strains lacking the AggR regulon.
Enteroinvasive E. coli (EIEC). EIEC are biochemically, genetically and pathogenically closely related to Shigella spp. Numerous studies have shown that Shigella and E. coli are taxonomically indistinguishable at the species level73,74, but, owing to the clinical significance of Shigella, a nomenclature distinction is still maintained. The four Shigella species that are responsible for human disease, S. dysenteriae, S. flexneri, Shigella sonnei and Shigella boydii, cause varying degrees of dysentery, which is characterized by fever, abdominal cramps and diarrhoea containing blood and mucous. EIEC might cause an invasive inflammatory colitis, and occasionally dysentery, but in most cases EIEC elicits watery diarrhoea that is indistinguishable from that due to infection by other E. coli pathogens2. EIEC are distinguished from Shigella by a few minor biochemical tests, but these pathotypes share essential virulence factors. EIEC infection is thought to represent an inflammatory colitis, although many patients seem to manifest secretory, small bowel syndrome. The early phase of EIEC/Shigella pathogenesis comprises epithelial cell penetration, followed by lysis of the endocytic vacuole, intracellular multiplication, directional movement through the cytoplasm and extension into adjacent epithelial cells (reviewed in Ref. 75). Movement within the cytoplasm is mediated by nucleation of cellular actin into a 'tail' that extends from one pole of the bacterium. In addition to invasion into and dissemination within epithelial cells, Shigella (and presumably EIEC) also induces apoptosis in infected macrophages76. Genes that are required to effect this complex pathogenetic scheme are present on a large virulence plasmid that is found in EIEC and all Shigella species. The sequence of the 213-kb virulence plasmid of S. flexneri (pWR100) indicates that this plasmid is a mosaic that includes genetic elements that were initially carried by four plasmids77. One-third of the plasmid is composed of insertion sequence (IS) elements, which are undoubtedly important in the evolution of the virulence plasmid. This plasmid encodes a type III secretion system (see below) and a 120-kDa outer-membrane protein called IcsA, which nucleates actin by the binding of N-WASP8,78. The growth of actin micofilaments at only one bacterial pole induces movement of the organism through the epithelial cell cytoplasm. This movement culminates in the formation of cellular protrusions that are engulfed by neighbouring cells, after which the process is repeated. Although EIEC are invasive, dissemination of the organism past the submucosa is rare.
Much of EIEC/Shigella pathogenesis seems to be the result of the multiple effects of its plasmid-borne type III secretion system. This type III secretion system secretes multiple proteins, such as IpaA, IpaB, IpaC and IpgD, which mediate epithelial signalling events, cytoskeletal rearrangements, cellular uptake, lysis of the endocytic vacuole and other actions (reviewed in Refs 79,80). The type III secretion system apparatus, which is encoded by mxi and spa genes, enables the insertion of a pore containing IpaB and IpaC proteins into host cell membranes. In addition to pore formation, IpaB has several functions, such as binding to the signalling protein CD44, thereby triggering cytoskeletal rearrangements and cell entry, and binding to the macrophage caspase 1, resulting in apoptosis and release of IL-1 from macrophages. IpaC induces actin polymerization, which leads to the formation of cell extensions by activating the GTPases Cdc42 and Rac. The actin polymerization activity resides in the carboxy terminus of IpaC, whereas the amino terminus of this protein is involved in lamellipodial extensions. Conversely, IpaA binds to vinculin and induces actin depolymerization, thereby helping to organize the extensions that are induced by IpaC into a structure that enables bacterial entry. The translocated effector protein IpgD is a potent inositol 4-phosphatase that helps to reorganize host-cell morphology by uncoupling the cellular plasma membrane from the actin cytoskeleton, which leads to membrane blebbing81. Although the extensively characterized type III secretion system is essential for the invasiveness characteristic of EIEC and Shigella species, additional virulence factors have been described, including the plasmid-encoded serine protease SepA, the chromosomally encoded aerobactin iron-acquisition system and other secreted proteases that are encoded by genes present on pathogenicity islands (see below).
Diffusely adherent E. coli (DAEC). DAEC are defined by the presence of a characteristic, diffuse pattern of adherence to HEp-2 cell monolayers. DAEC have been implicated as a cause of diarrhoea in several studies, particularly in children >12 months of age2,82. Approximately 75% of DAEC strains produce a fimbrial adhesin called F1845 or a related adhesin (Ref. 83; J.P.N., unpublished observations); F1845 belongs to the Dr family of adhesins, which use DAF, a cell-surface glycosylphosphatidylinositol-anchored protein, which normally protects cells from damage by the complement system, as the receptor84,85,86. DAEC strains induce a cytopathic effect that is characterized by the development of long cellular extensions, which wrap around the adherent bacteria (Fig. 1). This characteristic effect requires binding and clustering of the DAF receptor by Dr fimbriae85. All members of the Dr family (including UPEC as well as the DAEC strain C1845) elicit this effect83. Binding of Dr adhesins is accompanied by the activation of signal transduction cascades, including activation of PI-3 kinase86. Peiffer et al. have reported that infection of an intestinal cell line by strains of DAEC impairs the activities and reduces the abundance of brush-border-associated sucrase-isomaltase and dipeptidylpeptidase IV87. This effect is independent of the DAF-associated pathway described above, and therefore provides a feasible mechanism for DAEC-induced enteric disease and also indicates the presence of virulence factors in DAEC other than Dr adhesins. Tieng et al.7 have proposed that DAEC might induce expression of MICA by intestinal epithelial cells, indicating that DAEC infection could be pro-inflammatory; this effect could potentially be important in the induction of inflammatory bowel diseases.
Uropathogenic E. coli (UPEC). The urinary tract is among the most common sites of bacterial infection and E. coli is by far the most common infecting agent at this site. The subset of E. coli that causes uncomplicated cystitis and acute pyelonephritis is distinct from the commensal E. coli strains that comprise most of the E. coli populating the lower colon of humans. E. coli from a small number of O serogroups (six O groups cause 75% of UTIs) have phenotypes that are epidemiologically associated with cystitis and acute pyelonephritis in the normal urinary tract, which include expression of P fimbriae, haemolysin, aerobactin, serum resistance and encapsulation. Clonal groups and epidemic strains that are associated with UTIs have been identified88,89.
Although many UTI isolates seem to be clonal, there is no single phenotypic profile that causes UTIs. Specific adhesins, including P (Pap), type 1 and other fimbriae (such as F1C, S, M and Dr), seem to aid in colonization90,91. Several toxins are produced, including haemolysin, cytotoxic necrotizing factor and an autotransported protease known as Sat. These virulence factors are found in differing percentages among various subgroups of UPEC92. Uropathogenic strains possess large and small pathogenicity islands containing blocks of genes that are not found in the chromosome of faecal strains. Availability of the genome sequence of E. coli CFT073 (Ref. 93) and efforts by other investigators to identify virulence genes by SIGNATURE-TAGGED MUTAGENESIS94 and other methods have allowed the development of a model of pathogenesis for UPEC (Fig. 4).
It is likely that infection begins with the colonization of the bowel with a uropathogenic strain in addition to the commensal flora. This strain, by virtue of factors that are encoded in pathogenicity islands, is capable of infecting an immunocompetent host, as it colonizes the periurethral area and ascends the urethra to the bladder (Fig. 4). Between 4 and 24 hours after infection, the new environment in the bladder selects for the expression of type 1 fimbriae95, which have an important role early in the development of a UTI96. Type 1 fimbriated E. coli attach to mannose moieties of the uroplakin receptors that coat transitional epithelial cells97. Attachment triggers apoptosis and exfoliation; for at least one strain, invasion of the bladder epithelium is accompanied with formation of pod-like bulges on the bladder surface that contain bacteria encased in a polysaccharide-rich matrix surrounded by a shell of uroplakin98. It is argued that invaded epithelial cells containing a tightly packed bacterial 'biofilm' could act as a reservoir for recurrent infection97,98, and indeed, in some cases of recurrent infection, the same serotype is encountered. However, a number of studies have identified different serotypes as being responsible for the recurring infection, an observation that is not consistent with this hypothesis. Iron acquisition and the ability to grow in urine are also crucial for survival.
In strains that cause cystitis, type 1 fimbriae are continually expressed and the infection is confined to the bladder96. In pyelonephritis strains, the invertible element that controls type 1 fimbriae expression turns to the 'off' position and type 1 fimbriae are less well expressed95. It could be argued that this releases the E. coli strain from bladder epithelial cell receptors and allows the organism to ascend through the ureters to the kidneys, where the organism can attach by P fimbriae to digalactoside receptors that are expressed on the kidney epithelium99,100. At this stage, haemolysin could damage the renal epithelium101 and, together with other bacterial products including LPS, an acute inflammatory response recruits PMNs to the site. Haemolysin has also been shown to induce Ca2+ oscillations in renal epithelial cells, resulting in increased production of IL-6 and IL-8 (Ref. 102). Secretion of Sat, a vacuolating cytotoxin, damages glomeruli and is cytopathic for the surrounding epithelium103. In some cases, the barrier that is provided by the one-cell-thick proximal tubules can be breached and bacteria can penetrate the endothelial cell to enter the bloodstream, leading to bacteraemia.
