Key Points
-
Phages and bacteria are seemingly engaged in an endless battle. In the presence of bacterial antiviral barriers, phages rely on several diverse counter-strategies to usurp their hosts and ensure phage propagation.
-
Phages can readily modify their receptor-binding protein to adsorb to evolving bacterial populations. Phages can also use strategies such as diversity-generating retroelements to facilitate the recognition of multiple bacterial receptors. Moreover, if host receptors are masked by capsular polysaccharides, some phages possess degrading enzymes to hydrolyse this material and access the receptor.
-
When the phage genome enters its bacterial host, it can be targeted by various nucleic acid cleavage enzymes such as restriction–modification and CRISPR–Cas (clustered regularly interspaced short palindromic repeats–CRISPR-associated proteins) systems. To protect against these systems, phages use an array of strategies, including anti-restriction–modification and anti-CRISPR systems, and point mutations in specific sequences.
-
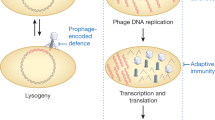
Abortive-infection mechanisms lead to the altruistic death of a phage-infected cell (for example, through induction of a toxin), thereby limiting phage propagation. Phages circumvent this hurdle mainly by mutating specific genes or through the production or hijacking of antitoxins that neutralize the bacterial toxin.
-
This is a rapidly expanding field of research, but many antiphage systems and many strategies used to escape these bacterial defences remain to be discovered. A better understanding of phage–host interactions is also needed to minimize the negative impact of phages on food production and biotechnology applications, as well as to maximize the use of phages as antibacterial agents.
Abstract
Bacteria and their viral predators (bacteriophages) are locked in a constant battle. In order to proliferate in phage-rich environments, bacteria have an impressive arsenal of defence mechanisms, and in response, phages have evolved counter-strategies to evade these antiviral systems. In this Review, we describe the various tactics that are used by phages to overcome bacterial resistance mechanisms, including adsorption inhibition, restriction–modification, CRISPR–Cas (clustered regularly interspaced short palindromic repeats–CRISPR-associated proteins) systems and abortive infection. Furthermore, we consider how these observations have enhanced our knowledge of phage biology, evolution and phage–host interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brüssow, H. & Hendrix, R. W. Phage genomics: small is beautiful. Cell 108, 13–16 (2002).
Labrie, S. J., Samson, J. E. & Moineau, S. Bacteriophage resistance mechanisms. Nature Rev. Microbiol. 8, 317–327 (2010). An exhaustive review on bacterial antiphage mechanisms.
Suttle, C. A. Marine viruses — major players in the global ecosystem. Nature Rev. Microbiol. 5, 801–812 (2007).
Moldovan, R., Chapman-McQuiston, E. & Wu, X. L. On kinetics of phage adsorption. Biophys. J. 93, 303–315 (2007).
Bertin, A., de Frutos, M. & Letellier, L. Bacteriophage–host interactions leading to genome internalization. Curr. Opin. Microbiol. 14, 492–496 (2011).
Rakhuba, D. V., Kolomiets, E. I., Dey, E. S. & Novik, G. I. Bacteriophage receptors, mechanisms of phage adsorption and penetration into host cell. Pol. J. Microbiol. 59, 145–155 (2010).
Rothenberg, E. et al. Single-virus tracking reveals a spatial receptor-dependent search mechanism. Biophys. J. 100, 2875–2882 (2011).
Sorensen, M. C. et al. Phase variable expression of capsular polysaccharide modifications allows Campylobacter jejuni to avoid bacteriophage infection in chickens. Front. Cell. Infect. Microbiol. 2, 11 (2012).
Kim, M. & Ryu, S. Spontaneous and transient defence against bacteriophage by phase-variable glucosylation of O-antigen in Salmonella enterica serovar Typhimurium. Mol. Microbiol. 86, 411–425 (2012).
Hoyland-Kroghsbo, N. M., Maerkedahl, R. B. & Svenningsen, S. L. A quorum-sensing-induced bacteriophage defense mechanism. mBio 4, e00362-12 (2013).
Chatterjee, S. & Rothenberg, E. Interaction of bacteriophage l with its E. coli receptor, LamB. Viruses 4, 3162–3178 (2012).
Meyer, J. R. et al. Repeatability and contingency in the evolution of a key innovation in phage lambda. Science 335, 428–432 (2012). An excellent study of phage λ evolution, showing that when faced with different bacterial receptors, phages can evolve the ability to recognize the new receptors.
Michel, A., Clermont, O., Denamur, E. & Tenaillon, O. Bacteriophage PhiX174's ecological niche and the flexibility of its Escherichia coli lipopolysaccharide receptor. Appl. Environ. Microbiol. 76, 7310–7313 (2010).
Pepin, K. M., Domsic, J. & McKenna, R. Genomic evolution in a virus under specific selection for host recognition. Infect. Genet. Evol. 8, 825–834 (2008).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006).
Qimron, U., Marintcheva, B., Tabor, S. & Richardson, C. C. Genomewide screens for Escherichia coli genes affecting growth of T7 bacteriophage. Proc. Natl Acad. Sci. USA 103, 19039–19044 (2006).
