Key Points
-
Mortality from influenza viruses is strongly linked to secondary bacterial invaders. In the most extreme example, more than 95% of the 50 million or more deaths during the 1918 pandemic were complicated by bacterial pneumonia.
-
Influenza viruses differ in their propensity to support bacterial superinfection, depending on the expression of particular virulence factors. Nascent pandemic strains that emerge from the avian reservoir naturally possess many such traits; often these are lost over time as the viruses reach an equilibrium with their hosts.
-
Similarly, different strains of bacteria express different virulence factors that enable them to take advantage of virus-mediated alterations to lung physiology and immunity. The regional distribution of these strains probably influences the severity of influenza epidemics and pandemics.
-
Disruption of lung physiology by respiratory viruses breaches natural barriers to infection and promotes bacterial co-infection. Receptors that can be used by bacteria for adherence and infection are uncovered and upregulated.
-
Viruses and bacteria express factors that subvert, inhibit or eliminate host immune responses. Paradoxically, the resulting pathogen overgrowth might lead to augmented inflammatory responses and immune-mediated host damage.
-
Although they are typically secondary invaders during influenza infections, bacteria express virulence factors that promote viral pathogenesis. This results in increased viral load and decreased clearance.
-
There are many key unanswered questions in the field of co-infections. A better understanding of the complex relationships between virus, host and bacteria will aid us in combating common manifestations, such as community-acquired pneumonia, and help us to prepare for the inevitable next severe influenza pandemic.
Abstract
Concern that a highly pathogenic virus might cause the next influenza pandemic has spurred recent research into influenza and its complications. Bacterial superinfection in the lungs of people suffering from influenza is a key element that promotes severe disease and mortality. This co-pathogenesis is characterized by complex interactions between co-infecting pathogens and the host, leading to the disruption of physical barriers, dysregulation of immune responses and delays in a return to homeostasis. The net effect of this cascade can be the outgrowth of the pathogens, immune-mediated pathology and increased morbidity. In this Review, advances in our understanding of the underlying mechanisms are discussed, and the key questions that will drive the field forwards are articulated.
Similar content being viewed by others
Main
During the 1918 influenza pandemic, more than 50 million individuals died from pneumonia exacerbated by bacterial superinfection1,2. Influenza viruses that have a similar potential to synergize with bacterial pathogens are currently circulating in the wild bird reservoirs of the world3. Much has changed since 1918 that might be expected to ameliorate a severe pandemic, including improved hygiene, vaccines, antibiotics and better insights into the interactions between influenza viruses and bacteria. However, the emergence of a new, highly pathogenic pandemic influenza virus strain would destabilize society, even if it were to cause only a fraction of the death toll of the 1918 strain. Over the past 10 years, research has increasingly focused on the co-pathogenesis of influenza viruses and their bacterial partners in co-infections. The general outlines of the mechanistic underpinning of these interactions can now be glimpsed, but the details remain to be worked out and translated to humans. This Review focuses on recent advances in our understanding of the pathogenesis of viral–bacterial pneumonia in the setting of influenza and articulates the key questions for further research in the field.
Epidemiology and microbiology of co-infections
The epidemiology of co-infections remains difficult to assess with any accuracy. The attribution of mortality to influenza or complications of influenza is complex4, as most deaths are from complications of influenza, rather than the primary disease, and a precise viral aetiology is infrequently confirmed by diagnostic testing or confounded by co-circulating pathogens and co-infections5. Statistical models have been developed to quantify this phenomenon, using the term 'excess mortality' to indicate the number of deaths that are in excess of those expected for the season and that are temporally linked to the circulation of influenza viruses4; however, the greater burden of disease in hospitals and outpatient settings, when death does not occur, is very poorly defined. Despite recent advances in diagnostics, current testing methods for identifying bacterial pathogens within the lungs suffer from a lack of sensitivity and specificity, as the lower respiratory tract is difficult to access and because many pathogens that can cause pneumonia (for example, Streptococcus pneumoniae) can colonize the nasopharynx, which is the typical sampling site for assessing the aetiology of respiratory tract infections6. In addition, sequential infections can be more difficult to accurately diagnose if the preceding pathogen (such as influenza or another virus) has resolved by the time the patient presents with secondary bacterial pneumonia7. Despite these challenges, there is an increasing appreciation that many episodes of community-acquired pneumonia result from co-infections, particularly when associated with influenza8,9.
Co-infections during the 1918 pandemic. The 1918 pandemic was comprehensively studied by leading microbiologists and pathologists of the era (Box 1), and many of their insights have been collated and reinterpreted in recent years10,11. Estimates from clinical case and autopsy series suggest that more than 95% of all severe illnesses and deaths were complicated by bacterial pathogens, most commonly by S. pneumoniae2. However, other common respiratory pathogens, including Staphylococcus aureus, Haemophilus influenzae and Streptococcus pyogenes (group A Streptococcus) were all identified as the predominant pathogen in various individual studies (reviewed in Ref. 12), which suggests that there was regional variation. Furthermore, pre-existing immunity after prior exposure to these bacterial pathogens modified the apparent attack rate and severity of pandemic disease. This is shown by an inverse correlation in members of the Australian army between length of service and mortality from secondary bacterial pneumonia13 and a lower case fatality rate in other military units for established medical care providers compared with newly arrived providers or patients14. Overall, these data indicate that severe disease and mortality were linked to the presence of secondary bacterial invaders, with factors such as variations in which bacterial strains were endemic being modified by personal or population immunity, dictating the epidemiology in a particular region9,12,15.
Co-infections during the 1957 and 1968 pandemics. The patterns of mortality in the next two pandemics in 1957 and 1968 resembled those of seasonal influenza in the respect that bacterial co-infections were a less likely cause of death than they were during the 1918 pandemic. Still, pneumonia accounted for a higher percentage of deaths in 1957–1958 (44%)16 than during most of the preceding inter-pandemic years (∼20%; range 4–44%)17,18, but did not approach the 95% of 1918 (Ref. 2). In addition, most deaths resulting from pneumonia were in people with chronic medical conditions19. In 1957–1958, S. aureus was the most frequent secondary invader to cause fatal pneumonia20,21,22,23,24. The clinical course was often fulminant, with death occurring in less than 7 days, and severe pulmonary oedema and haemorrhage were commonly found on autopsy25. The higher incidence of S . aureus was a striking departure from previous pandemics and seasonal epidemics and was initially blamed on the use of antibiotics that were effective against S. pneumoniae, S. pyogenes and many strains of H. influenzae but not against emerging antibiotic-resistant strains of S. aureus that were prevalent in hospitals at the time24,26. However, the data from the 1968 pandemic suggested that antibiotics were not the only reason for the changes to the microbiology. In 1968–1969, the incidence of pneumonia was comparable to that of 1957 and most mortality was observed in people with underlying chronic diseases, but S. pneumoniae was once again the predominant pathogen27,28,29,30. Thus, it is more likely that strain-related differences in either the virus or the bacterial co-pathogens are responsible for these shifts in aetiology between pandemics.
Recent epidemiology of co-infections. Although S. pneumoniae was considered to be the most common cause of secondary pneumonia in the decades after the 1968 pandemic, S. aureus is emerging as a cause of fulminant pneumonia in association with influenza in many parts of the world31,32. The USA300 and USA400 clonotypes of S. aureus seem to be particularly likely to cause secondary pneumonia with influenza, compared with other circulating strains, for unclear reasons that are probably related to the altered expression or regulation of particular bacterial virulence factors, such as cytotoxins or adherence factors9,33. H. influenzae has become less prominent as a cause of secondary bacterial pneumonia following the introduction of the H. influenzae type B conjugate vaccine in 1985, although it remains important in regions of the world that have poor vaccine coverage34, and non-typeable strains that are not covered by the vaccine continue to be seen in a minority of cases in adults35. Group A Streptococcus is entirely absent from many case series that describe community-acquired pneumonia in association with viruses and is typically third in incidence when it does appear36. Viruses other than influenza also participate in viral–bacterial co-infections, including respiratory syncytial virus (RSV), parainfluenzaviruses, rhinoviruses and adenoviruses37,38,39,40,41,42,43 (Box 2).