Meningitis/sepsis-associated E. coli (MNEC). This E. coli pathotype is the most common cause of Gram-negative neonatal meningitis, with a case fatality rate of 15–40% and severe neurological defects in many of the survivors104,105. The incidence of infants with early-onset sepsis owing to E. coli infection seems to be increasing, while infection by Gram-positive organisms decreases106. As with E. coli pathotypes that have a well-defined genetic basis for virulence, strains that cause meningitis are represented by only a limited number of O serogroups, and 80% of the strains are of the K1 capsule type. One interesting difference between MNEC and E. coli that cause intestinal or urinary tract infections is that although the latter strains can be readily transmitted by urine or faeces, infection of the central nervous system offers no obvious advantage for the selection and transmission of virulent MNEC strains.
E. coli that cause meningitis are spread haematogenously. Levels of bacteraemia correlate with the development of meningitis107; for example, bacteraemias of >103 colony forming units per ml of blood are significantly more likely to lead to the development of meningitis than in individuals with lower colony forming units per ml in their blood. These bacteria translocate from the blood to the central nervous system without apparent damage to the blood–brain barrier, which indicates a transcytosis process. Electron micrographs imply entry by a zippering mechanism in a process that does not affect transendothelial electrical resistance108. This indicates that the host-cell membrane is not significantly disrupted during entry of the bacterium. Two models for studying MNEC have been developed: a monolayer of brain microvascular endothelial cells109 and an intact animal model using 5-day-old rats110.
As for other E. coli pathotypes, the genomes of these extraintestinal K1 strains have additional genes that are not found in the commensal E. coli K-12 strains. In genomic comparisons, the genome of E. coli RS218, a meningitis-associated strain, was found to have at least 500 kb of additional genes inserted in at least 12 loci compared with E. coli K-12 (Refs 111,112). In addition, strain RS218 harbours a 100-kb plasmid, on which at least one virulence factor has been localized113.
Some insights into the mechanism of pathogenesis of these strains have been obtained. K1 strains use S fimbriae to bind to the lumenal surfaces of brain microvascular endothelium in neonatal rats114. Invasion requires the outer-membrane protein OmpA to bind to the GlcNAcβ1-4GlcNAc epitope of the brain microvascular endothelial cell receptor glycoprotein115. Other membrane proteins — for example, IbeA, IbeB, IbeC and AslA — are also required for invasion (reviewed in Ref. 116). Invasion correlates with microaerobic growth and iron supplementation117. CNF1 is required for invasion113, as is the K1 capsule, which elicits serum resistance and has antiphagocytic properties. In an experimental model, strains that express K1 capsule proteins and those that do not were able to cross the blood–brain barrier, but only the K1-expressing strains survived118. As a consequence of invasion, actin cytoskeletal rearrangement occurs and tyrosine phosphorylation of focal adhesion kinase (FAK) and paxillin is induced119. In addition, a substantial list of in vivo-induced genes, including those that encode iron-acquisition systems, was compiled using in vivo expression technology (IVET) in conjunction with a murine model of septicaemic infection120.
Other potential E. coli pathotypes. Several other potential E. coli pathotypes have been described, but none of these are as well established as the pathotypes described above (Box 1). Among the most intriguing of these potential pathogens are strains of E. coli that are associated with Crohn's Disease, which are known as adherent-invasive E. coli (AIEC)121. No unique genetic sequences have yet been described for AIEC strains, but such strains can invade and replicate within macrophages without inducing host-cell death and can induce the release of high amounts of tumour-necrosis factor (TNF)-α, a characteristic which could lead to the intestinal inflammation that is characteristic of Crohn's Disease. An inflammatory process, together with necrosis of the intestinal epithelium, are characteristics of necrotizing enterocolitis (NEC), an important cause of mortality and long-term morbidity in pre-term infants. The ability of some E. coli strains to transcytose through epithelial cell monolayers has been hypothesized to contribute to NEC122. Necrotoxic E. coli (NTEC) produce either CNF1 or CNF2 and have been associated with disease in both humans and animals123. Strains that are known as cell-detaching E. coli (CDEC) have been isolated from children with diarrhoea and the characteristic ability of these strains to detach cultured epithelial cells from glass or plastic has been associated with the production of haemolysin124. The relationships among the NEC-associated strains, NTEC and CDEC, have not yet been clearly established. The genes encoding CDT are infrequently present in E. coli strains and no significant association with disease has yet been found for this toxin. CDT is usually found in strains that possess other virulence factors, such as CNF, Stx and the LEE. However, recent information indicates that CDT can be encoded by four distinct genetic variants in E. coli and so earlier epidemiological studies using only one or two cdt genes as probes should be re-evaluated125. In at least one strain, the cdt genes are contained on a bacteriophage126, which could account for the presence of this toxin in a number of different E. coli pathotypes.
A poorly characterized subset of E. coli infections outside the gastrointestinal or urinary tract is a group implicated in intra-abdominal infections (IAIs), including abscesses, wounds, appendicitis and peritonitis. The initial microflora at the site of an IAI is polymicrobial, but E. coli and the strictly anaerobic Bacteroides fragilis are often isolated from these abscesses. A recent study indicates that a novel haem-binding protein, known as the 'haemoglobin-binding protease' (Hbp), is significantly associated with E. coli strains isolated from IAIs compared with those E. coli strains isolated from blood, urine or faeces127. Purified Hbp was shown to be capable of delivering haem to B. fragilis, indicating a synergy in abscess formation whereby E. coli provides iron from haem to B. fragilis to overcome iron restrictions imposed by the host. Interestingly, Hbp is identical to Tsh, which is an autotransporter haemagglutinin that is associated with APEC, thereby indicating that this protein can contribute to at least two different infectious diseases — IAIs in humans and respiratory tract infections in poultry127.
Genetics
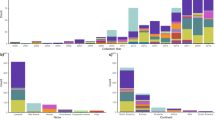
Mobile genetic elements. A striking feature of pathogenic E. coli is the association of genes that encode virulence factors with mobile genetic elements (Fig. 5). This was first shown more than 30 years ago with ETEC strains, in which enterotoxic activity was transferred together with a self-transmissible plasmid. In many cases, these 'Ent' plasmids were also shown to encode antibiotic resistance. There are now numerous examples of plasmids that encode crucial virulence factors of pathogenic E. coli, including plasmids in EAEC that encode fimbriae and toxins, plasmids in EIEC/Shigella that encode a type III secretion system and invasion factors, the EPEC EAF plasmid, which encodes BFP, and the pO157 plasmid of EHEC, which encodes accessory toxins. Although many of these plasmids are self-transmissible, some lack conjugation genes and can only be transferred with a conjugative plasmid. For ETEC, the genes that encode both LT and ST are found on plasmids, but some estA genes encoding STa are on transposons that can be inserted into either plasmids or the chromosome. One IS element has been described that contains the astA gene encoding the EAST1 toxin, completely embedded in a large putative transposase gene, the coding sequence of which is on the same strand but in the −1 reading frame relative to astA128.
E. coli virulence factors can be encoded by several mobile genetic elements, including transposons (Tn) (for example, heat stable enterotoxin (ST) of ETEC), plasmids (for example, heat-labile enterotoxin (LT) of ETEC and invasion factors of EIEC), bacteriophage (for example, Shiga toxin of EHEC) and pathogenicity islands (PAIs) — for example, the locus of enterocyte effacement (LEE) of EPEC/EHEC and PAIs I and II of UPEC. Commensal E. coli can also undergo deletions resulting in 'black holes', point mutations or other DNA rearrangements that can contribute to virulence. These additions, deletions and other genetic changes can give rise to pathogenic E. coli forms capable of causing diarrhoea (EPEC, EHEC, EAEC DAEC), dysentery (EIEC), haemolytic uremic syndrome (EHEC), urinary tract infections (UPEC) and meningitis (MNEC). HUS, haemolytic uremic syndrome; UTI, urinary tract infection.
