Key Points
-
CRISPR–Cas systems provide archaea and bacteria with adaptive immunity against viruses and plasmids.
-
CRISPR–Cas genomic loci show extreme diversity in sequence and gene arrangement.
-
We developed a computational approach for CRISPR–Cas classification, combining comparisons of Cas protein sequences and locus architectures.
-
Two classes, five types and 16 subtypes of CRISPR–Cas systems were identified based on this approach.
-
An automated classifier was developed for assigning CRISPR–Cas loci from sequenced genomes to specific subtypes.
-
The evolution of CRISPR–Cas systems is marked by extensive horizontal transfer and recombination of functional modules.
Abstract
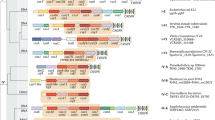
The evolution of CRISPR–cas loci, which encode adaptive immune systems in archaea and bacteria, involves rapid changes, in particular numerous rearrangements of the locus architecture and horizontal transfer of complete loci or individual modules. These dynamics complicate straightforward phylogenetic classification, but here we present an approach combining the analysis of signature protein families and features of the architecture of cas loci that unambiguously partitions most CRISPR–cas loci into distinct classes, types and subtypes. The new classification retains the overall structure of the previous version but is expanded to now encompass two classes, five types and 16 subtypes. The relative stability of the classification suggests that the most prevalent variants of CRISPR–Cas systems are already known. However, the existence of rare, currently unclassifiable variants implies that additional types and subtypes remain to be characterized.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Deveau, H., Garneau, J. E. & Moineau, S. CRISPR/Cas system and its role in phage-bacteria interactions. Annu. Rev. Microbiol. 64, 475–493 (2010).
Marraffini, L. A. & Sontheimer, E. J. CRISPR interference: RNA-directed adaptive immunity in bacteria and archaea. Nat. Rev. Genet. 11, 181–190 (2010).
Koonin, E. V. & Makarova, K. S. CRISPR–Cas: evolution of an RNA-based adaptive immunity system in prokaryotes. RNA Biol. 10, 679–686 (2013).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. The basic building blocks and evolution of CRISPR–Cas systems. Biochem. Soc. Trans. 41, 1392–1400 (2013).
Barrangou, R. & Marraffini, L. A. CRISPR-Cas systems: prokaryotes upgrade to adaptive immunity. Mol. Cell 54, 234–244 (2014).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Barrangou, R. CRISPR–Cas systems and RNA-guided interference. Wiley Interdiscip. Rev. RNA 4, 267–278 (2013).
Westra, E. R. et al. The CRISPRs, they are a-changin': how prokaryotes generate adaptive immunity. Annu. Rev. Genet. 46, 311–339 (2012).
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Magadán, A. H., Dupuis, M. E., Villion, M. & Moineau, S. Cleavage of phage DNA by the Streptococcus thermophilus CRISPR3–Cas system. PLoS ONE 7, e40913 (2012).
van der Oost, J., Jore, M. M., Westra, E. R., Lundgren, M. & Brouns, S. J. CRISPR-based adaptive and heritable immunity in prokaryotes. Trends Biochem. Sci. 34, 401–407 (2009).
Makarova, K. S. et al. Evolution and classification of the CRISPR–Cas systems. Nat. Rev. Microbiol. 9, 467–477 (2011).
Westra, E. R., Buckling, A. & Fineran, P. C. CRISPR–Cas systems: beyond adaptive immunity. Nat. Rev. Microbiol. 12, 317–326 (2014).
Sampson, T. R. & Weiss, D. S. CRISPR–Cas systems: new players in gene regulation and bacterial physiology. Front. Cell. Infect. Microbiol. 4, 37 (2014).
Louwen, R., Staals, R. H., Endtz, H. P., van Baarlen, P. & van der Oost, J. The role of CRISPR–Cas systems in virulence of pathogenic bacteria. Microbiol. Mol. Biol. Rev. 78, 74–88 (2014).
Nunez, J. K. et al. Cas1–Cas2 complex formation mediates spacer acquisition during CRISPR–Cas adaptive immunity. Nat. Struct. Mol. Biol. 21, 528–534 (2014).
Yosef, I., Goren, M. G. & Qimron, U. Proteins and DNA elements essential for the CRISPR adaptation process in Escherichia coli. Nucleic Acids Res. 40, 5569–5576 (2012).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Shah, S. A., Erdmann, S., Mojica, F. J. & Garrett, R. A. Protospacer recognition motifs: mixed identities and functional diversity. RNA Biol. 10, 891–899 (2013).