In 2009, a novel H1N1 virus emerged from swine and caused the first pandemic in more than 40 years44. In contrast to the 1957 and 1968 pandemics, mortality rates were similar to recent seasonal epidemics and most deaths occurred in young adults, often with no underlying chronic conditions45. Respiratory deaths that were associated with an aberrant immune response to the virus accounted for most of this mortality in previously healthy people45,46,47. The precise effect of secondary bacterial disease remains unclear; some estimates put excess mortality from influenza and pneumonia as low as 10% (Ref. 48), which is lower than that of many seasonal influenza epidemics49. In careful studies of severe or fatal cases, bacterial pneumonia was found to complicate between one-quarter and one-half of infections50,51,52,53,54. S. pneumoniae and S. aureus were the most common aetiologies of secondary bacterial pneumonia, with regional variations in which pathogen was more frequent51,52,55,56. The prominence of S. aureus in 2009–2010 was probably due to strain-specific features of the recently emerged USA300 clonotype, such as Panton–Valentine leukotoxin (PVL) expression31, coupled with increased penetrance of pneumococcal vaccination in the past decade57. Of note, the role of PVL in the pathogenesis of pneumonia is controversial, and its expression might instead be a USA300 genotype marker that commonly assorts with as yet unknown virulence factors that facilitate pneumonia in virus-infected hosts58. S. pyogenes was absent as a secondary pathogen from many case series but was surprisingly frequent in others, which suggests that there are further regional differences in common co-pathogens51,59. In some carefully conducted studies, no serious bacterial superinfections were seen at all, despite extensive sampling60. However, when present, it was clear that bacterial superinfections from S. pneumoniae or S. aureus resulted in worse outcomes55,56,61,62.
Mechanisms of co-pathogenesis
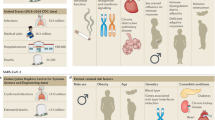
Although mostly based on animal-model data, it is clear that co-pathogenesis between influenza and superinfecting bacteria has a multifactorial basis63,64 (Fig. 1). Several virulence factors that are expressed by the virus have viral strain-specific effects on the host that enable bacteria to cause disease. Multiple host pathways are affected, and specific host states, or factors such as the timing between exposures to the virus and the bacterium, might favour certain outcomes. Although much less is known about the importance of specific bacterial virulence factors, there is growing evidence that strain-specific differences in expression are also important on the bacterial 'side of the equation'.
a | Several virulence factors that are expressed by influenza viruses can directly interact with the lungs or with the host immune system. Haemagglutinin mediates attachment by binding to terminal sialic acids on cell surface proteins and initiating endocytosis. A variety of extracellular proteins can bind to glycans on haemagglutinin and neutralize or help to eliminate viruses from the lower respiratory tract. Viruses with a poorly glycosylated haemagglutinin and the ability to engage both α2,3- and α2,6-linked sialic acids as receptors are able to penetrate deep into the lungs. The sialidase activity of the neuraminidase protein cleaves sialic acids from the surface of epithelial cells and from mucins that try to bind and eliminate virions — this facilitates bacterial access to receptors. The non-structural proteins PB1-F2 and NS1 are made in infected cells. PB1-F2 causes cytotoxicity and promotes inflammatory responses to co-pathogens; NS1 modulates innate pathways, including interferon signalling. b | These virus-mediated effects engender changes in the physical properties of the lungs and compromise innate immunity at several levels. Epithelial damage and increased receptor availability enable bacteria to adhere and grow. Depletion of the specific subset of lung macrophages that is functionally capable of phagocytosing bacteria enables escape from early innate immunity. Anergy of the primary bacterial sensing apparatus of the immune system, pattern recognition receptors, such as the Toll-like receptors (TLRs), prolongs this window of susceptibility for weeks, while a dysfunctional and paradoxically over-exuberant inflammatory response that is characterized by neutrophil influx and cytokine storm furthers the acute lung injury that has been started by the virus. c | Bacteria that express specific virulence factors may take advantage of these changes to the host, grow unchecked and cause disease. Adherence factors, such as pneumococcal surface protein A or staphylococcal MSCRAMMs (microbial surface components recognizing adhesive matrix molecules), enable bacteria to attach to newly uncovered receptors or to the matrix of collagen and fibrin that has been laid down as a scaffold for repair. Bacterial cytotoxins synergize with their viral counterparts to further the physical and immune-mediated damage to the lungs. Specific characteristics of some bacterial strains, such as a thick, complement-resistant capsule, and a set of unknown proteins whose existence can be inferred but have not yet been described, enable improved survival, growth and pathology in virus-infected hosts.
Dysfunction of lung physiology. Influenza virus infection causes multiple changes in the lungs that can facilitate secondary bacterial invasion. Epithelial damage, disruption of surfactant and the sloughing of cells into the airways provide access and a rich source of nutrients, promoting rapid bacterial growth65. Combined with the release of fibrinous materials and the secretion of mucins, small airways become obstructed, leading to dead space, decreased oxygen and carbon dioxide diffusion capacities and lung dysfunction66,67. Ciliary beat frequency is decreased and ciliary motion becomes uncoordinated68. The physiological effects of these functional changes in the respiratory tract are decreased oxygen exchange, airway hyper-reactivity and decreased mechanical clearance of bacteria. These changes are most problematic in hosts with pre-existing conditions that limit pulmonary function, such as patients with chronic obstructive pulmonary disease, who are more likely to have exacerbations, chronic bronchitis and pneumonia during the course of an influenza virus infection69.
Increased receptor availability. The prevailing dogma in the field since 1918 has been that the respiratory epithelial sloughing that is associated with highly pathogenic influenza viruses provides a foothold for bacteria by exposing sites for attachment70,71. Bacteria express a range of virulence factors, such as pneumococcal surface protein A (PsaP), choline-binding protein A (CbpA) and pneumococcal serine-rich repeat protein (PsrP) in S. pneumoniae72, and members of the MSCRAMM (microbial surface components recognizing adhesive matrix molecules) family (for example, fibronectin-binding protein A (FnBPA), FnBPB, clumping factor A (ClfA) and ClfB) and non-MSCRAMM members of the serine–aspartate dipeptide repeat-containing (Sdr) family in S. aureus73, that can be used for adherence to the basement membrane or elements of the extracellular matrix such as fibrin, fibrinogen and collagen74,75 (Fig. 1c). Virulent viruses, such as the mouse-adapted influenza virus strain PR8, cause substantial epithelial cell death in vivo, which exposes sites for adherence in the tracheobronchial tree70,76. In humans, autopsy studies show the adherence of bacteria to sites of epithelial damage during pandemic influenza2,19,21,25,51. As virulence in influenza viruses is a multigenic trait, any mutations that increase viral fitness or cytotoxic potential can contribute to epithelial damage and subsequent secondary bacterial infection. In particular, the viral cytotoxin PB1-F2 (see below) can cause cell death and a cytokine storm77,78,79.
However, other receptor-mediated mechanisms must also be used as most seasonal influenza strains do not cause severe lung injury but can still facilitate bacterial superinfection — although to a lesser degree80. Three additional mechanisms might increase receptor availability. First, the influenza virus neuraminidase cleaves sialic acids, which can expose cryptic receptors for pneumococcal adherence on host cells and disrupt sialylated mucins that can function as decoy receptors for the bacteria81,82 (Fig. 1a). Bacteria that can cause productive infections in the lungs, such as S. pneumoniae, often produce neuraminidases themselves in order to access receptors and avoid host defences by cleaving sialic acids from protective mucins, thereby preventing aggregation and removal via the ciliary ladder83. The presence of influenza virus in the lungs before bacterial incursion can thus facilitate access to the lower respiratory tract by providing a zone in which bacteria that are inhaled from the environment or that are transitioning from a colonizing state in the upper respiratory tract can establish an initial infection81 (Box 3). Second, the host inflammatory response to viral infections can alter the regulatory state and surface display of multiple proteins, including some, such as the platelet activating factor receptor, that can be used to facilitate pneumococcal invasion84,85. Third, changes in the airway during wound healing might provide adherence sites during recovery71,86,87. Injured cells or cells that are in an intermediate differentiation state may express apical receptors, including asialylated glycans (for example, GalNacβ1-4Gal) or α5β1 integrins, to which bacteria such as S. aureus or Pseudomonas aeruginosa can attach (reviewed in Ref. 88). Areas of incomplete healing, where basement membrane elements, such as laminin or type I and type IV collagen, are exposed, or fibrin and fibrinogen deposition have taken place, might also lead to more avid bacterial binding by S. pneumoniae, H. influenzae or S. aureus, mediated by traditional bacterial adhesins. This contributes to the clinical observation that some secondary bacterial infections occur as the patient begins to recover from the primary illness89.