The main virulence factor of EHEC, Stx, is encoded on a lambda-like bacteriophage; acquisition of this phage was a key step in the evolution of EHEC from EPEC45. The EHEC EDL933 genome sequence contains 18 regions with homology to known bacteriophages, but most seem to be incomplete phage genomes49. Although only the Stx phage seems to be capable of lytic growth and production of infectious particles, these cryptic phage sequences enable the continued evolution of these strains by homologous recombination of phages into different chromosomal sites. The ability to produce Stx can be readily transmitted by transduction of the genes encoding Stx phage to K-12 or commensal E. coli, but this step is probably insufficient to confer virulence because non-O157:H7 E. coli strains containing stx genes without other EHEC virulence factor genes can be readily isolated from commercial meat products. This observation reinforces the concept that a single gene is insufficient to convert commensal E. coli to pathogenic E. coli, and that instead a combination of genes encoding toxins, colonization factors and other functions are required to make E. coli pathogenic.
PAIs are large genomic regions (10–200 kb) that are present in the genomes of pathogenic strains but absent from the genomes of non-pathogenic members of the same or related species (reviewed in Ref. 129). PAIs are typically associated with tRNA genes, have a different G+C content compared with the host DNA and often carry cryptic or functional genes that encode mobility factors, such as integrases, transposases and IS elements. PAIs were first described in pathogenic E. coli and have subsequently been described in several Gram-negative and Gram-positive bacteria. The first PAIs were described in UPEC strain 536, which contains at least four such islands130. The PAI II536 island is 100 kb in size, is inserted at the leuX tRNA gene at minute 97 on the E. coli chromosome and encodes haemolysin and P fimbriae. This island is flanked by 18-bp direct repeats, which facilitate deletion of the entire island at a relatively high frequency.
The first PAI to be described in diarrhoeagenic E. coli was the LEE PAI in EPEC and EHEC21. As described above, the LEE encodes a type III secretion system and other factors that are responsible for the A/E histopathology. In EPEC strain E2348/69 and EHEC strain O157:H7, the LEE is inserted at the selC tRNA gene, which is also the site of insertion of the PAI I536 island of UPEC. The insertion of two different PAIs at the same chromosomal site in EPEC/EHEC and UPEC indicates the presence of 'hot spots' in the E. coli chromosome into which different PAIs can insert and give rise to different E. coli pathotypes. The 35-kb LEE from E2348/69 contains 41 open reading frames that are highly conserved among EPEC and EHEC strains, as well as rabbit and other animal strains of EPEC that produce A/E lesions. In some E. coli strains, the LEE PAI is immediately adjacent to genes that encode other potential virulence factors, such as the efa1/lifA gene, to form a larger PAI of 59.5 kb131. The LEE of one rabbit strain is contained on a ∼85-kb PAI that contains an intact integrase gene and is flanked by direct repeats. This PAI is capable of spontaneous deletion and site-specific integration into the pheU tRNA locus of K-12 (Ref. 131). The prototypic LEE of E2348/69 contains no direct repeats or mobility genes and seems to be incapable of spontaneous deletion or transfer, which indicates that this PAI has evolved to the point that it has lost the genetic elements that were responsible for the initial integration into the chromosome.
PAIs have also been described for EAEC, EIEC/Shigella, MNEC and some ETEC strains (reviewed in Refs 132–134). Some PAIs are unique to individual pathotypes, whereas other PAIs are found in multiple pathotypes. The she (Shi-I) PAI is present in EAEC, where it encodes the ShET1 enterotoxin and the autotransporter toxin Pic. The high pathogenicity island (HPI) was originally described in Yersinia, but is also present in most strains of EAEC, DAEC and UPEC, and in some strains of EIEC, ETEC, EPEC and EHEC, as well as some Klebsiella and Citrobacter strains135. The HPI contains genes that are involved in regulation, biosynthesis and uptake of the siderophore yersiniabactin.
The inverse of PAIs are 'black holes', which refers to the deletion of blocks of genes in commensal or K-12 E. coli that lead to increased virulence. In EIEC/Shigella, lack of the cadA gene, which encodes lysine decarboxylase (LDC) in K-12, enables activity of an enterotoxin which is normally inhibited by the product of the LDC reaction — cadaverine136. In many EIEC strains, the cadC gene that encodes a regulator of cadA is preferentially mutated, which results in the same phenotype137. EIEC/Shigella also have a large number of pseudogenes (see below), which might also comprise functional 'black holes'. Although the genes encoding E. coli virulence factors are usually either present or absent, single-nucleotide polymorphisms (SNPs) that contribute to virulence have been found in the genes that encode the FimH and Dr adhesins138.
Genomic sequences. Prior to the determination of the complete genomic sequence for a pathogenic strain of E. coli it was anticipated that these pathogens differed from K-12 primarily by the presence of a limited number of PAIs, plasmids and phage that encoded specific virulence factors. However, when the first pathotype was sequenced — namely two different strains of EHEC O157:H7 — the extent of lateral gene transfer was found to be far greater than had been anticipated. EHEC strain EDL933 contains nearly 1,400 novel genes scattered throughout 177 discrete regions of DNA greater than 50 bp in size called O-islands; these regions total 1.34 Mb of DNA that is not present in K-12 (Ref. 49). Almost as surprising was the fact that although the two strains shared a 4.1-Mb 'backbone' of common sequences, EDL933 lacked 0.53 Mb of DNA that was present in K-12 in 234 'K-islands' (>50 bp). The absence of a substantial amount of K-12 DNA in other E. coli pathotypes was shown in a recent DNA array study in which up to 10% of E. coli K-12 open reading frames were not detected in several pathogenic and non-pathogenic E. coli strains139.
The striking mosaic structure of EHEC was further shown by the determination of the UPEC genome sequence, which at 5.2 Mb is similar in size to that of EHEC93. UPEC strain CFT073 contains 2,004 genes in 247 islands that are not present in K-12. In contrast to the striking conservation of the core LEE PAI in EPEC and EHEC, substantial differences were seen between the large PAIs of CFT073 and two other well-studied UPEC strains — J96 and 536. The analyses indicated that extraintestinal pathogenic E. coli strains arose independently from multiple clonal lineages. Interestingly, when the predicted proteins from all three strains, K-12, EHEC and UPEC, were compared, only 39.2% of the combined (nonredundant) set of proteins are common to all three strains93.
As noted above, several studies using DNA hybridization, multilocus enzyme electrophoresis and sequencing of a small number of genes indicates that Shigella species clearly fall taxonomically within the E. coli species74. The genome sequence of S. flexneri 2a further supports this grouping and exhibits the backbone and island mosaic structure of the genomes of the E. coli pathogens73. The 4.599-Mb genome size is closer to that of K-12 (4.639 Mb) than to EHEC and UPEC, and the 70.6% of K-12 genes that are found in S. flexneri is of a similar magnitude to the 74.3% of K-12 genes that are found in UPEC CFT073. However, UPEC contains an additional 1,827 proteins that are not found in K-12, whereas S. flexneri contains only 205 proteins that are not found in K-12, thereby indicating that S. flexneri is more similar to K-12 than is UPEC CFT073. The S. flexneri genome is notable for its large number of IS elements — which constitute 6.7% (309.4 kb) of the chromosome — and for the large number (372) of pseudogenes present — which constitute 8.1% of the genome. These pseudogenes arose by several mechanisms, including single-nucleotide insertions or deletions, point mutations and IS-element insertions. Interestingly, phenotypic tests that have traditionally been used to distinguish E. coli from S. flexneri, such as lack of motility, utilization of various carbon sources and the requirement for NAD, are largely the result of pseudogenes. Whether these pseudogenes are advantageous, disadvantageous, or neutral cannot be stated at this time.
Regulation. Consistent with the fact that E. coli virulence factors are typically encoded on 'foreign' DNA that is not contained in commensal E. coli strains, the expression of many virulence factor genes is frequently regulated by transcriptional regulators that are also encoded on pathogenicity islands or plasmids. One such pathogen-specific regulator is the LEE-encoded Ler protein, which positively regulates the EPEC/EHEC genes encoding the type III secretion system that are also found on the LEE140. Another example is the PapB regulator of the pap operon encoded on PAIs in UPEC141. In some instances, a plasmid-encoded regulator can activate transcription of chromosomal genes — for example, regulators such as the regulatory cascade formed by the EPEC plasmid-encoded regulator (Per) that regulates the LEE-encoded regulator, Ler (Fig. 6). Many pathogen-specific regulators belong to the AraC family of transcriptional activators, such as Per (EPEC), AggR (EAEC), VirF (EIEC) and Rns (ETEC).