Bolotin, A., Quinquis, B., Sorokin, A. & Ehrlich, S. D. Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151, 2551–2561 (2005).
Mojica, F. J., Díez-Villaseñor, C., García-Martínez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155, 733–740 (2009).
Wang, R., Preamplume, G., Terns, M. P., Terns, R. M. & Li, H. Interaction of the Cas6 riboendonuclease with CRISPR RNAs: recognition and cleavage. Structure 19, 257–264 (2011).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Samai, P. et al. Co-transcriptional DNA and RNA cleavage during type III CRISPR–Cas immunity. Cell 161, 1164–1174 (2015).
Hale, C. R. et al. Essential features and rational design of CRISPR RNAs that function with the Cas RAMP module complex to cleave RNAs. Mol. Cell 45, 292–302 (2012).
Sashital, D. G., Wiedenheft, B. & Doudna, J. A. Mechanism of foreign DNA selection in a bacterial adaptive immune system. Mol. Cell 46, 606–615 (2012).
van Duijn, E. et al. Native tandem and ion mobility mass spectrometry highlight structural and modular similarities in clustered-regularly-interspaced shot-palindromic-repeats (CRISPR)-associated protein complexes from Escherichia coli and Pseudomonas aeruginosa. Mol. Cell Proteom. 11, 1430–1441 (2012).
Zhang, J. et al. Structure and mechanism of the CMR complex for CRISPR-mediated antiviral immunity. Mol. Cell 45, 303–313 (2012).
Wiedenheft, B. et al. Structures of the RNA-guided surveillance complex from a bacterial immune system. Nature 477, 486–489 (2011).
Haft, D. H., Selengut, J., Mongodin, E. F. & Nelson, K. E. A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput. Biol. 1, e60 (2005).
Makarova, K. S., Aravind, L., Wolf, Y. I. & Koonin, E. V. Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR–Cas systems. Biol. Direct 6, 38 (2011).
Makarova, K. S., Grishin, N. V., Shabalina, S. A., Wolf, Y. I. & Koonin, E. V. A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol. Direct 1, 7 (2006).
Vestergaard, G., Garrett, R. A. & Shah, S. A. CRISPR adaptive immune systems of Archaea. RNA Biol. 11, 156–167 (2014).
Staals, R. H. et al. Structure and activity of the RNA-targeting Type III-B CRISPR–Cas complex of Thermus thermophilus. Mol. Cell 52, 135–145 (2013).
Spilman, M. et al. Structure of an RNA silencing complex of the CRISPR–Cas immune system. Mol. Cell 52, 146–152 (2013).
Staals, R. H. et al. RNA targeting by the type III-A CRISPR–Cas Csm complex of Thermus thermophilus. Mol. Cell 56, 518–530 (2014).
Tamulaitis, G. et al. Programmable RNA shredding by the type III-A CRISPR–Cas system of Streptococcus thermophilus. Mol. Cell 56, 506–517 (2014).
Benda, C. et al. Structural model of a CRISPR RNA-silencing complex reveals the RNA-target cleavage activity in Cmr4. Mol. Cell 56, 43–54 (2014).
Hale, C. R., Cocozaki, A., Li, H., Terns, R. M. & Terns, M. P. Target RNA capture and cleavage by the Cmr type III-B CRISPR–Cas effector complex. Genes Dev. 28, 2432–2443 (2014).
van der Oost, J., Westra, E. R., Jackson, R. N. & Wiedenheft, B. Unravelling the structural and mechanistic basis of CRISPR–Cas systems. Nat. Rev. Microbiol. 12, 479–492 (2014).
Jackson, R. N. Lavin, M., Carter, J., & Wiedenheft, B. Fitting CRISPR-associated Cas3 into the helicase family tree. Curr Opin Struct Biol. 24, 106–114 (2014).
Mulepati, S., Heroux, A. & Bailey, S. Crystal structure of a CRISPR RNA-guided surveillance complex bound to a ssDNA target. Science 345, 1479–1484 (2014).
Zhao, H. et al. Crystal structure of the RNA-guided immune surveillance Cascade complex in Escherichia coli. Nature 515, 147–150 (2014).
Taylor, D. W. et al. Structures of the CRISPR–Cmr complex reveal mode of RNA target positioning. Science 348, 581–585 (2015).