Altered tropism of the virus. Viral polymorphisms that alter the tropism of influenza viruses from the upper to lower respiratory tract facilitate the co-pathogenesis of bacteria in the lungs. Two features of the influenza virus haemagglutinin dictate its tropism within the respiratory tract (Fig. 1a). The first feature is receptor use; influenza viruses can bind to terminal sialic acids that are attached via either an α2,3 or α2,6 linkage90 and, depending on the configuration of the receptor-binding site, specific viruses might bind to one more efficiently than the other. Although both types of sialic acids are present in both the upper and lower respiratory tract, a preference for α2,3-linked receptors is thought to favour deep lung infections owing to poorly understood differences in the prevalence and distribution of the receptors on specific cells types91. Lower respiratory tract infection by the virus then enables deep lung infection by the bacteria by the other mechanisms that are discussed above. Secondly, the glycosylation status of haemagglutinin affects lung tropism. Collagenous lectins in surfactant can bind to haemagglutinin proteins that are highly glycosylated, and thus highly glycosylated viruses are eliminated from the airways via the ciliary ladder, which limits infection to the upper respiratory tract92,93. A poorly glycosylated virus that has an ability to bind to α2,3-linked sialic acid receptors, such as the 1918 pandemic strain, is likely to be most capable of causing deep lung infection and facilitating secondary bacterial pneumonia.
Virus effects on immunity to bacteria. Bacteria such as pneumococci activate, modulate and are eventually controlled by multiple responses of the immune system during invasion of the lungs94,95. Respiratory viruses including influenza virus, RSV, human metapneumovirus and parainfluenzaviruses also manipulate many of these pathways by the expression of multifunctional accessory proteins, such as the influenza virus non-structural protein 1 (NS1) (Refs 96, 97), which potentially interfere with lung immune responses to bacterial invaders42,98,99,100,101 (Box 2). Early innate responses to bacteria have been shown to be compromised by the preceding induction of interferons102,103,104,104,105, which participate in immune sensing and the control of Gram-positive pathogens106,107. This apparently paradoxical finding is mediated by type I interferon, which is produced following the recognition of viral nucleic acids by Toll-like receptors104, and probably functions by suppressing the macrophage and neutrophil responses that normally assist in the clearance of bacteria from the lungs102,105. Inhibition of acute pro-inflammatory cytokines by the impairment of natural killer cell responses108 and the direct suppression of chemokines, which is mediated by the antiviral state promoted by type I interferon105, depress the normal phagocytic activity of macrophages and neutrophils. In addition, influenza viruses specifically deplete the airway-resident alveolar macrophages that are responsible for early bacterial clearance, which leads to a deficit in early bacterial surveillance and killing109 (Fig. 1b). These alveolar macrophages die during the initial stages of infection and are replaced over the next 2 weeks by the proliferation and differentiation of macrophages from other classes, which creates a window of primary susceptibility that extends beyond the immediate viral infection. During the clearance of influenza virus and the onset of wound healing, a general anti-inflammatory state that is orchestrated by systems dedicated to the restoration of lung immune homeostasis110,111 and characterized by increased interleukin-10 (IL-10) production112 broadly suppresses multiple mechanisms that are involved in pathogen recognition and clearance (reviewed in Ref. 113). For example, increased expression of the receptor for the negative regulatory ligand CD200 on myeloid cells during viral infections raises the activation threshold for these cells to superinfecting bacteria, enabling bacterial outgrowth114. Downregulation of other receptors, such as MARCO, on phagocytic cells might impair phagocytosis and killing102. Desensitization of pattern recognition receptors that are used by these phagocytes to detect and respond to bacteria in this setting can persist for weeks or months and can contribute to late secondary infections after apparent recovery from the preceding viral illness115 (Fig. 1b); however, there seem to be virus-specific differences in the longevity of this suppression116. Broadly, it is clear that normal pathogen recognition and effector responses are globally impaired during, and for some time after, influenza (and possibly other) viral infections, although the precise mechanisms are still being elucidated.
Increased inflammation. Pneumonia is an inflammatory condition of the lungs. Therefore, viral and bacterial factors or host responses that increase inflammation in response to the pathogens contribute to the co-pathogenesis of secondary bacterial pneumonia. Some influenza A viruses express a cytotoxic accessory protein, PB1-F2 (Ref. 117), that drives inflammatory responses, which manifest as increased cellular infiltration of the lungs and airways, together with cytokine storm77,78,118 (Fig. 1a). This pro-inflammatory activity is strongly linked to the induction and severity of pneumonia from superinfecting bacteria, as animal models reveal only minor pathological effects in the context of single viral infections but a robust effect on morbidity and mortality during bacterial superinfection33,77,78,79,119. Bacteria also express cytotoxins that contribute to inflammation120,121, such as pneumolysin and PVL, and these might synergize with the effects of PB1-F2 via increased cell death related to pore formation or via increased inflammatory signalling, leading to cytokine storm122 (Fig. 1c). Multiple innate immune mechanisms that involve pattern recognition receptors generate inflammatory responses to respiratory viruses and/or bacteria (reviewed in Refs 94, 123). Many of the inflammatory pathways that are involved overlap, leading to the synergistic activation of immune responses, with resulting morbidity95,96,98,124. For example, viruses such as RSV and influenza virus can activate Toll-like receptor 4 (TLR4), and RSV also activates TLR2 (Refs 125, 126). As these are prominent receptors that are involved in the innate sensing of many Gram-positive bacteria including S. pneumoniae, co-infection or lysis of bacteria in the setting of viral pre-infection can generate inflammation and acute lung injury127. In the setting of depressed phagocytotic activity102,105,108, neutrophil-mediated inflammatory damage may occur in the lungs without effective pathogen control127,128. In concert with these aberrant responses, the virus might 'cripple' wound-repair mechanisms in the lungs, preventing healing and prolonging the period of susceptibility to secondary invasion via the mechanisms discussed above129.
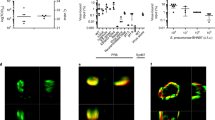
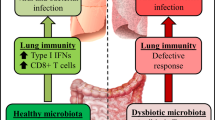
Facilitation of the viral infection. Although most of the mechanisms that are discussed in this Review involve viral facilitation of subsequent bacterial superinfection, it is likely that factors that are expressed by co-infecting bacteria affect the virus as well. Previous studies have indicated that there is a directional effect; that is, synergism occurs when the viral infection precedes bacterial exposure and no effect — or even protection — occurs when the order of challenge is reversed. Bacterial modulation of viral infection may be mediated via direct interactions, via bacterial interference with antiviral immunity or by synergism or complementation by virulence factors that have similar functions. Animal models consistently show an increase in influenza virus titres during bacterial superinfections in which bacterial challenge follows the viral infection; this increased viral lung load is typically accompanied by delayed clearance via unclear mechanisms33,76,119 (Fig. 2). However, viral titres may be suppressed and morbidity may be diminished via the induction of innate immune responses if the bacterial infection precedes viral challenge76,130. It has been proposed that bacterial proteases from S. aureus are capable of cleaving the nascent haemagglutinin from its basal state to a fusion-active complex131, which would increase viral titres and spread. A mostly unexplored area is the effect of bacterial virulence factors on antiviral immunity. Although no data are available, it is reasonable to hypothesize that the bacterial mechanisms that broadly interfere with innate immune mechanisms95,98 might also affect the immune response to respiratory viruses. There is evidence that the composition of the microbiome alters immune responses to influenza both by changing the activation set-point for antiviral responses and by influencing the development of adaptive immune responses132,133. Whether pathogens such as S. pneumoniae and S. aureus have similar effects during colonization is an interesting area for exploration. Finally, there is some limited evidence from animal models that viral and bacterial gene products that have similar functions (for example, viral and bacterial neuraminidases and cytotoxins) might synergize or functionally complement each other to increase growth, avoid immune surveillance or cause additional inflammation and damage33,81,119.