The attaching and effacing histopathology induced by EPEC and EHEC is encoded by the locus of enterocyte effacement (LEE) pathogenicity island, which contains five major polycistronic operons designated LEE1–5. Expression of the LEE genes is regulated by EPEC-specific regulators (depicted in green) and generic E. coli regulators (depicted in yellow). The first open reading frame of the LEE1 operon encodes the LEE-encoded regulator, Ler, which positively regulates expression of other LEE operons by counteracting the repressive effects of H-NS140,148. Ler also regulates expression of the EspC enterotoxin that is produced by many EPEC strains and potentially other virulence factors. Expression of Ler is itself regulated by several factors, including IHF149, FIS150 and BipA151, and quorum sensing through the QseA regulator152. Quorum sensing also regulates other factors that are potentially involved in virulence, such as flagella, through the QseBC two-component regulator153. In EPEC, but not EHEC, expression of Ler is positively regulated by the products of the per (plasmid-encoded regulator)154 locus, which consists of three open reading frames, perA, perB and perC; PerA (BfpT) also regulates the bfp genes encoding a type IV pilus155. In acidic conditions, the per genes are repressed by GadX, which activates the gadAB genes involved in acid resistance156. This dual action of GadX could prevent premature expression of virulence factors in the stomach while enhancing survival of the organism until it reaches more alkaline conditions in the small intestine where expression of virulence factors is induced. Bip, Ig heavy chain binding protein; FIS, factor for inversion stimulation; IHF, integration host factor.
Expression of E. coli virulence factors is not solely regulated by pathogen-specific regulators. A common theme among the various E. coli pathotypes is the exploitation of regulators present in commensal E. coli for the regulation of virulence factor genes that are present only in pathogenic E. coli. For example, the stx1 gene encoding Shiga toxin is transcribed from the PR′ promoter that also controls expression of late lambda phage lysis genes, thereby linking toxin expression with a lytic function, which allows release of the toxin142. This linkage leads to induction of transcription of both toxin genes and lysis genes by certain antibiotics, causing increased toxin production, increased release of toxin by lysis and increased death in a mouse model143. Another example is the EPEC Ler, which in addition to being regulated by Per is also regulated by integrative host factor (IHF), factor for inversion stimulation (FIS) and Ig heavy chain binding protein (BipA) — global regulators of housekeeping genes in K-12 (Fig. 6). Another regulatory system present in K-12 that regulates expression of both housekeeping and virulence factor genes is the AI-2/luxS quorum sensing (QS) system. QS is a method of intercellular communication that allows unicellular organisms such as E. coli to behave as multi-cellular organisms. A small autoinducer (AI) molecule is produced by many organisms, including E. coli; AIs can activate the expression of a subset of genes when the microbial population, and therefore the AI concentration, reaches a crucial level. QS regulates the expression of the EPEC and EHEC LEE operons by Ler as well as flagella expression144. As the infectious dose of EHEC (10–100 organisms) is too low to make use of QS, a model has been proposed in which EHEC detect the AI signals that are produced by the large concentration of commensal E. coli and other bacteria present in the large intestine144. In response to this signal, expression of key virulence factors, including the LEE and Stx, is induced, thereby initiating the disease process. This regulatory mechanism can also be activated by mammalian hormones, such as adrenaline and noradrenaline, in an example of regulatory 'cross-talk' between eukaryotic and prokaryotic organisms145.
Regulation of virulence factor expression by physical DNA rearrangements is uncommon in pathogenic E. coli but phase variation is seen with type 1 fimbriae. Transcription of the fim operon that encodes type 1 fimbriae is primarily under the control of an invertible element that contains the promoter responsible for transcription of the main structural subunit. Individual bacterial cells either express the fimbriae over their entire surface or do not express any fimbriae. This phase variation of type 1 fimbriae is controlled at the transcriptional level by the invertible element, which is regulated by the FimB and FimE recombinases146. The inversion seems to be regulated during the course of infection, and the orientation of the element correlates with whether UPEC strains remain localized to the bladder. In cystitis infections most of the strains have the invertible element in the 'on' position and express type 1 fimbriae, whereas when they leave the bladder and ascend to the kidneys to cause pyelonephritis, most of the strains have the element in the 'off' position and do not express type 1 fimbriae95. The regulation of type 1 fimbriae in UPEC is further complicated by cross-talk between two different adhesion operons, whereby PapB, a key regulator of the pap operon, inhibits type 1 phase variation141.
Conclusions
The evolution of pathogenic E. coli that has resulted in formation of distinct pathotypes capable of colonizing the gastrointestinal tract, urinary tract or meninges illustrates how key genetic elements can adapt a strain to distinct host environments. Using E. coli K-12 as a 'base-model', several features can be added (PAIs, plasmids, transposons or phage) or subtracted (black holes or pseudogenes) to modify the base model to adapt to specific environments and to enable these modified strains to cause disease in an immunocompetent human or animal host. This genomic plasticity complicates efforts to categorize the various clusters of pathogenic E. coli strains into sharply delineated pathotypes. The evolutionary process, clearly ongoing, has resulted in a highly versatile species that is capable of colonizing, multiplying in and damaging diverse environments. The host cell activities that are affected by these pathogenic strains of E. coli encompass a broad spectrum of functions, including signal transduction, protein synthesis, mitochondrial function, cytoskeletal function, cell division, ion secretion, transcription and apoptosis. The ability of various E. coli virulence factors to affect such a wide range of cellular functions has led to the use of the various toxins, effectors and cell surface structures as tools to better understand these fundamental eukaryotic processes. Our increased understanding of the mechanisms by which E. coli can cause disease has dramatically changed our perspective of this species that was once dismissed as a harmless commensal of the intestinal tract.
References
Sweeney, N. J. et al. The Escherichia coli K-12 gntP gene allows E. coli F-18 to occupy a distinct nutritional niche in the streptomycin-treated mouse large intestine. Infect. Immun. 64, 3497–3503 (1996).
Nataro, J. P. & Kaper, J. B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 11, 142–201 (1998). A comprehensive review of the pathogenesis, epidemiology, diagnosis and clinical aspects of diarrhoeagenic E. coli.
Russo, T. A. & Johnson, J. R. Proposal for a new inclusive designation for extraintestinal pathogenic isolates of Escherichia coli: ExPEC. J. Infect. Dis. 181, 1753–1754 (2000).
Whittam, T. S. in Escherichia coli and Salmonella (eds Neidhardt, F. C. et al.) 2708–2720 (ASM Press, Washington DC, USA, 1996).
Cassels, F. J. & Wolf, M. K. Colonization factors of diarrheagenic E. coli and their intestinal receptors. J. Ind. Microbiol. 15, 214–226 (1995).
Keller, R. et al. Afa, a diffuse adherence fibrillar adhesin associated with enteropathogenic Escherichia coli. Infect. Immun. 70, 2681–2689 (2002).
Tieng, V. et al. Binding of Escherichia coli adhesin AfaE to CD55 triggers cell-surface expression of the MHC class I-related molecule MICA. Proc. Natl Acad. Sci. USA 99, 2977–2982 (2002).
Goldberg, M. B. & Theriot, J. A. Shigella flexneri surface protein IcsA is sufficient to direct actin-based motility. Proc. Natl Acad. Sci. USA 92, 6572–6576 (1995).
Tapping, R. I., Akashi, S., Miyake, K., Godowski, P. J. & Tobias, P. S. Toll-like receptor 4, but not toll-like receptor 2, is a signaling receptor for Escherichia and Salmonella lipopolysaccharides. J. Immunol. 165, 5780–5787 (2000).
Hayashi, F. et al. The innate immune response to bacterial flagellin is mediated by Toll-like receptor 5. Nature 410, 1099–1103 (2001).
Sears, C. L. & Kaper, J. B. Enteric bacterial toxins: mechanisms of action and linkage to intestinal secretion. Microbiol. Rev. 60, 167–215 (1996).
Melton-Celsa, A. R. & O'Brien, A. D. in Escherichia coli O157:H7 and Other Shiga Toxin-Producing E. coli Strains (eds Kaper, J. B. & O'Brien, A. D.) 121–128 (ASM Press, Washington DC, USA, 1998).
De Rycke, J. & Oswald, E. Cytolethal distending toxin (CDT): a bacterial weapon to control host cell proliferation? FEMS Microbiol Lett. 203, 141–148 (2001).
Marches, O. et al. Enteropathogenic and enterohaemorrhagic Eschericha coli deliver a novel effector called Cif, which blocks cell cycle G2/M transition. Mol. Microbiol 50, 1553–1567 (2003).
Lerm, M. et al. Deamidation of Cdc42 and Rac by Escherichia coli cytotoxic necrotizing factor 1: activation of c-Jun N-terminal kinase in HeLa cells. Infect. Immun. 67, 496–503 (1999).
Kenny, B. et al. Co-ordinate regulation of distinct host cell signalling pathways by multifunctional enteropathogenic Escherichia coli effector molecules. Mol. Microbiol. 44, 1095–1107 (2002).
Tauschek, M., Gorrell, R. J., Strugnell, R. A. & Robins-Browne, R. M. Identification of a protein secretory pathway for the secretion of heat-labile enterotoxin by an enterotoxigenic strain of Escherichia coli. Proc. Natl Acad. Sci. USA 99, 7066–7071 (2002).
Henderson, I. R., Navarro-Garcia, F. & Nataro, J. P. The great escape: structure and function of the autotransporter proteins. Trends Microbiol. 6, 370–378 (1998).