Mali, P., Esvelt, K. M. & Church, G. M. Cas9 as a versatile tool for engineering biology. Nat. Methods 10, 957–963 (2013).
Sander, J. D. & Joung, J. K. CRISPR–Cas systems for editing, regulating and targeting genomes. Nat. Biotech. 32, 347–355 (2014).
Altschul, S. F. & Koonin, E. V. PSI-BLAST — a tool for making discoveries in sequence databases. Trends Biochem. Sci. 23, 444–447 (1998).
Sinkunas, T. et al. Cas3 is a single-stranded DNA nuclease and ATP-dependent helicase in the CRISPR/Cas immune system. EMBO J. 30, 1335–1342 (2011).
Gong, B. et al. Molecular insights into DNA interference by CRISPR-associated nuclease-helicase Cas3. Proc. Natl Acad. Sci. USA 111, 16359–16364 (2014).
Huo, Y. et al. Structures of CRISPR Cas3 offer mechanistic insights into Cascade-activated DNA unwinding and degradation. Nat. Struct. Mol. Biol. 21, 771–777 (2014).
Mulepati, S. & Bailey, S. Structural and biochemical analysis of nuclease domain of clustered regularly interspaced short palindromic repeat (CRISPR)-associated protein 3 (Cas3). J. Biol. Chem. 286, 31896–31903 (2011).
Makarova, K. S. & Koonin, E. V. Annotation and classification of CRISPR–Cas systems. Methods Mol. Biol. 1311, 47–75 (2015).
Nam, K. H. et al. Cas5d protein processes pre-crRNA and assembles into a cascade-like interference complex in subtype I-C/Dvulg CRISPR–Cas system. Structure 20, 1574–1584 (2012).
Makarova, K. S., Aravind, L., Grishin, N. V., Rogozin, I. B. & Koonin, E. V. A DNA repair system specific for thermophilic Archaea and bacteria predicted by genomic context analysis. Nucleic Acids Res. 30, 482–496 (2002).
Hale, C. R. et al. RNA-guided RNA cleavage by a CRISPR RNA–Cas protein complex. Cell 139, 945–956 (2009).
Marraffini, L. A. & Sontheimer, E. J. CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322, 1843–1845 (2008).
Goldberg, G. W., Jiang, W., Bikard, D. & Marraffini, L. A. Conditional tolerance of temperate phages via transcription-dependent CRISPR–Cas targeting. Nature 514, 633–637 (2014).
Deng, L., Garrett, R. A., Shah, S. A., Peng, X. & She, Q. A novel interference mechanism by a type IIIB CRISPR–Cmr module in Sulfolobus. Mol. Microbiol. 87, 1088–1099 (2013).
Peng, W., Feng, M., Feng, X., Liang, Y. X. & She, Q. An archaeal CRISPR type III-B system exhibiting distinctive RNA targeting features and mediating dual RNA and DNA interference. Nucleic Acids Res. 43, 406–417 (2015).
White, M. F. Structure, function and evolution of the XPD family of iron-sulfur-containing 5′→3′ DNA helicases. Biochem. Soc. Trans. 37, 547–551 (2009).
Heler, R. et al. Cas9 specifies functional viral targets during CRISPR–Cas adaptation. 519, 199–202 Nature (2015).
Wei, Y., Terns, R. M. & Terns, M. P. Cas9 function and host genome sampling in Type II-A CRISPR–Cas adaptation. Genes Dev. 29, 356–361 (2015).
Chylinski, K., Makarova, K. S., Charpentier, E. & Koonin, E. V. Classification and evolution of type II CRISPR–Cas systems. Nucleic Acids Res. 42, 6091–6105 (2014).
Chylinski, K., Le Rhun, A. & Charpentier, E. The tracrRNA and Cas9 families of type II CRISPR–Cas immunity systems. RNA Biol. 10, 726–737 (2013).
Briner, A. E. et al. Guide RNA functional modules direct Cas9 activity and orthogonality. Mol. Cell 56, 333–339 (2014).
Fonfara, I. et al. Phylogeny of Cas9 determines functional exchangeability of dual-RNA and Cas9 among orthologous type II CRISPR–Cas systems. Nucleic Acids Res. 42, 2577–2590 (2014).
Zhang, Y. et al. Processing-independent CRISPR RNAs limit natural transformation in Neisseria meningitidis. Mol. Cell 50, 488–503 (2013).