Influenza viruses replicate rapidly when a primary infection is established, reaching a peak titre (blue line) in the lungs 2–3 days after inoculation. Impairment of host defences, including a swift depletion of alveolar macrophages (green line) over the first 3 days of infection enables superinfecting bacteria to grow rapidly (red line) and cause pneumonia. The rapid growth of bacteria is associated with a rebound in viral titre via unclear mechanisms. Failure of secondary immunity to control the co-infection can lead to unchecked bacterial growth after viral clearance, resulting in morbidity and mortality109,153.
Spatiotemporal differences in virulence
The epidemiological data reviewed above, such as the differences in incidence of bacterial superinfections between seasonal and pandemic viruses, suggest that viruses differ in their potential to facilitate bacterial superinfections. All influenza A viruses are zoonoses that originate in the wild bird reservoir and transition through other intermediate hosts before emerging as epidemic or pandemic strains after undergoing gene reassortment in the different host134. Viruses that successfully establish themselves in humans then adapt and evolve over time. Virulence is a multigenic trait in influenza viruses, and one would expect that any features of the viral genome that increase replication, fitness in the lungs or pathogenicity would have downstream effects on co-infections via one or several of the mechanisms described above. However, specific changes that affect co-pathogenesis that are distinct from traditional virulence mechanisms have also been identified in animal models. It is becoming increasingly evident that there are certain gene constellations that favour co-pathogenesis with bacteria, such as low glycosylation of haemagglutinin, an inflammatory PB1-F2 and high neuraminidase activity (Fig. 3). Viruses that have these attributes should be targeted by surveillance and prioritized for study and pandemic preparation119.
Viruses with a gene constellation that is predicted to efficiently support secondary bacterial infections exist in the animal reservoirs of the world119. It can be assumed that, should such a virus cross over into humans and initiate a pandemic, bacterial superinfections would be the primary cause of mortality in a similar manner to the 1918 pandemic. Once established in humans, these viruses adapt to their new hosts, changing over time and losing many of the features that most strongly support co-pathogenesis with bacteria. Many virulence factors change over decades during adaptation as a side effect of negative selection for traits that are associated with increased immune recognition. As examples, N-linked glycans on the globular head of haemagglutinin accumulate to provide immune escape from antibodies in the human population. Highly glycosylated viruses are eliminated via the ciliary ladder after being bound by molecules such as collagenous lectins and neutralized. Poorly glycosylated viruses, such as pandemic strains from the avian reservoir, can evade these protective measures and cause deeper lung infections. The activity of the neuraminidase is in equilibrium with the strength of receptor binding of haemagglutinin and decreases to maintain a balance with the decreased affinity of haemagglutinin related to its adaptation. High-activity neuraminidase strains cleave host sialic acids more effectively, exposing receptors and disrupting the binding of sialylated mucins, which better supports secondary bacterial infections. As this neuraminidase activity is lost, the incidence of secondary bacterial infections decreases. A polymerase complex that provides an appropriate balance, enabling replication and transmission without killing the host, is selected over time. As viral replication efficiency directly relates to virulence and the associated cytotoxicity of the virus, decreased polymerase activity has global effects on the synergistic damage that occurs during co-infections. The cytotoxic and inflammatory PB1-F2 contributes to the severity of bacterial superinfection. Over decades during adaptation, it loses its activity as a result of truncation or mutation via negative selection for a virus that is less likely to trigger a robust immune response. The net effect of these changes in multiple virulence factors, and their functions that support bacterial superinfections, is that well-adapted seasonal strains often lack identifiable features associated with viral–bacterial synergism78,93,154.
Many of the features that support synergism with bacteria are lost or diminished during adaptation and evolution in mammalian hosts, which creates a gradient between seasonal and pandemic strains9. For example, the glycosylation state of haemagglutinin increases over decades of circulation in humans to facilitate escape from immune pressure; this is associated with decreased virulence, a shift in tropism to the upper respiratory tract and diminished support of bacterial lung infections9,93. Similarly, the inflammatory properties of PB1-F2 are lost during adaptation to mammalian lungs, abrogating its contribution to secondary bacterial infections78. Although the timescale for this evolutionary loss of activity is measured in decades, the consistency of the finding suggests that the cytotoxic or pro-inflammatory activity of PB1-F2 is detrimental when expressed in the lungs but not when expressed in the gut — which is the host niche of these viruses in birds — leading to negative selection after crossing species3,78. Generally, as zoonotic viruses that have recently been introduced into humans adapt to their new hosts, they become less virulent, probably via negative selection for traits that stimulate immune responses, and their propensity to support bacterial superinfections decreases. This accounts, in part, for the differing epidemiological patterns of complications and mortality that are associated with seasonal and pandemic strains.
However, it also seems probable that geographic differences in the regional distribution of bacteria can help to determine local epidemiology during influenza epidemics and pandemics135. Although there are currently little data available, one intriguing hypothesis is that specific bacterial virulence factors that contribute to the co-pathogenesis of lung infections with viruses are unevenly distributed across genotypes, such that the local endemicity of bacteria that express these factors influences outcomes9. In a broad sense, this is likely to have occurred with the spread of the USA300 clonotype of S. aureus across North America136; the incidence of serious influenza and S. aureus co-infections in children increased in parallel in the United States with the emergence and dominance of this strain32,137. Animal model data support the idea that there are serotype- and clonotype-specific differences in the ability to participate in co-infections, including a propensity for USA300 strains to cause severe disease together with influenza virus33,119,138. In the United States between 2004 and 2009, specific serotypes of S. pneumoniae seemed to be associated with influenza in humans in an age-related manner: serotype 19A dominated co-infections in children less than 5 years old, 7F dominated in older children and young adults of up to 24 years old, and other serotypes, mostly those not contained in the current 13-valent vaccine, were most common in older adults139. Whether these associations represent baseline prevalence in a highly vaccinated population at the time of the study or whether they are tied to specific virulence factors linked to these serotypes is not clear. Recent data from an influenza virus and H. influenzae mouse model of co-infection suggest that certain virulence factors contribute to pathogenesis only in the context of dual infection140; these genes would not have been found in traditional virulence screens during single-agent infections.
Key questions for the field
The study of viral–bacterial interactions is expanding rapidly with our increased appreciation that most pneumonia results from co-infections. This growing recognition that the interplay between multiple pathogens, the immune system and, more recently, the microbiome is tremendously complex and is breaking down the linear one-pathogen one-disease paradigm that has been accepted since the statement of Koch's postulates. Although much has been done — as reviewed here — to understand the specific mechanisms underlying interactions such as those between influenza viruses, immunity and bacterial respiratory pathogens such as S. pneumoniae, there are key unanswered questions that will drive the field forward over the next decade. Imperatives for the molecular biology laboratory include determining the specific bacterial factors that enable some bacterial strains to take advantage of virus-induced changes in immune pathways and cause disease, understanding the relationship of timing between encounters with pathogens on the relevance of different mechanisms of synergy and discovering host-gene polymorphisms that facilitate co-infections or favour disease. A great deal of the work on novel modes of treatment and prevention must take place in animals before translation to humans. The relevance of the composition of the human and animal microbiome to the acquisition of co-infections and the development of disease, the role of colonization and the relevance of the mechanisms that are discussed in this Review to the upper respiratory tract and the details of the changes to innate and adaptive immunity in the setting of co-infection will be exciting new areas to explore.
However, the most important issues may surround the relevance and relative effect of bench discoveries — which mechanisms that have been discovered in vitro or in animal models are truly important in humans and which have a significant impact on actual epidemiology and pathogenesis? Little work had been done in humans on this area before the 2009 pandemic, and much remains for future study. Questions that can only be answered by large-scale studies that involve consortia or clinical networks include: determining the geographic distribution and diversity as well as the evolution, dispersion and replacement over time of bacteria that express virulence factors that support co-pathogenesis with viruses; developing a set of molecular signatures in viruses and bacteria that promote co-pathogenesis in pneumonia and surveying the animal and human reservoirs for their frequency and distributions to establish a threat level for each; and translating strategies that have been developed at the bench for treatment and prevention into clinical practice. Beyond the laudable goal of increasing our general understanding of the biology that underlies co-infections, the scientific community must help the world to prepare for the next pandemic through focused research so that another pandemic similar to that of 1918 can be averted or managed with limited loss of life.