Hueck, C. J. Type III protein secretion systems in bacterial pathogens of animals and plants. Microbiol. Mol. Biol. Rev. 62, 379–433 (1998).
Balakrishnan, L., Hughes, C. & Koronakis, V. Substrate-triggered recruitment of the TolC channel-tunnel during type I export of hemolysin by Escherichia coli. J. Mol. Biol. 313, 501–510 (2001).
McDaniel, T. K., Jarvis, K. G., Donnenberg, M. S. & Kaper, J. B. A genetic locus of enterocyte effacement conserved among diverse enterobacterial pathogens. Proc. Natl Acad. Sci. USA 92, 1664–1668 (1995). The first description of a pathogenicity island in enteric E. coli pathotypes.
Jerse, A. E., Yu, J., Tall, B. D. & Kaper, J. B. A genetic locus of enteropathogenic Escherichia coli necessary for the production of attaching and effacing lesions on tissue culture cells. Proc. Natl Acad. Sci. USA 87, 7839–7843 (1990).
Higgins, L. M. et al. Role of bacterial intimin in colonic hyperplasia and inflammation. Science 285, 588–591 (1999).
Kenny, B. et al. Enteropathogenic E. coli (EPEC) transfers its receptor for intimate adherence into mammalian cells. Cell 91, 511–520 (1997). The first report of a bacterium translocating its receptor into mammalian cells by a type III secretion system.
Muza-Moons, M. M., Koutsouris, A. & Hecht, G. Disruption of cell polarity by enteropathogenic Escherichia coli enables basolateral membrane proteins to migrate apically and to potentiate physiological consequences. Infect. Immun. 71, 7069–7078 (2003).
Sinclair, J. F. & O'Brien, A. D. Cell surface-localized nucleolin is a eukaryotic receptor for the adhesin intimin-γ of enterohemorrhagic Escherichia coli O157:H7. J. Biol. Chem. 277, 2876–2885 (2002).
Kalman, D. et al. Enteropathogenic E. coli acts through WASP and Arp2/3 complex to form actin pedestals. Nature Cell Biol. 1, 389–391 (1999).
Campellone, K. G. & Leong, J. M. Tails of two Tirs: actin pedestal formation by enteropathogenic E. coli and enterohemorrhagic E. coli O157:H7. Curr. Opin. Microbiol. 6, 82–90 (2003).
Vallance, B. A. & Finlay, B. B. Exploitation of host cells by enteropathogenic Escherichia coli. Proc. Natl Acad. Sci. USA 97, 8799–8806 (2000).
Sanger, J. M., Chang, R., Ashton, F., Kaper, J. B. & Sanger, J. W. Novel form of actin-based motility transports bacteria on the surface of infected cells. Cell Motil. Cytoskeleton 34, 279–287 (1996).
Crane, J. K., McNamara, B. P. & Donnenberg, M. S. Role of EspF in host cell death induced by enteropathogenic Escherichia coli. Cell Microbiol. 3, 197–211 (2001).
McNamara, B. P. et al. Translocated EspF protein from enteropathogenic Escherichia coli disrupts host intestinal barrier function. J. Clin. Invest. 107, 621–629 (2001).
Klapproth, J. M. et al. A large toxin from pathogenic Escherichia coli strains that inhibits lymphocyte activation. Infect. Immun. 68, 2148–2155 (2000).
Nicholls, L., Grant, T. H. & Robins-Browne, R. M. Identification of a novel genetic locus that is required for in vitro adhesion of a clinical isolate of enterohaemorrhagic Escherichia coli to epithelial cells. Mol. Microbiol. 35, 275–288 (2000).
Tobe, T. et al. Complete DNA sequence and structural analysis of the enteropathogenic Escherichia coli adherence factor plasmid. Infect. Immun. 67, 5455–5462 (1999).
Girón, J. A., Ho, A. S. Y. & Schoolnik, G. K. An inducible bundle-forming pilus of enteropathogenic Escherichia coli. Science 254, 710–713 (1991).
Trabulsi, L. R., Keller, R. & Tardelli Gomes, T. A. Typical and atypical enteropathogenic Escherichia coli. Emerg. Infect. Dis. 8, 508–513 (2002).
Hecht, G. Microbes and microbial toxins: paradigms for microbial–mucosal interactions. VII. Enteropathogenic Escherichia coli: physiological alterations from an extracellular position. Am. J. Physiol Gastrointest. Liver Physiol. 281, G1–G7 (2001).
Frankel, G. et al. Enteropathogenic and enterohaemorrhagic Escherichia coli: more subversive elements. Mol. Microbiol. 30, 911–921 (1998).
Kenny, B. Mechanism of action of EPEC type III effector molecules. Int. J. Med. Microbiol. 291, 469–477 (2002).
Hecht, G. et al. Pathogenic Escherichia coli increase Cl− secretion from intestinal epithelia by upregulating galanin-1 receptor expression. J. Clin. Invest. 104, 253–262 (1999).
Varma, J. K. et al. An outbreak of Escherichia coli O157 infection following exposure to a contaminated building. JAMA 290, 2709–2712 (2003).
Andreoli, S. P., Trachtman, H., Acheson, D. W., Siegler, R. L. & Obrig, T. G. Hemolytic uremic syndrome: epidemiology, pathophysiology, and therapy. Pediatr. Nephrol. 17, 293–298 (2002).
Jones, N. L. et al. Escherichia coli Shiga toxins induce apoptosis in epithelial cells that is regulated by the Bcl-2 family. Am. J. Physiol. Gastrointest. Liver Physiol. 278, G811–G819 (2000).
Reid, S. D., Herbelin, C. J., Bumbaugh, A. C., Selander, R. K. & Whittam, T. S. Parallel evolution of virulence in pathogenic Escherichia coli. Nature 406, 64–67 (2000).
Tatsuno, I. et al. toxB gene on pO157 of enterohemorrhagic Escherichia coli O157:H7 is required for full epithelial cell adherence phenotype. Infect. Immun. 69, 6660–6669 (2001).
Burland, V. et al. The complete DNA sequence and analysis of the large virulence plasmid of Escherichia coli O157:H7. Nucleic Acids Res. 26, 4196–4204 (1998).
Lathem, W. W. et al. StcE, a metalloprotease secreted by Escherichia coli O157:H7, specifically cleaves C1 esterase inhibitor. Mol. Microbiol. 45, 277–288 (2002).
Perna, N. T. et al. Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature 409, 529–533 (2001). The first reported genome sequence for a pathogenic E. coli strain.
Heimer, S. R. et al. Urease of enterohemorrhagic Escherichia coli: evidence for regulation by fur and a trans-acting factor. Infect. Immun. 70, 1027–1031 (2002).
Wolf, M. K. Occurrence, distribution, and associations of O and H serogroups, colonization factor antigens, and toxins of enterotoxigenic Escherichia coli. Clin. Microbiol. Rev. 10, 569–584 (1997).
Spangler, B. D. Structure and function of cholera toxin and the related Escherichia coli heat-labile enterotoxin. Microbiol. Rev. 56, 622–647 (1992).
Pizza, M. et al. Mucosal vaccines: non-toxic derivatives of LT and CT as mucosal adjuvants. Vaccine 19, 2534–2541 (2001).
Currie, M. G. et al. Guanylin: an endogenous activator of intestinal guanylate cyclase. Proc. Natl Acad. Sci. USA 89, 947–951 (1992). This paper suggests that STa evolved as a molecular mimic of an endogenous ligand. This model is necessary, not only to understand ETEC pathogenesis and evolution, but also to provide a context for future studies of microbial evolution.
Dubreuil, J. D. Escherichia coli STb enterotoxin. Microbiology 143, 1783–1795 (1997).
Pitari, G. M. et al. Bacterial enterotoxins are associated with resistance to colon cancer. Proc. Natl Acad. Sci. USA 100, 2695–2699 (2003).
Nataro, J. P., Steiner, T. S. & Guerrant, R. L. Enteroaggregative Escherichia coli. Emerg. Infect. Dis. 4, 251–261 (1998).
Hicks, S., Candy, D. C. A. & Phillips, A. D. Adhesion of enteroaggregative Escherichia coli to pediatric intestinal mucosa in vitro. Infect. Immun. 64, 4751–4760 (1996).
Vial, P. A. et al. Characterization of enteroadherent-aggregative Escherichia coli, a putative agent of diarrheal disease. J. Infect. Dis. 158, 70–79 (1988).
Benjamin, P., Federman, M. & Wanke, C. A. Characterization of an invasive phenotype associated with enteroaggregative Escherichia coli. Infect. Immun. 63, 3417–3421 (1995).