Schunder, E., Rydzewski, K., Grunow, R. & Heuner, K. First indication for a functional CRISPR/Cas system in Francisella tularensis. Int. J. Med. Microbiol. 303, 51–60 (2013).
Makarova, K. S. et al. Dark matter in archaeal genomes: a rich source of novel mobile elements, defense systems and secretory complexes. Extremophiles 18, 877–893 (2014).
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. Comparative genomics of defense systems in archaea and bacteria. Nucleic Acids Res. 41, 4360–4377 (2013).
Grissa, I., Vergnaud, G. & Pourcel, C. The CRISPRdb database and tools to display CRISPRs and to generate dictionaries of spacers and repeats. BMC Bioinformatics 8, 172 (2007).
Bland, C. et al. CRISPR recognition tool (CRT): a tool for automatic detection of clustered regularly interspaced palindromic repeats. BMC Bioinformatics 8, 209 (2007).
Lange, S. J., Alkhnbashi, O. S., Rose, D., Will, S. & Backofen, R. CRISPRmap: an automated classification of repeat conservation in prokaryotic adaptive immune systems. Nucleic Acids Res. 41, 8034–8044 (2013).
Alkhnbashi, O. S. et al. CRISPRstrand: predicting repeat orientations to determine the crRNA-encoding strand at CRISPR loci. Bioinformatics 30, i489–i496 (2014).
Kunin, V., Sorek, R. & Hugenholtz, P. Evolutionary conservation of sequence and secondary structures in CRISPR repeats. Genome Biol. 8, R61 (2007).
Koonin, E. V. & Wolf, Y. I. Is evolution Darwinian or/and Lamarckian? Biol. Direct 4, 42 (2009).
Leplae, R. et al. Diversity of bacterial type II toxin-antitoxin systems: a comprehensive search and functional analysis of novel families. Nucleic Acids Res. 39, 5513–5525 (2011).
Koonin, E. V. & Wolf, Y. I. Evolution of microbes and viruses: a paradigm shift in evolutionary biology? Front. Cell Infect. Microbiol. 2, 119 (2012).
Godde, J. S. & Bickerton, A. The repetitive DNA elements called CRISPRs and their associated genes: evidence of horizontal transfer among prokaryotes. J. Mol. Evol. 62, 718–729 (2006).
Almendros, C., Mojica, F. J., Díez-Villaseñor, C., Guzmán, N. M. & García-Martínez, J. CRISPR–Cas functional module exchange in Escherichia coli. mBio 5, e00767–e00713 (2014).
Shah, S. A. & Garrett, R. A. CRISPR/Cas and Cmr modules, mobility and evolution of adaptive immune systems. Res. Microbiol. 162, 27–38 (2011).
Yutin, N., Puigbo, P., Koonin, E. V. & Wolf, Y. I. Phylogenomics of prokaryotic ribosomal proteins. PLoS ONE 7, e36972 (2012).
Takeuchi, N., Wolf, Y. I., Makarova, K. S. & Koonin, E. V. Nature and intensity of selection pressure on CRISPR-associated genes. J. Bacteriol. 194, 1216–1225 (2012).
Krupovic, M., Makarova, K. S., Forterre, P., Prangishvili, D. & Koonin, E. V. Casposons: a new superfamily of self-synthesizing DNA transposons at the origin of prokaryotic CRISPR–Cas immunity. BMC Biol. 12, 36 (2014).
Koonin, E. V. & Krupovic, M. Evolution of adaptive immunity from transposable elements combined with innate immune systems. Nat. Rev. Genet. 16, 184–192 (2015).
Garrett, R. A., Vestergaard, G. & Shah, S. A. Archaeal CRISPR-based immune systems: exchangeable functional modules. Trends Microbiol. 19, 549–556 (2011).
Nunez, J. K., Lee, A. S., Engelman, A. & Doudna, J. A. Integrase-mediated spacer acquisition during CRISPR–Cas adaptive immunity. Nature 519, 193–198 (2015).
Puigbo, P., Wolf, Y. I. & Koonin, E. V. Search for a 'Tree of Life' in the thicket of the phylogenetic forest. J. Biol. 8, 59 (2009).
Hooton, S. P. & Connerton, I. F. Campylobacter jejuni acquire new host-derived CRISPR spacers when in association with bacteriophages harboring a CRISPR-like Cas4 protein. Front. Microbiol. 5, 744 (2014).