References
Potter, C. W. in Textbook of Influenza (eds Nicholson, K. G., Webster, R. G. and Hay, A. J.) 3–18 (Blackwell Scientific Publications, 1998).
Morens, D. M., Taubenberger, J. K. & Fauci, A. S. Predominant role of bacterial pneumonia as a cause of death in pandemic influenza: implications for pandemic influenza preparedness. J. Infect. Dis. 198, 962–970 (2008). This paper presents a comprehensive review of the aetiology of fatal cases during the 1918 pandemic.
Smith, A. M. & McCullers, J. A. Molecular signatures of virulence in the PB1-F2 proteins of H5N1 influenza viruses. Virus Res. 178, 146–150 (2013).
Collins, S. D. Influenza–pneumonia mortality in a group of about 95 cities in the United States, 1920–1929. Public Health Rep. 45, 361–406 (1930).
van Asten, L. et al. Mortality attributable to 9 common infections: significant effect of influenza A, respiratory syncytial virus, influenza B, norovirus, and parainfluenza in elderly persons. J. Infect. Dis. 206, 628–639 (2012).
Nolte, F. S. Molecular diagnostics for detection of bacterial and viral pathogens in community-acquired pneumonia. Clin. Infect. Dis. 47, S123–S126 (2008).
Weinberger, D. M. et al. Impact of the 2009 influenza pandemic on pneumococcal pneumonia hospitalizations in the United States. J. Infect. Dis. 205, 458–465 (2012).
Falsey, A. R. et al. Bacterial complications of respiratory tract viral illness: a comprehensive evaluation. J. Infect. Dis. 208, 432–441 (2013). This recent paper documents a high rate of co-infections in a large study of hospitalized patients with pneumonia.
McCullers, J. A. Do specific virus–bacteria pairings drive clinical outcomes of pneumonia? Clin. Microbiol. Infect. 19, 113–118 (2013).
Brundage, J. F. & Shanks, G. D. What really happened during the 1918 influenza pandemic? The importance of bacterial secondary infections. J. Infect. Dis. 196, 1717–1718 (2007).
Chien, Y. W., Klugman, K. P. & Morens, D. M. Bacterial pathogens and death during the 1918 influenza pandemic. N. Engl. J. Med. 361, 2582–2583 (2009).
Brundage, J. F. & Shanks, G. D. Deaths from bacterial pneumonia during 1918–1919 influenza pandemic. Emerg. Infect. Dis. 14, 1193–1199 (2008).
Shanks, G. D. et al. Mortality risk factors during the 1918–1919 influenza pandemic in the Australian army. J. Infect. Dis. 201, 1880–1889 (2010).
Shanks, G. D., MacKenzie, A., Waller, M. & Brundage, J. F. Low but highly variable mortality among nurses and physicians during the influenza pandemic of 1918–1919. Influenza Other Respir. Viruses 5, 213–219 (2011).
Jordan, E. O. in Epidemic Influenza 356–438 (American Medical Association, 1927). This comprehensive and historically important textbook explores all aspects of influenza epidemiology and research in the nineteenth and early twentieth centuries.
Trotter, Y. Jr et al. Asian influenza in the United States, 1957–1958. Am. J. Hyg. 70, 34–50 (1959).
Dauer, C. C. Mortality in the 1957–1958 influenza epidemic. Public Health Rep. 73, 803–810 (1958).
Collins, S. D. & Lehmann, J. Trends and epidemics of influenza and pneumonia: 1918–1951. Public Health Rep. 66, 1487–1516 (1951).
Giles, C. & Shuttleworth, E. M. Postmortem findings in 46 influenza deaths. Lancet 273, 1224–1225 (1957).
Martin, C. M., Kunin, C. M., Gottlieb, L. S. & Finland, M. Asian influenza A in Boston, 1957–1958. II. Severe staphylococcal pneumonia complicating influenza. AMA. Arch. Intern. Med. 103, 532–542 (1959).
Oseasohn, R., Adelson, L. & Kaji, M. Clinicopathologic study of thirty-three fatal cases of Asian influenza. N. Engl. J. Med. 260, 509–518 (1959).
Hers, J. F. P., Masurel, N. & Mulder, J. Bacteriology and histopathology of the respiratory tract and lungs of fatal Asian influenza. Lancet 2, 1164–1165 (1958).
Hers, J. F., Goslings, W. R., Masurel, N. & Mulder, J. Death from Asiatic influenza in the Netherlands. Lancet 273, 1164–1165 (1957).
Robertson, L., Caley, J. P. & Moore, J. Importance of Staphylococcus aureus in pneumonia in the 1957 epidemic of influenza A. Lancet 2, 233–236 (1958).
Herzog, H., Staub, H. & Richterich, R. Gas-analytical studies in severe pneumonia; observations during the 1957 influenza epidemic. Lancet 1, 593–597 (1959).
Jevons, M. P., Taylor, C. E. D. & Williams, R. E. O. Bacteria and influenza. Lancet 273, 835–836 (1957).
Schwarzmann, S. W., Adler, J. L., Sullivan, R. J. Jr. & Marine, W. M. Bacterial pneumonia during the Hong Kong influenza epidemic of 1968–1969. Arch. Intern. Med. 127, 1037–1041 (1971).
Burk, R. F., Schaffner, W. & Koenig, M. G. Severe influenza virus pneumonia in the pandemic of 1968–1969. Arch. Intern. Med. 127, 1122–1128 (1971).
Bisno, A. L., Griffin, J. P., Van Epps, K. A., Niell, H. B. & Rytel, M. W. Pneumonia and Hong Kong influenza: a prospective study of the 1968–1969 epidemic. Am. J. Med. Sci. 261, 251–263 (1971).
Sharrar, R. G. National influenza experience in the USA, 1968–1969. Bull World Health Organ. 41, 361–366 (1969).
Gillet, Y. et al. Association between Staphylococcus aureus strains carrying gene for Panton-Valentine leukocidin and highly lethal necrotising pneumonia in young immunocompetent patients. Lancet 359, 753–759 (2002).
Finelli, L. et al. Influenza-associated pediatric mortality in the United States: increase of Staphylococcus aureus coinfection. Pediatrics 122, 805–811 (2008). This large epidemiological study shows an increase in influenza virus– S. aureus co-infections with the emergence of the USA300 genotype in the United States.
Iverson, A. R. et al. Influenza virus primes mice for pneumonia from Staphylococcus aureus. J. Infect. Dis. 203, 880–888 (2011).
Watt, J. P. et al. Burden of disease caused by Haemophilus influenzae type b in children younger than 5 years: global estimates. Lancet 374, 903–911 (2009).
Jennings, L. C. et al. Incidence and characteristics of viral community-acquired pneumonia in adults. Thorax 63, 42–48 (2008).
Chaussee, M. S. et al. Inactivated and live, attenuated influenza vaccines protect mice against influenza: Streptococcus pyogenes super-infections. Vaccine 29, 3773–3781 (2011).
Michelow, I. C. et al. Epidemiology and clinical characteristics of community-acquired pneumonia in hospitalized children. Pediatrics 113, 701–707 (2004).
Berkley, J. A. et al. Viral etiology of severe pneumonia among Kenyan infants and children. JAMA 303, 2051–2057 (2010).
Olsen, S. J. et al. Incidence of respiratory pathogens in persons hospitalized with pneumonia in two provinces in Thailand. Epidemiol. Infect. 138, 1811–1822 (2010).
Hammitt, L. L. et al. A preliminary study of pneumonia etiology among hospitalized children in Kenya. Clin. Infect. Dis. 54, S190–S199 (2012).
Chen, C. J. et al. Etiology of community-acquired pneumonia in hospitalized children in northern Taiwan. Pediatr. Infect. Dis. J. 31, e196–e201 (2012).
Techasaensiri, B. et al. Viral coinfections in children with invasive pneumococcal disease. Pediatr. Infect. Dis. J. 29, 519–523 (2010).
Peltola, V. et al. Temporal association between rhinovirus circulation in the community and invasive pneumococcal disease in children. Pediatr. Infect. Dis. J. 30, 456–461 (2011).