Abe, C. M., Knutton, S., Pedroso, M. Z., Freymuller, E. & Gomes, T. A. An enteroaggregative Escherichia coli strain of serotype O111:H12 damages and invades cultured T84 cells and human colonic mucosa. FEMS Microbiol. Lett. 203, 199–205 (2001).
Czeczulin, J. R. et al. Aggregative adherence fimbria II, a second fimbrial antigen mediating aggregative adherence in enteroaggregative Escherichia coli. Infect. Immun. 65, 4135–4145 (1997).
Nataro, J. P. et al. Aggregative adherence fimbriae I of enteroaggregative Escherichia coli mediate adherence to HEp-2 cells and hemagglutination of human erythrocytes. Infect. Immun. 60, 2297–2304 (1992).
Nataro, J. P., Yikang, D., Yingkang, D. & Walker, K. AggR, a transcriptional activator of aggregative adherence fimbria I expression in enteroaggregative Escherichia coli. J. Bacteriol. 176, 4691–4699 (1994). Describes the emergence of AggR as a global regulator of virulence genes in EAEC
Sheikh, J. et al. A novel dispersin protein in enteroaggregative Escherichia coli. J. Clin. Invest. 110, 1329–1337 (2002).
Steiner, T. S., Nataro, J. P., Poteet-Smith, C. E., Smith, J. A. & Guerrant, R. L. Enteroaggregative Escherichia coli expresses a novel flagellin that causes IL-8 release from intestinal epithelial cells. J. Clin. Invest 105, 1769–1777 (2000). The pathogenesis of EAEC is not completely understood, but inflammation might be an important component.
Henderson, I. R., Czeczulin, J., Eslava, C., Noriega, F. & Nataro, J. P. Characterization of Pic, a secreted protease of Shigella flexneri and enteroaggregative Escherichia coli. Infect. Immun. 67, 5587–5596 (1999).
Noriega, F. R., Liao, F. M., Formal, S. B., Fasano, A. & Levine, M. M. Prevalence of Shigella enterotoxin 1 among Shigella clinical isolates of diverse serotypes. J. Infect. Dis. 172, 1408–1410 (1995).
Savarino, S. J. et al. Enteroaggregative Escherichia coli heat-stable enterotoxin 1 represents another subfamily of E. coli heat-stable toxin. Proc. Natl Acad. Sci. USA 90, 3093–3097 (1993).
Menard, L. P. & Dubreuil, J. D. Enteroaggregative Escherichia coli heat-stable enterotoxin 1 (EAST1): a new toxin with an old twist. Crit. Rev. Microbiol. 28, 43–60 (2002).
Navarro-Garcia, F. et al. In vitro effects of a high-molecular weight heat-labile enterotoxin from enteroaggregative Escherichia coli. Infect. Immun. 66, 3149–3154 (1998).
Jiang, Z. D., Greenberg, D., Nataro, J. P., Steffen, R. & DuPont, H. L. Rate of occurrence and pathogenic effect of enteroaggregative Escherichia coli virulence factors in international travelers. J. Clin. Microbiol. 40, 4185–4190 (2002).
Wei, J. et al. Complete genome sequence and comparative genomics of Shigella flexneri serotype 2a strain 2457T. Infect. Immun. 71, 2775–2786 (2003).
Pupo, G. M., Lan, R. & Reeves, P. R. Multiple independent origins of Shigella clones of Escherichia coli and convergent evolution of many of their characteristics. Proc. Natl Acad. Sci. USA 97, 10567–10572 (2000). This paper suggests that Shigella should be considered within the species Escherichia coli , and that their evolution represents adaptation to a specific pathogenetic niche, a phenomenon that has occurred on several occasions over many years.
Sansonetti, P. Host–pathogen interactions: the seduction of molecular cross talk. Gut 50, Suppl. 3 S2–S8 (2002).
Zychlinsky, A., Prevost, M. C. & Sansonetti, P. J. Shigella flexneri induces apoptosis in infected macrophages. Nature 358, 167–169 (1992).
Buchrieser, C. et al. The virulence plasmid pWR100 and the repertoire of proteins secreted by the type III secretion apparatus of Shigella flexneri. Mol. Microbiol. 38, 760–771 (2000).
Egile, C. et al. Activation of the CDC42 effector N-WASP by the Shigella flexneri IcsA protein promotes actin nucleation by Arp2/3 complex and bacterial actin- based motility. J. Cell Biol. 146, 1319–1332 (1999).
Sansonetti, P. J. et al. Caspase-1 activation of IL-1β and IL-18 are essential for Shigella flexneri-induced inflammation. Immunity. 12, 581–590 (2000). The pathogenesis of Shigella infection represents a complex manipulation of the immune response, in ways that are beneficial to both pathogen and host.
Tran Van Nhieu, G., Bourdet-Sicard, R., Dumenil, G., Blocker, A. & Sansonetti, P. J. Bacterial signals and cell responses during Shigella entry into epithelial cells. Cell. Microbiol. 2, 187–193 (2000).
Niebuhr, K. et al. Conversion of PtdIns(4,5)P2 into PtdIns5P by the S. flexneri effector IpgD reorganizes host cell morphology. EMBO J. 21, 5069–5078 (2002).
Scaletsky, I. C. et al. Diffusely adherent Escherichia coli as a cause of acute diarrhea in young children in northeast Brazil: a case-control study. J. Clin. Microbiol. 40, 645–648 (2002).
Bilge, S. S., Clausen, C. R., Lau, W. & Moseley, S. L. Molecular characterization of a fimbrial adhesin, F1845, mediating diffuse adherence of diarrhea-associated Escherichia coli to HEp-2 cells. J. Bacteriol. 171, 4281–4289 (1989).
Hasan, R. J. et al. Structure–function analysis of decayaccelerating factor: identification of residues important for binding of the Escherichia coli Dr adhesin and complement regulation. Infect. Immun. 70, 4485–4493 (2002).
Bernet-Camard, M. F., Coconnier, M. H., Hudault, S. & Servin, A. L. Pathogenicity of the diffusely adhering strain Escherichia coli C1845: F1845 adhesin-decay accelerating factor interaction, brush border microvillus injury, and actin disassembly in cultured human intestinal epithelial cells. Infect. Immun. 64, 1918–1928 (1996).
Peiffer, I., Servin, A. L. & Bernet-Camard, M. F. Piracy of decay-accelerating factor (CD55) signal transduction by the diffusely adhering strain Escherichia coli C1845 promotes cytoskeletal F-actin rearrangements in cultured human intestinal INT407 cells. Infect. Immun. 66, 4036–4042 (1998). DAEC exhibits a unique pathogenetic scheme that includes cytoskeletal sabotage.
Peiffer, I., Bernet-Camard, M. F., Rousset, M. & Servin, A. L. Impairments in enzyme activity and biosynthesis of brush border-associated hydrolases in human intestinal Caco-2/TC7 cells infected by members of the Afa/Dr family of diffusely adhering Escherichia coli. Cell. Microbiol. 3, 341–357 (2001).
Phillips, I. et al. Epidemic multiresistant Escherichia coli infection in West Lambeth Health District. Lancet 1, 1038–1041 (1988).
Manges, A. R. et al. Widespread distribution of urinary tract infections caused by a multidrug-resistant Escherichia coli clonal group. N. Engl. J. Med. 345, 1007–1013 (2001). This work identified specific clonal groups of E. coli that cause widespread antibiotic resistant bacteria.
Nowicki, B., Svanborg-Eden, C., Hull, R. & Hull, S. Molecular analysis and epidemiology of the Dr hemagglutinin of uropathogenic Escherichia coli. Infect. Immun. 57, 446–451 (1989).
Johnson, J. R. Virulence factors in Escherichia coli urinary tract infection. Clin. Microbiol. Rev. 4, 80–128 (1991).
Johnson, J. R. & Stell, A. L. Extended virulence genotypes of Escherichia coli strains from patients with urosepsis in relation to phylogeny and host compromise. J. Infect. Dis. 181, 261–272 (2000).
Welch, R. A. et al. Extensive mosaic structure revealed by the complete genome sequence of uropathogenic Escherichia coli. Proc. Natl Acad. Sci. USA 99, 17020–17024 (2002). The first complete nucleotide sequence of a representative uropathogenic strain of E. coli and shows that EHEC, UPEC and E. coli K-12 share only 39.2% of the combined set of predicted proteins.
Bahrani-Mougeot, F. K. et al. Type 1 fimbriae and extracellular polysaccharides are preeminent uropathogenic Escherichia coli virulence determinants in the murine urinary tract. Mol. Microbiol. 45, 1079–1093 (2002).
Gunther, N. W., Lockatell, V., Johnson, D. E. & Mobley, H. L. In vivo dynamics of type 1 fimbria regulation in uropathogenic Escherichia coli during experimental urinary tract infection. Infect. Immun. 69, 2838–2846 (2001).