Wiedenheft, B. et al. Structural basis for DNase activity of a conserved protein implicated in CRISPR-mediated genome defense. Structure 17, 904–912 (2009).
Kwon, A. R. et al. Structural and biochemical characterization of HP0315 from Helicobacter pylori as a VapD protein with an endoribonuclease activity. Nucleic Acids Res. 40, 4216–4228 (2012).
Makarova, K. S., Anantharaman, V., Aravind, L. & Koonin, E. V. Live virus-free or die: coupling of antivirus immunity and programmed suicide or dormancy in prokaryotes. Biol. Direct 7, 40 (2012).
Beloglazova, N. et al. A novel family of sequence-specific endoribonucleases associated with the clustered regularly interspaced short palindromic repeats. J. Biol. Chem. 283, 20361–20371 (2008).
Nam, K. H. et al. Double-stranded endonuclease activity in Bacillus halodurans clustered regularly interspaced short palindromic repeats (CRISPR)-associated Cas2 protein. J. Biol. Chem. 287, 35943–35952 (2012).
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008).
Rouillon, C. et al. Structure of the CRISPR interference complex CSM reveals key similarities with cascade. Mol. Cell 52, 124–134 (2013).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 343, 1247997 (2014).
Beloglazova, N. et al. Structure and activity of the Cas3 HD nuclease MJ0384, an effector enzyme of the CRISPR interference. EMBO J. 30, 4616–4627 (2011).
Ramia, N. F. et al. Essential structural and functional roles of the Cmr4 subunit in RNA cleavage by the Cmr CRISPR–Cas complex. Cell Rep. 9, 1610–1617 (2014).
Zhu, X. & Ye, K. Cmr4 is the slicer in the RNA-targeting Cmr CRISPR complex. Nucleic Acids Res. 43, 1257–1267 (2015).
Brendel, J. et al. A complex of Cas proteins 5, 6, and 7 is required for the biogenesis and stability of clustered regularly interspaced short palindromic repeats (crispr)-derived rnas (crrnas) in Haloferax volcanii. J. Biol. Chem. 289, 7164–7177 (2014).
Osawa, T., Inanaga, H., Sato, C. & Numata, T. Crystal structure of the CRISPR–Cas RNA silencing Cmr complex bound to a target analog. Mol. Cell 58, 418–430 (2015).
Jung, T. Y. et al. Crystal structure of the Csm1 subunit of the Csm complex and its single-stranded DNA-specific nuclease activity. Structure 23, 782–790 (2015).
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011).
Makarova, K. S., Anantharaman, V., Grishin, N. V., Koonin, E. V. & Aravind, L. CARF and WYL domains: ligand-binding regulators of prokaryotic defense systems. Front. Genet. 5, 102 (2014).
Nam, K. H., Kurinov, I. & Ke, A. Crystal structure of clustered regularly interspaced short palindromic repeats (CRISPR)-associated Csn2 protein revealed Ca2+-dependent double-stranded DNA binding activity. J. Biol. Chem. 286, 30759–30768 (2011).
Koo, Y., Jung, D. K. & Bae, E. Crystal structure of Streptococcus pyogenes Csn2 reveals calcium-dependent conformational changes in its tertiary and quaternary structure. PLoS ONE 7, e33401 (2012).
Arslan, Z. et al. Double-strand DNA end-binding and sliding of the toroidal CRISPR-associated protein Csn2. Nucleic Acids Res. 41, 6347–6359 (2013).
Lee, K. H. et al. Identification, structural, and biochemical characterization of a group of large Csn2 proteins involved in CRISPR-mediated bacterial immunity. Proteins 80, 2573–2582 (2012).
Zhu, X. & Ye, K. Crystal structure of Cmr2 suggests a nucleotide cyclase-related enzyme in type III CRISPR–Cas systems. FEBS Lett. 586, 939–945 (2012).
Shao, Y. et al. Structure of the Cmr2–Cmr3 subcomplex of the Cmr RNA silencing complex. Structure 21, 376–384 (2013).
Guy, C. P., Majernik, A. I., Chong, J. P. & Bolt, E. L. A novel nuclease-ATPase (Nar71) from archaea is part of a proposed thermophilic DNA repair system. Nucleic Acids Res. 32, 6176–6186 (2004).
Cass, S. D. et al. The role of Cas8 in type I CRISPR interference. Biosci. Rep. 35, e00197 (2015).