Dawood, F. S. et al. Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N. Engl. J. Med. 360, 2605–2615 (2009).
Dawood, F. S. et al. Estimated global mortality associated with the first 12 months of 2009 pandemic influenza A H1N1 virus circulation: a modelling study. Lancet Infect. Dis. 12, 687–695 (2012).
Reichert, T., Chowell, G., Nishiura, H., Christensen, R. A. & McCullers, J. A. Does glycosylation as a modifier of Original Antigenic Sin explain the case age distribution and unusual toxicity in pandemic novel H1N1 influenza? BMC Infect. Dis. 10, 5 (2010).
Monsalvo, A. C. et al. Severe pandemic 2009 H1N1 influenza disease due to pathogenic immune complexes. Nature Med. 17, 195–199 (2011).
Charu, V. et al. Mortality burden of the A/H1N1 pandemic in Mexico: a comparison of deaths and years of life lost to seasonal influenza. Clin. Infect. Dis. 53, 985–993 (2011).
Simonsen, L. et al. The impact of influenza epidemics on mortality: introducing a severity index. Am. J. Publ. Health 87, 1944–1950 (1997).
Dominguez-Cherit, G. et al. Critically ill patients with 2009 influenza A (H1N1) in Mexico. JAMA 302, 1880–1887 (2009).
Centers for Disease Control Bacterial coinfections in lung tissue specimens from fatal cases of 2009 pandemic influenza A (H1N1) — United States, May–August 2009. Morb. Mortal. Wkly Rep. 58, 1071–1074 (2009).
Mauad, T. et al. Lung pathology in fatal novel human influenza A (H1N1) infection. Am. J. Respir. Crit. Care Med. 181, 72–79 (2010).
Estenssoro, E. et al. Pandemic 2009 influenza A in Argentina: a study of 337 patients on mechanical ventilation. Am. J. Respir. Crit. Care Med. 182, 41–48 (2010).
Shieh, W. J. et al. 2009 pandemic influenza A (H1N1): pathology and pathogenesis of 100 fatal cases in the United States. Am. J. Pathol. 177, 166–175 (2010). This paper comprehensively describes the pathology of co-infections during the 2009 pandemic.
Rice, T. W. et al. Critical illness from 2009 pandemic influenza A virus and bacterial coinfection in the United States. Crit. Care Med. 40, 1487–1498 (2012).
Nguyen, T. et al. Coinfection with Staphylococcus aureus increases risk of severe coagulopathy in critically ill children with influenza A (H1N1) virus infection. Crit. Care Med. 40, 3246–3250 (2012).
Nelson, J. C. et al. Impact of the introduction of pneumococcal conjugate vaccine on rates of community acquired pneumonia in children and adults. Vaccine 26, 4947–4954 (2008).
Watkins, R. R., David, M. Z. & Salata, R. A. Current concepts on the virulence mechanisms of meticillin-resistant Staphylococcus aureus. J. Med. Microbiol. 61, 1179–1193 (2012).
Ampofo, K. et al. Association of 2009 pandemic influenza A (H1N1) infection and increased hospitalization with parapneumonic empyema in children in Utah. Pediatr. Infect. Dis. J. 29, 905–909 (2010).
Kumar, S. et al. Clinical and epidemiologic characteristics of children hospitalized with 2009 pandemic H1N1 influenza A infection. Pediatr. Infect. Dis. J. 29, 591–594 (2010).
Palacios, G. et al. Streptococcus pneumoniae coinfection is correlated with the severity of H1N1 pandemic influenza. PLoS ONE 4, e8540 (2009). This paper shows a link between co-infections and more severe outcomes.
Dhanoa, A., Fang, N. C., Hassan, S. S., Kaniappan, P. & Rajasekaram, G. Epidemiology and clinical characteristics of hospitalized patients with pandemic influenza A (H1N1) 2009 infections: the effects of bacterial coinfection. Virol. J. 8, 501 (2011).
McCullers, J. A. Insights into the interaction between influenza virus and pneumococcus. Clin. Microbiol. Rev. 19, 571–582 (2006).
McCullers, J. A. Preventing and treating secondary bacterial infections with antiviral agents. Antivir. Ther. 16, 123–135 (2011). This paper reviews the mechanisms and our current knowledge of related treatment modalities for viral–bacterial co-infections.
Loosli, C. G. et al. The destruction of type 2 pneumocytes by airborne influenza PR8-A virus; its effect on surfactant and lecithin content of the pneumonic lesions of mice. Chest 67, 7S–14S (1975).
Johanson, W. G. Jr., Pierce, A. K. & Sanford, J. P. Pulmonary function in uncomplicated influenza. Am. Rev. Respir. Dis. 100, 141–146 (1969).
Horner, G. J. & Gray, F. D. Jr. Effect of uncomplicated, presumptive influenza on the diffusing capacity of the lung. Am. Rev. Respir. Dis. 108, 866–869 (1973).
Levandowski, R. A., Gerrity, T. R. & Garrard, C. S. Modifications of lung clearance mechanisms by acute influenza A infection. J. Lab. Clin. Med. 106, 428–432 (1985).
Glezen, W. P., Greenberg, S. B., Atmar, R. L., Piedra, P. A. & Couch, R. B. Impact of respiratory virus infections on persons with chronic underlying conditions. JAMA 283, 499–505 (2000).
Plotkowski, M. C., Puchelle, E., Beck, G., Jacquot, J. & Hannoun, C. Adherence of type I Streptococcus pneumoniae to tracheal epithelium of mice infected with influenza A/PR8 virus. Am. Rev. Respir. Dis. 134, 1040–1044 (1986). This paper is a classic study that shows bacterial adherence in areas of virus-mediated epithelial damage.
Plotkowski, M. C., Bajolet-Laudinat, O. & Puchelle, E. Cellular and molecular mechanisms of bacterial adhesion to respiratory mucosa. Eur. Respir. J. 6, 903–916 (1993).
Dockrell, D. H., Whyte, M. K. & Mitchell, T. J. Pneumococcal pneumonia: mechanisms of infection and resolution. Chest 142, 482–491 (2012).
Heilmann, C. Adhesion mechanisms of staphylococci. Adv. Exp. Med. Biol. 715, 105–123 (2011).
Foster, T. J. & Hook, M. Surface protein adhesins of Staphylococcus aureus. Trends Microbiol. 6, 484–488 (1998).
McCullers, J. A. & Tuomanen, E. I. Molecular pathogenesis of pneumococcal pneumonia. Front. Biosci. 6, D877–D889 (2001).
McCullers, J. A. & Rehg, J. E. Lethal synergism between influenza virus and Streptococcus pneumoniae: characterization of a mouse model and the role of platelet-activating factor receptor. J. Infect. Dis. 186, 341–350 (2002).
McAuley, J. L. et al. Expression of the 1918 influenza A virus PB1-F2 enhances the pathogenesis of viral and secondary bacterial pneumonia. Cell Host Microbe 2, 240–249 (2007).
McAuley, J. L. et al. PB1-F2 proteins from H5N1 and 20th century pandemic influenza viruses cause immunopathology. PLoS Pathog. 6, e1001014 (2010).
Alymova, I. V. et al. Immunopathogenic and antibacterial effects of H3N2 influenza A virus PB1-F2 map to amino acid residues 62, 75, 79, and 82. J. Virol. 85, 12324–12333 (2011).
Guarner, J. & Falcon-Escobedo, R. Comparison of the pathology caused by H1N1, H5N1, and H3N2 influenza viruses. Arch. Med. Res. 40, 655–661 (2009).
McCullers, J. A. & Bartmess, K. C. Role of neuraminidase in lethal synergism between influenza virus and Streptococcus pneumoniae. J. Infect. Dis. 187, 1000–1009 (2003).
McCullers, J. A. Effect of antiviral treatment on the outcome of secondary bacterial pneumonia after influenza. J. Infect. Dis. 190, 519–526 (2004).
Camara, M., Mitchell, T. J., Andrew, P. W. & Boulnois, G. J. Streptococcus pneumoniae produces at least two distinct enzymes with neuraminidase activity: cloning and expression of a second neuraminidase gene in Escherichia coli. Infect. Immun. 59, 2856–2858 (1991).