Connell, I. et al. Type 1 fimbrial expression enhances Escherichia coli virulence for the urinary tract. Proc. Natl Acad. Sci. USA 93, 9827–9832 (1996). Demonstrates that type 1 fimbriae satisfies molecular Koch's postulates.
Mulvey, M. A. et al. Induction and evasion of host defenses by type 1-piliated uropathogenic Escherichia coli. Science 282, 1494–1497 (1998).
Anderson, G. G. et al. Intracellular bacterial biofilm-like pods in urinary tract infections. Science 301, 105–107 (2003).
Svanborg-Eden, C. & Hansson, H. A. Escherichia coli pili as possible mediators of attachment to human urinary tract epithelial cells. Infect. Immun. 21, 229–237 (1978).
Korhonen, T. K., Virkola, R. & Holthofer, H. Localization of binding sites for purified Escherichia coli P fimbriae in the human kidney. Infect. Immun. 54, 328–332 (1986).
Trifillis, A. L. et al. Binding to and killing of human renal epithelial cells by hemolytic P-fimbriated E. coli. Kidney Int. 46, 1083–1091 (1994).
Uhlen, P. et al. α-haemolysin of uropathogenic E. coli induces Ca2+ oscillations in renal epithelial cells. Nature 405, 694–697 (2000).
Guyer, D. M., Henderson, I. R., Nataro, J. P. & Mobley, H. L. Identification of Sat, an autotransporter toxin produced by uropathogenic Escherichia coli. Mol. Microbiol. 38, 53–66 (2000). Describes the identification of a new toxin of uropathogenic E. coli
Unhanand, M., Mustafa, M. M., McCracken, G. H. Jr & Nelson, J. D. Gram-negative enteric bacillary meningitis: a twenty-one-year experience. J. Pediatr. 122, 15–21 (1993).
Dawson, K. G., Emerson, J. C. & Burns, J. L. Fifteen years of experience with bacterial meningitis. Pediatr. Infect. Dis. J. 18, 816–822 (1999).
Stoll, B. J. et al. Changes in pathogens causing early-onset sepsis in very-low-birth-weight infants. N. Engl. J. Med. 347, 240–247 (2002).
Dietzman, D. E., Fischer, G. W. & Schoenknecht, F. D. Neonatal Escherichia coli septicemia-bacterial counts in blood. J. Pediatr. 85, 128–130 (1974).
Stins, M. F., Badger, J. L. & Kim, K. S. Bacterial invasion and transcytosis in transfected human brain microvascular endothelial cells. Microb. Pathog. 30, 19–28 (2001).
Stins, M. F., Nemani, P. V., Wass, C. & Kim, K. S. Escherichia coli binding to and invasion of brain microvascular endothelial cells derived from humans and rats of different ages. Infect. Immun. 67, 5522–5525 (1999).
Kim, K. S. et al. The K1 capsule is the critical determinant in the development of Escherichia coli meningitis in the rat. J. Clin. Invest. 90, 897–905 (1992). Recognition of the importance of capsule for virulence of MNEC.
Rode, C. K., Melkerson-Watson, L. J., Johnson, A. T. & Bloch, C. A. Type-specific contributions to chromosome size differences in Escherichia coli. Infect. Immun. 67, 230–236 (1999).
Bonacorsi, S. P. et al. Identification of regions of the Escherichia coli chromosome specific for neonatal meningitis-associated strains. Infect. Immun. 68, 2096–2101 (2000).
Badger, J. L., Wass, C. A., Weissman, S. J. & Kim, K. S. Application of signature-tagged mutagenesis for identification of Escherichia coli K1 genes that contribute to invasion of human brain microvascular endothelial cells. Infect. Immun. 68, 5056–5061 (2000).
Parkkinen, J., Korhonen, T. K., Pere, A., Hacker, J. & Soinila, S. Binding sites in the rat brain for Escherichia coli S fimbriae associated with neonatal meningitis. J. Clin. Invest 81, 860–865 (1988).
Prasadarao, N. V., Wass, C. A. & Kim, K. S. Endothelial cell GlcNAc β,1-4GlcNAc epitopes for outer membrane protein A enhance traversal of Escherichia coli across the blood–brain barrier. Infect. Immun. 64, 154–160 (1996).
Kim, K. S. Escherichia coli translocation at the blood–brain barrier. Infect. Immun. 69, 5217–5222 (2001).
Badger, J. L. & Kim, K. S. Environmental growth conditions influence the ability of Escherichia coli K1 to invade brain microvascular endothelial cells and confer serum resistance. Infect. Immun. 66, 5692–5697 (1998).
Hoffman, J. A., Wass, C., Stins, M. F. & Kim, K. S. The capsule supports survival but not traversal of Escherichia coli K1 across the blood–brain barrier. Infect. Immun. 67, 3566–3570 (1999).
Reddy, M. A., Wass, C. A., Kim, K. S., Schlaepfer, D. D. & Prasadarao, N. V. Involvement of focal adhesion kinase in Escherichia coli invasion of human brain microvascular endothelial cells. Infect. Immun. 68, 6423–6430 (2000).
Khan, M. A. & Isaacson, R. E. Identification of Escherichia coli genes that are specifically expressed in a murine model of septicemic infection. Infect. Immun. 70, 3404–3412 (2002).
Darfeuille-Michaud, A. Adherent-invasive Escherichia coli: a putative new E. coli pathotype associated with Crohn's disease. Int. J. Med. Microbiol. 292, 185–193 (2002).
Panigrahi, P., Bamford, P., Horvath, K., Morris, J. G. Jr & Gewolb, I. H. Escherichia coli transcytosis in a Caco-2 cell model: implications in neonatal necrotizing enterocolitis. Pediatr. Res. 40, 415–421 (1996).
De Rycke, J., Milon, A. & Oswald, E. Necrotoxic Escherichia coli (NTEC): two emerging categories of human and animal pathogens. Vet. Res. 30, 221–233 (1999).
Elliott, S. J. et al. Characterization of the roles of hemolysin and other toxins in enteropathy caused by α-hemolytic Escherichia coli linked to human diarrhea. Infect. Immun. 66, 2040–2051 (1998).
Toth, I., Herault, F., Beutin, L. & Oswald, E. Production of cytolethal distending toxins by pathogenic Escherichia coli strains isolated from human and animal sources: establishment of the existence of a new cdt variant (type IV). J. Clin. Microbiol. 41, 4285–4291 (2003).
Janka, A. et al. Cytolethal distending toxin gene cluster in enterohemorrhagic Escherichia coli O157:H- and O157:H7: characterization and evolutionary considerations. Infect. Immun. 71, 3634–3638 (2003).
Otto, B. R., van Dooren, S. J., Dozois, C. M., Luirink, J. & Oudega, B. Escherichia coli hemoglobin protease autotransporter contributes to synergistic abscess formation and heme-dependent growth of Bacteroides fragilis. Infect. Immun. 70, 5–10 (2002).
McVeigh, A. et al. IS 1414, an Escherichia coli insertion sequence with a heat-stable enterotoxin gene embedded in a transposase-like gene. Infect. Immun. 68, 5710–5715 (2000).
Hacker, J. & Kaper, J. B. Pathogenicity islands and the evolution of microbes. Annu. Rev. Microbiol. 54, 641–679 (2000).
Dobrindt, U. et al. Genetic structure and distribution of four pathogenicity islands (PAI I(536) to PAI IV(536)) of uropathogenic Escherichia coli strain 536. Infect. Immun. 70, 6365–6372 (2002).
Tauschek, M., Strugnell, R. A. & Robins-Browne, R. M. Characterization and evidence of mobilization of the LEE pathogenicity island of rabbit-specific strains of enteropathogenic Escherichia coli. Mol. Microbiol. 44, 1533–1550 (2002).
Torres, A. G. & Kaper, J. B. Pathogenicity islands of intestinal E. coli. Curr. Top. Microbiol. Immunol. 264, 31–48 (2002).
Ingersoll, M., Groisman, E. A. & Zychlinsky, A. Pathogenicity islands of Shigella. Curr. Top. Microbiol. Immunol. 264, 49–65 (2002).
Redford, P. & Welch, R. A. Extraintestinal Escherichia coli as a model system for the study of pathogenicity islands. Curr. Top. Microbiol. Immunol. 264, 15–30 (2002).
Schubert, S., Cuenca, S., Fischer, D. & Heesemann, J. High-pathogenicity island of Yersinia pestis in enterobacteriaceae isolated from blood cultures and urine samples: prevalence and functional expression. J. Infect. Dis. 182, 1268–1271 (2000).
Maurelli, A. T., Fernández, R. E., Bloch, C. A., Rode, C. K. & Fasano, A. 'Black holes' and bacterial pathogenicity: a large genomic deletion that enhances the virulence of Shigella spp. and enteroinvasive Escherichia coli. Proc. Natl Acad. Sci. USA 95, 3943–3948 (1998).