Reeks, J. et al. Structure of the archaeal Cascade subunit Csa5: relating the small subunits of CRISPR effector complexes. RNA Biol. 10, 762–769 (2013).
Jackson, R. N. & Wiedenheft, B. A conserved structural chassis for mounting versatile CRISPR RNA-guided immune responses. Mol. Cell 58, 722–728 (2015).
Acknowledgements
K.S.M., Y.I.W., D.H. and E.V.K. are supported by the National Institutes of Health (NIH) Intramural Research Program at the National Library of Medicine, US Department of Health and Human Services. R.M.T. and M.P.T. are supported by NIH grants RO1 GM54682 and RO1 GM99876. J.v.d.O. was partly supported by SIAM Gravitation Grant 024.002.002 from the Netherlands Organization for Scientific Research (N.W.O.). S.J.J.B. was financially supported by an NWO Vidi grant (864.11.005) and European Research Council (ERC) Stg (639707). A.F.Y. is supported by the Natural Sciences and Engineering Research Council (NSERC) Strategic Network Grant IBN and NSERC Discovery grant. S.M. acknowledges funding from Natural Sciences and Engineering Research Council of Canada (Discovery program) and holds a Tier 1 Canada Research Chair in Bacteriophages. F.J.M.M. is supported by the Ministerio de Economía y Competitividad (BIO2014-53029). R.B. is supported by the Deutsche Forschungsgemeinschaft (DFG) grant (BA 2168/5-2). S.A.S. and R.A.G. were funded primarily by the Danish Natural Science Research Council. O.S.A., F.C., S.J.S., R.B., S.A.S and R.A.G. are grateful to all members of the FOR1680 for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Supplementary information
Supplementary information S1 (table)
The core proteins of CRISPR-Cas systems (PDF 257 kb)
Supplementary Information S2 (table)
The list of all Cas profiles with assigned gene names, system types and subtypes and a role in CRISPR-Cas function (XLSX 23 kb)
Supplementary information S3 (box)
Methods (PDF 444 kb)
Supplementary information S4 (table)
The classes, types and subtypes of CRISPR-Cas systems, their signature proteins and key features (PDF 254 kb)
Supplementary information S5 (box)
Cas1 phylogenetic tree in the Newick format (DOCX 48 kb)
Supplementary information S6 (box)
Cas3 phylogenetic tree in the Newick format (DOCX 177 kb)
Supplementary Information S7 (table)
The list of all cas loci, profiles and cas gene names assigned for each gene in the locus (XLSX 1239 kb)
Supplementary information S8 (table)
Distribution of different types and subtypes of CRISPR-Cas in archaeal and bacterial phyla (PDF 169 kb)
Supplementary information S9 (box)
Cas10 phylogenetic trees in the Newick format (DOCX 164 kb)
Supplementary Information S10 (box)
Distribution of array to loci distances. (XLSX 123 kb)
Supplementary information S11 (table)
Comparison of different classifications of CRISPR-Cas systems (PDF 137 kb)
Supplementary information S12 (box)
Locus architecture phylogentic tree in the Newick format (DOCX 46 kb)
Supplementary information S13 (box)
Sequence similarity-based dendrogram of the crRNA-effector complexes in the Newick format (DOCX 179 kb)
Supplementary information S14 (box)
Sequence similarity-based dendrogram of the adaptation cas genes in the Newick format (DOCX 56 kb)
Supplementary information S15 (figure)
Performance of an automated classifier for annotation of CRISPR–cas loci (PDF 112 kb)
Rights and permissions
About this article
Cite this article
Makarova, K., Wolf, Y., Alkhnbashi, O. et al. An updated evolutionary classification of CRISPR–Cas systems. Nat Rev Microbiol 13, 722–736 (2015). https://doi.org/10.1038/nrmicro3569
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3569
This article is cited by
-
Potential therapeutic strategies for osteoarthritis via CRISPR/Cas9 mediated gene editing
Reviews in Endocrine and Metabolic Disorders (2024)
-
An Update on the Application of CRISPR Technology in Clinical Practice
Molecular Biotechnology (2024)
-
The applications of CRISPR/Cas-mediated genome editing in genetic hearing loss
Cell & Bioscience (2023)
-
Potential of CRISPR/Cas system as emerging tools in the detection of viral hepatitis infection
Virology Journal (2023)
-
Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy
Molecular Cancer (2023)