Cundell, D. R. & Tuomanen, E. I. Receptor specificity of adherence of Streptococcus pneumoniae to human type-II pneumocytes and vascular endothelial cells in vitro. Microb. Pathog. 17, 361–374 (1994).
Miller, M. L., Gao, G., Pestina, T., Persons, D. & Tuomanen, E. Hypersusceptibility to invasive pneumococcal infection in experimental sickle cell disease involves platelet-activating factor receptor. J. Infect. Dis. 195, 581–584 (2007).
de Bentzmann, S., Plotkowski, C. & Puchelle, E. Receptors in the Pseudomonas aeruginosa adherence to injured and repairing airway epithelium. Am. J. Respir. Crit. Care Med. 154, S155–S162 (1996).
Martin, P. & Leibovich, S. J. Inflammatory cells during wound repair: the good, the bad and the ugly. Trends Cell Biol. 15, 599–607 (2005).
Puchelle, E., Zahm, J. M., Tournier, J. M. & Coraux, C. Airway epithelial repair, regeneration, and remodeling after injury in chronic obstructive pulmonary disease. Proc. Am. Thorac. Soc. 3, 726–733 (2006).
Louria, D., Blumenfeld, H., Ellis, J., Kilbourne, E. D. & Rogers, D. Studies on influenza in the pandemic of 1957–1958. II. Pulmonary complications of influenza. J. Clin. Invest. 38, 213–265 (1959).
Webster, R. G., Bean, W. J., Gorman, O. T., Chambers, T. M. & Kawaoka, Y. Evolution and ecology of influenza A viruses. Microbiol. Rev. 56, 152–179 (1992).
Chutinimitkul, S. et al. In vitro assessment of attachment pattern and replication efficiency of H5N1 influenza A viruses with altered receptor specificity. J. Virol. 84, 6825–6833 (2010).
Reading, P. C., Morey, L. S., Crouch, E. C. & Anders, E. M. Collectin-mediated antiviral host defense of the lung: evidence from influenza virus infection of mice. J. Virol. 71, 8204–8212 (1997).
Vigerust, D. J. et al. N-linked glycosylation attenuates H3N2 influenza viruses. J. Virol. 81, 8593–8600 (2007).
Koppe, U., Suttorp, N. & Opitz, B. Recognition of Streptococcus pneumoniae by the innate immune system. Cell. Microbiol. 14, 460–466 (2012).
Joyce, E. A., Popper, S. J. & Falkow, S. Streptococcus pneumoniae nasopharyngeal colonization induces type I interferons and interferon-induced gene expression. BMC Genomics 10, 404 (2009).
Bucasas, K. L. et al. Global gene expression profiling in infants with acute respiratory syncytial virus broncholitis demonstrates systemic activation of interferon signaling networks. Pediatr. Infect. Dis. J. 32, e68–e76 (2013).
Hale, B. G., Randall, R. E., Ortin, J. & Jackson, D. The multifunctional NS1 protein of influenza A viruses. J. Gen. Virol. 89, 2359–2376 (2008).
Kimaro Mlacha, S. Z. et al. Transcriptional adaptation of pneumococci and human pharyngeal cells in the presence of a virus infection. BMC Genomics 14, 378 (2013).
Kukavica-Ibrulj, I. et al. Infection with human metapneumovirus predisposes mice to severe pneumococcal pneumonia. J. Virol. 83, 1341–1349 (2009).
Stark, J. M., Stark, M. A., Colasurdo, G. N. & LeVine, A. M. Decreased bacterial clearance from the lungs of mice following primary respiratory syncytial virus infection. J. Med. Virol. 78, 829–838 (2006).
Alymova, I. V. et al. The novel parainfluenza virus hemagglutinin-neuraminidase inhibitor BCX 2798 prevents lethal synergism between a paramyxovirus and Streptococcus pneumoniae. Antimicrob. Agents Chemother. 49, 398–405 (2005).
Sun, K. & Metzger, D. W. Inhibition of pulmonary antibacterial defense by interferon-γ during recovery from influenza infection. Nature Med. 14, 558–564 (2008).
Li, W., Moltedo, B. & Moran, T. M. Type I interferon induction during influenza virus infection increases susceptibility to secondary Streptococcus pneumoniae infection by negative regulation of γδ T cells. J. Virol. 86, 12304–12312 (2012).
Tian, X. et al. Poly I:C enhances susceptibility to secondary pulmonary infections by Gram-positive bacteria. PLoS ONE 7, e41879 (2012).
Shahangian, A. et al. Type I IFNs mediate development of postinfluenza bacterial pneumonia in mice. J. Clin. Invest. 119, 1910–1920 (2009).
Koppe, U. et al. Streptococcus pneumoniae stimulates a STING- and IFN regulatory factor 3-dependent type I IFN production in macrophages, which regulates RANTES production in macrophages, cocultured alveolar epithelial cells, and mouse lungs. J. Immunol. 188, 811–817 (2012).
Parker, D. et al. Streptococcus pneumoniae DNA initiates type I interferon signaling in the respiratory tract. mBio 2, e00016–00011 (2011).
Small, C. L. et al. Influenza infection leads to increased susceptibility to subsequent bacterial superinfection by impairing NK cell responses in the lung. J. Immunol. 184, 2048–2056 (2010).
Ghoneim, H. E., Thomas, P. G. & McCullers, J. A. Depletion of alveolar macrophages during influenza infection facilitates bacterial superinfections. J. Immunol. 191, 1250–1259 (2013).
Snelgrove, R. J. et al. A critical function for CD200 in lung immune homeostasis and the severity of influenza infection. Nature Immunol. 9, 1074–1083 (2008).
Hussell, T. & Cavanagh, M. M. The innate immune rheostat: influence on lung inflammatory disease and secondary bacterial pneumonia. Biochem. Soc. Trans. 37, 811–813 (2009).
van der Sluijs, K. F. et al. IL-10 is an important mediator of the enhanced susceptibility to pneumococcal pneumonia after influenza infection. J. Immunol. 172, 7603–7609 (2004).
Metzger, D. W. & Sun, K. Immune dysfunction and bacterial coinfections following influenza. J. Immunol. 191, 2047–2052 (2013).
Goulding, J. et al. Lowering the threshold of lung innate immune cell activation alters susceptibility to secondary bacterial superinfection. J. Infect. Dis. 204, 1086–1094 (2011).
Didierlaurent, A. et al. Sustained desensitization to bacterial Toll-like receptor ligands after resolution of respiratory influenza infection. J. Exp. Med. 205, 323–329 (2008). This paper shows that innate immune defects persist for months after influenza in mice.
Ludewick, H. P., Aerts, L., Hamelin, M. E. & Boivin, G. Long-term impairment of Streptococcus pneumoniae lung clearance is observed after initial infection with influenza A virus but not human metapneumovirus in mice. J. Gen. Virol. 92, 1662–1665 (2011).
Chen, W. et al. A novel influenza A virus mitochondrial protein that induces cell death. Nature Med. 7, 1306–1312 (2001).
Conenello, G. M., Zamarin, D., Perrone, L. A., Tumpey, T. & Palese, P. A single mutation in the PB1-F2 of H5N1 (HK/97) and 1918 influenza A viruses contributes to increased virulence. PLoS Pathog. 3, 1414–1421 (2007).
Weeks-Gorospe, J. N. et al. Naturally occurring swine influenza A virus PB1-F2 phenotypes that contribute to superinfection with Gram-positive respiratory pathogens. J. Virol. 86, 9035–9043 (2012).
Tuomanen, E. I., Austrian, R. & Masure, H. R. Pathogenesis of pneumococcal infection. N. Engl. J. Med. 332, 1280–1284 (1995).
Rogolsky, M. Nonenteric toxins of Staphylococcus aureus. Microbiol. Rev. 43, 320–360 (1979).
Boulnois, G. J., Paton, J. C., Mitchell, T. J. & Andrew, P. W. Structure and function of pneumolysin, the multifunctional, thiol-activated toxin of Streptococcus pneumoniae. Mol. Microbiol. 5, 2611–2616 (1991).
Ramos, I. & Fernandez-Sesma, A. Cell receptors for influenza A viruses and the innate immune response. Front. Microbiol. 3, 117 (2012).