Casalino, M., Latella, M. C., Prosseda, G. & Colonna, B. CadC is the preferential target of a convergent evolution driving enteroinvasive Escherichia coli toward a lysine decarboxylase-defective phenotype. Infect. Immun. 71, 5472–5479 (2003).
Weissman, S. J., Moseley, S. L., Dykhuizen, D. E. & Sokurenko, E. V. Enterobacterial adhesins and the case for studying SNPs in bacteria. Trends Microbiol. 11, 115–117 (2003).
Dobrindt, U. et al. Analysis of genome plasticity in pathogenic and commensal Escherichia coli isolates by use of DNA arrays. J. Bacteriol. 185, 1831–1840 (2003).
Mellies, J. L., Elliott, S. J., Sperandio, V., Donnenberg, M. S. & Kaper, J. B. The Per regulon of enteropathogenic Escherichia coli: identification of a regulatory cascade and a novel transcriptional activator, the locus of enterocyte effacement (LEE)-encoded regulator (Ler). Mol. Microbiol. 33, 296–306 (1999).
Xia, Y., Gally, D., Forsman-Semb, K. & Uhlin, B. E. Regulatory cross-talk between adhesin operons in Escherichia coli: inhibition of type 1 fimbriae expression by the PapB protein. EMBO J. 19, 1450–1457 (2000).
Wagner, P. L. et al. Bacteriophage control of Shiga toxin 1 production and release by Escherichia coli. Mol. Microbiol. 44, 957–970 (2002).
Zhang, X. et al. Quinolone antibiotics induce Shiga toxin-encoding bacteriophages, toxin production, and death in mice. J. Infect. Dis. 181, 664–670 (2000).
Sperandio, V., Mellies, J. L., Nguyen, W., Shin, S. & Kaper, J. B. Quorum sensing controls expression of the type III secretion gene transcription and protein secretion in enterohemorrhagic and enteropathogenic Escherichia coli. Proc. Natl Acad. Sci. USA 96, (1999). First report that enteric bacterial virulence factors are regulated by quorum sensing.
Sperandio, V., Torres, A. G., Jarvis, B., Nataro, J. P. & Kaper, J. B. Bacteria–host communication: the language of hormones. Proc. Natl Acad. Sci. USA 100, 8951–8956 (2003).
Abraham, J. M., Freitag, C. S., Clements, J. R. & Eisenstein, B. I. An invertible element of DNA controls phase variation of type 1 fimbriae of Escherichia coli. Proc. Natl Acad. Sci. USA 82, 5724–5727 (1985).
Kuehn, M. J., Heuser, J., Normark, S. & Hultgren, S. J. P pili in uropathogenic E. coli are composite fibres with distinct fibrillar adhesive tips. Nature 356, 252–255 (1992).
Bustamante, V. H., Santana, F. J., Calva, E. & Puente, J. L. Transcriptional regulation of type III secretion genes in enteropathogenic Escherichia coli: Ler antagonizes H-NS-dependent repression. Mol. Microbiol. 39, 664–678 (2001).
Friedberg, D., Umanski, T., Fang, Y. & Rosenshine, I. Hierarchy in the expression of the locus of enterocyte effacement genes of enteropathogenic Escherichia coli. Mol. Microbiol. 34, 941–952 (1999).
Goldberg, M. D., Johnson, M., Hinton, J. C. & Williams, P. H. Role of the nucleoid-associated protein Fis in the regulation of virulence properties of enteropathogenic Escherichia coli. Mol. Microbiol. 41, 549–559 (2001).
Grant, A. J. et al. Co-ordination of pathogenicity island expression by the BipA GTPase in enteropathogenic Escherichia coli (EPEC). Mol. Microbiol. 48, 507–521 (2003).
Sperandio, V., Li, C. C. & Kaper, J. B. Quorum-sensing Escherichia coli regulator A: a regulator of the LysR family involved in the regulation of the locus of enterocyte effacement pathogenicity island in enterohemorrhagic E. coli. Infect. Immun. 70, 3085–3093 (2002).
Sperandio, V., Torres, A. G. & Kaper, J. B. Quorum sensing Escherichia coli regulators B and C (QseBC): a novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in E. coli. Mol. Microbiol. 43, 809–821 (2002).
Gómez-Duarte, O. G. & Kaper, J. B. A plasmid-encoded regulatory region activates chromosomal eaeA expression in enteropathogenic Escherichia coli. Infect. Immun. 63, 1767–1776 (1995).
Tobe, T., Schoolnik, G. K., Sohel, I., Bustamante, V. H. & Puente, J. L. Cloning and characterization of bfpTVW, genes required for the transcriptional activation of bfpA in enteropathogenic Escherichia coli. Mol. Microbiol. 21, 963–975 (1996).
Shin, S. et al. An activator of glutamate decarboxylase genes regulates the expression of enteropathogenic Escherichia coli virulence genes through control of the plasmid-encoded regulator, Per. Mol. Microbiol. 41, 1133–1150 (2001).
Mellies, J. L. et al. espC pathogenicity island of enteropathogenic Escherichia coli encodes an enterotoxin. Infect. Immun. 69, 315–324 (2001)
Acknowledgements
Work in the authors' laboratories is supported by the National Institutes of Health. We thank J. Girón for providing electron micrographs. We apologize to the numerous investigators whose papers could not be cited due to space constraints.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez
LocusLink
The Protein Data Bank
SwissProt
FURTHER INFORMATION
Glossary
- PATHOTYPES
-
A group of strains of a single species that cause a common disease using a common set of virulence factors.
- SEROGROUP
-
An antigenically distinct variety of serotype, based only on O (LPS) antigens.
- SEROTYPE
-
An antigenically distinct variety within a bacterial species. For E. coli, a specific combination of O (lipopolysaccharide), H (flagellar) and sometimes K (capsular) antigens defines a serotype.
- DECAY-ACCELERATING FACTOR
-
(DAF). A plasma membrane protein, also called CD55, that regulates the complement cascade by interfering with the formation of the C3bBb complex.
- MICA
-
A homologue of MHC (major histocompatibility complex) I molecules. Two homologues have been described called MICA (MHC class I chain-related gene A) and MICB (MHC class I chain-related gene B).
- TH1 IMMUNE RESPONSE
-
A response that is characterized by a subset of helper T cells that secrete a particular set of cytokines, including IL-2, interferon-γ and TNF-α, the main function of which is to stimulate phagocytosis-mediated defences against intracellular pathogens.
- NUCLEOLIN
-
A nucleolar protein that functions as a shuttle protein between the nucleus and the cytoplasm and is also found on the cell surface.
- GAP
-
GTPase-activating protein. A family of eukaryotic proteins that modulate the activity of Rac, Rho and Cdc42.
- GALANIN
-
A neuropeptide that is widely distributed in the central nervous system and the gastrointestinal tract. Binding to the galanin-1 receptor can alter intestinal ion flux.
- USSING CHAMBER
-
A device that is used to measure ion flow across an epithelium. Bacterial enterotoxins that induce ion fluxes are frequently studied in Ussing chambers.
- SIGNATURE-TAGGED MUTAGENESIS
-
(STM). A technique to screen large numbers of distinct mutants for those that fail to survive an animal infection. Each mutant is tagged with a unique DNA sequence (called a signature tag), which allows a specific mutant to be tracked within a large pool of bacteria.
- IVET
-
In vivo expression technology is a promoter trap technique that uses cloned promoters fused to a reporter gene. A library of such constructs is introduced into an animal model to detect promoters that are activated in vivo.
Rights and permissions
About this article
Cite this article
Kaper, J., Nataro, J. & Mobley, H. Pathogenic Escherichia coli. Nat Rev Microbiol 2, 123–140 (2004). https://doi.org/10.1038/nrmicro818
Issue Date:
DOI: https://doi.org/10.1038/nrmicro818
This article is cited by
-
Impact of an oligosaccharide-based polymer on the metabolic profiles and microbial ecology of weanling pigs experimentally infected with a pathogenic E. coli
Journal of Animal Science and Biotechnology (2024)
-
Immunity to Cryptosporidium: insights into principles of enteric responses to infection
Nature Reviews Immunology (2024)
-
Avian reservoirs or vectors? Unravelling the relationship between waterbirds and E. coli contamination
Ornithology Research (2024)
-
Synthesis and pharmacological evaluation of novel coumarin based triazolyl glycoconjugates as potential antibacterial and anti-proliferative agents
Medicinal Chemistry Research (2024)
-
Evaluation of stability of (1R,2 S)-(−)-2-methylamino-1-phenyl-1-propanol hydrochloride in plasma and urine samples-inoculated with Escherichia coli using high-performance liquid chromatography (HPLC)
World Journal of Microbiology and Biotechnology (2024)