Kuri, T. et al. Influenza A virus-mediated priming enhances cytokine secretion by human dendritic cells infected with Streptococcus pneumoniae. Cell. Microbiol. 15, 1385–1400 (2013).
Imai, Y. et al. Identification of oxidative stress and Toll-like receptor 4 signaling as a key pathway of acute lung injury. Cell 133, 235–249 (2008).
Klein, K. P., Tan, L., Werkman, W., van Bleek, G. M. & Coenjaerts, F. The role of Toll-like receptors in regulating the immune response against respiratory syncytial virus. Crit. Rev. Immunol. 29, 531–550 (2009).
Karlstrom, A., Heston, S. M., Boyd, K. L., Tuomanen, E. I. & McCullers, J. A. Toll-like receptor 2 mediates fatal immunopathology in mice during treatment of secondary pneumococcal pneumonia following influenza. J. Infect. Dis. 204, 1358–1366 (2011).
Weiss, S. J. Tissue destruction by neutrophils. N. Engl. J. Med. 320, 365–376 (1989).
Kash, J. C. et al. Lethal synergism of 2009 pandemic H1N1 influenza virus and Streptococcus pneumoniae coinfection is associated with loss of murine lung repair responses. mBio 2, e00172 (2011).
Wang, J. et al. Bacterial colonization dampens influenza-mediated acute lung injury via induction of M2 alveolar macrophages. Nature Commun. 4, 2106 (2013).
Tashiro, M., Ciborowski, P., Klenk, H. D., Pulverer, G. & Rott, R. Role of Staphylococcus protease in the development of influenza pneumonia. Nature 325, 536–537 (1987).
Ichinohe, T. et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl Acad. Sci. USA 108, 5354–5359 (2011).
Abt, M. C. et al. Commensal bacteria calibrate the activation threshold of innate antiviral immunity. Immunity 37, 158–170 (2012).
Kasowski, E. J., Garten, R. J. & Bridges, C. B. Influenza pandemic epidemiologic and virologic diversity: reminding ourselves of the possibilities. Clin. Infect. Dis. 52, S44–S49 (2011).
Klugman, K. P., Chien, Y. W. & Madhi, S. A. Pneumococcal pneumonia and influenza: a deadly combination. Vaccine 27, C9–C14 (2009).
Mediavilla, J. R., Chen, L., Mathema, B. & Kreiswirth, B. N. Global epidemiology of community-associated methicillin resistant Staphylococcus aureus (CA-MRSA). Curr. Opin. Microbiol. 15, 588–595 (2012).
Peebles, P. J., Dhara, R., Brammer, L., Fry, A. M. & Finelli, L. Influenza-associated mortality among children — United States: 2007–2008. Influenza Other Respir. Viruses 5, 25–31 (2011).
McCullers, J. A. et al. Influenza enhances susceptibility to natural acquisition of and disease due to Streptococcus pneumoniae in ferrets. J. Infect. Dis. 202, 1287–1295 (2010).
Fleming-Dutra, K. E. et al. Effect of the 2009 influenza A (H1N1) pandemic on invasive pneumococcal pneumonia. J. Infect. Dis. 207, 1135–1143 (2013). This paper presents the first large study associating specific pneumococcal serotypes with influenza.
Wong, S. M., Bernui, M., Shen, H. & Akerley, B. J. Genome-wide fitness profiling reveals adaptations required by Haemophilus in coinfection with influenza A virus in the murine lung. Proc. Natl Acad. Sci. USA 110, 15413–15418 (2013).
Morens, D. M. & Taubenberger, J. K. Pandemic influenza: certain uncertainties. Rev. Med. Virol. 21, 262–284 (2011).
Laennec, R. T. H. in Translation of Selected Passages from De l'Auscultation Mediate 88–95 (Williams Wood & Co., 1923).
Wherry, W. B. & Butterfield, C. T. Inhalation experiments on influenza and pneumonia, and on the importance of spray-borne bacteria in respiratory infections. J. Infect. Dis. 27, 315–326 (1920).
Shope, R. E. Swine influenza. I. Experimental transmission and pathology. II. A hemophilic bacillus from the respiratory tract of infected swine. III. Filtration experiments and etiology. J. Exp. Med. 54, 349–385 (1931). This is a classic paper showing that the manifestation of disease during viral infection requires a co-infecting pathogen.
Francis, T. E. & de Torregrosa, M. V. Combined infection of mice with H. influenzae and influenza virus by the intranasal route. J. Infect. Dis. 76, 70–77 (1945).
Avadhanula, V. et al. Respiratory viruses augment the adhesion of bacterial pathogens to respiratory epithelium in a viral species- and cell type-dependent manner. J. Virol. 80, 1629–1636 (2006).
Vissers, M. et al. Respiratory syncytial virus infection augments NOD2 signaling in an IFN-β-dependent manner in human primary cells. Eur. J. Immunol. 42, 2727–2735 (2012).
Ren, J. et al. Human metapneumovirus inhibits IFN-β signaling by downregulating Jak1 and Tyk2 cellular levels. PLoS ONE 6, e24496 (2011).
Radin, J. N. et al. β-Arrestin 1 participates in platelet-activating factor receptor-mediated endocytosis of Streptococcus pneumoniae. Infect. Immun. 73, 7827–7835 (2005).
Vernatter, J. & Pirofski, L. A. Current concepts in host–microbe interaction leading to pneumococcal pneumonia. Curr. Opin. Infect. Dis. 26, 277–283 (2013).
Marks, L. R., Davidson, B. A., Knight, P. R. & Hakansson, A. P. Interkingdom signaling induces Streptococcus pneumoniae biofilm dispersion and transition from asymptomatic colonization to disease. mBio 4, e00438 (2013).
Shrestha, S. et al. Identifying the interaction between influenza and pneumococcal pneumonia using incidence data. Sci. Transl. Med. 5, 191ra84 (2013).
Smith, A. M. et al. Kinetics of coinfection with influenza A virus and Streptococcus pneumoniae. PLoS Pathog. 9, e1003238 (2013).
Peltola, V. T., Murti, K. G. & McCullers, J. A. Influenza virus neuraminidase contributes to secondary bacterial pneumonia. J. Infect. Dis. 192, 249–257 (2005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Glossary
- Influenza pandemic
-
A worldwide and severe influenza epidemic caused by a virus with a novel haemagglutinin.
- Clonotypes
-
Groups of staphylococcal strains that share similar characteristics, as distinguished by pulsed-field gel electrophoresis.
- Dead space
-
An area of lung that has a mismatch between aeration and perfusion, such that gas exchange cannot take place.
- Cytokine storm
-
A state characterized by cytokine levels that are higher than physiological norms; it is associated with inflammation and organ dysfunction.
- α2,3 or α2,6 linkage
-
Linkages between terminal sialic acids and galactose on host cells; influenza viruses preferentially attach to different sialic acids depending on linkage type.
- Intermediate hosts
-
Hosts such as pigs or domestic poultry, in which a virus first adapts before successfully jumping to humans.
- Reassortment
-
A process by which segmented viruses, such as influenza, can swap genes with one another while co-infecting a cell.
- Serotypes
-
Groups of pneumococcal strains that are distinguished by the antigenicity of their capsules.
- Molecular signatures
-
Specific amino acids that are associated with virulence and co-pathogenesis with bacteria.
Rights and permissions
About this article
Cite this article
McCullers, J. The co-pathogenesis of influenza viruses with bacteria in the lung. Nat Rev Microbiol 12, 252–262 (2014). https://doi.org/10.1038/nrmicro3231
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3231
This article is cited by
-
Intracranial complications of sinogenic and otogenic infections in children: an ESPN survey on their occurrence in the pre-COVID and post-COVID era
Child's Nervous System (2024)
-
Staphylococcus epidermidis induced toxic shock syndrome (TSS) secondary to influenza infection
BMC Infectious Diseases (2023)
-
Targeting complement hyperactivation: a novel therapeutic approach for severe pneumonia induced by influenza virus/staphylococcus aureus coinfection
Signal Transduction and Targeted Therapy (2023)
-
Human Identical Sequences, hyaluronan, and hymecromone ─ the new mechanism and management of COVID-19
Molecular Biomedicine (2022)
-
N-glycosylation, a leading role in viral infection and immunity development
Molecular Biology Reports (2022)