Ravin, V., Raisanen, L. & Alatossava, T. A conserved C-terminal region in Gp71 of the small isometric-head phage LL-H and ORF474 of the prolate-head phage JCL1032 is implicated in specificity of adsorption of phage to its host, Lactobacillus delbrueckii. J. Bacteriol. 184, 2455–2459 (2002).
Avrani, S., Wurtzel, O., Sharon, I., Sorek, R. & Lindell, D. Genomic island variability facilitates Prochlorococcus–virus coexistence. Nature 474, 604–608 (2011).
Scanlan, P. D., Hall, A. R., Lopez-Pascua, L. D. & Buckling, A. Genetic basis of infectivity evolution in a bacteriophage. Mol. Ecol. 20, 981–989 (2011).
Paterson, S. et al. Antagonistic coevolution accelerates molecular evolution. Nature 464, 275–278 (2010).
Scholl, D., Adhya, S. & Merril, C. Escherichia coli K1's capsule is a barrier to bacteriophage T7. Appl. Environ. Microbiol. 71, 4872–4874 (2005).
Leiman, P. G. et al. The structures of bacteriophages K1E and K1-5 explain processive degradation of polysaccharide capsules and evolution of new host specificities. J. Mol. Biol. 371, 836–849 (2007).
Stummeyer, K. et al. Evolution of bacteriophages infecting encapsulated bacteria: lessons from Escherichia coli K1-specific phages. Mol. Microbiol. 60, 1123–1135 (2006).
Baker, J. R., Dong, S. & Pritchard, D. G. The hyaluronan lyase of Streptococcus pyogenes bacteriophage H4489A. Biochem. J. 365, 317–322 (2002).
Cornelissen, A. et al. Identification of EPS-degrading activity within the tail spikes of the novel Pseudomonas putida phage AF. Virology 434, 251–256 (2012).
Glonti, T., Chanishvili, N. & Taylor, P. W. Bacteriophage-derived enzyme that depolymerizes the alginic acid capsule associated with cystic fibrosis isolates of Pseudomonas aeruginosa. J. Appl. Microbiol. 108, 695–702 (2010).
Lampson, B. C., Inouye, M. & Inouye, S. Retrons, msDNA, and the bacterial genome. Cytogenet. Genome Res. 110, 491–499 (2005).
Liu, M. et al. Reverse transcriptase-mediated tropism switching in Bordetella bacteriophage. Science 295, 2091–2094 (2002).
Doulatov, S. et al. Tropism switching in Bordetella bacteriophage defines a family of diversity-generating retroelements. Nature 431, 476–481 (2004).
Medhekar, B. & Miller, J. F. Diversity-generating retroelements. Curr. Opin. Microbiol. 10, 388–395 (2007).
Stockdale, S. R. et al. The lactococcal phages Tuc2009 and TP901-1 incorporate two alternate forms of their tail fiber into their virions for infection specialization. J. Biol. Chem. 288, 5581–5590 (2013). The first example of lactococcal proteolytic processing of a tail fibre, which results in a heterogeneous population consisting of two phage types, leading to increased host adsorption efficiency.
Tetart, F., Repoila, F., Monod, C. & Krisch, H. M. Bacteriophage T4 host range is expanded by duplications of a small domain of the tail fiber adhesin. J. Mol. Biol. 258, 726–731 (1996).
Lin, T. Y. et al. A T3 and T7 recombinant phage acquires efficient adsorption and a broader host range. PLoS ONE 7, e30954 (2012).
Schwarzer, D. et al. A multivalent adsorption apparatus explains the broad host range of phage phi92: a comprehensive genomic and structural analysis. J. Virol. 86, 10384–10398 (2012).
Roberts, R. J. et al. A nomenclature for restriction enzymes, DNA methyltransferases, homing endonucleases and their genes. Nucleic Acids Res. 31, 1805–1812 (2003).
Tock, M. R. & Dryden, D. T. The biology of restriction and anti-restriction. Curr. Opin. Microbiol. 8, 466–472 (2005).
Murray, N. E. Fred Griffith review lecture. Immigration control of DNA in bacteria: self versus non-self. Microbiology 148, 3–20 (2001).
Lacks, S. & Greenberg, B. A deoxyribonuclease of Diplococcus pneumoniae specific for methylated DNA. J. Biol. Chem. 250, 4060–4066 (1975).
Kruger, D. H. & Bickle, T. A. Bacteriophage survival: multiple mechanisms for avoiding the deoxyribonucleic acid restriction systems of their hosts. Microbiol. Rev. 47, 345–360 (1983).
Bickle, T. A. & Kruger, D. H. Biology of DNA restriction. Microbiol. Rev. 57, 434–450 (1993).
Kruger, D. H., Barcak, G. J., Reuter, M. & Smith, H. O. EcoRII can be activated to cleave refractory DNA recognition sites. Nucleic Acids Res. 16, 3997–4008 (1988).
Meisel, A., Bickle, T. A., Kruger, D. H. & Schroeder, C. Type III restriction enzymes need two inversely oriented recognition sites for DNA cleavage. Nature 355, 467–469 (1992).
Warren, R. A. Modified bases in bacteriophage DNAs. Annu. Rev. Microbiol. 34, 137–158 (1980).
Bair, C. L. & Black, L. W. A type IV modification dependent restriction nuclease that targets glucosylated hydroxymethyl cytosine modified DNAs. J. Mol. Biol. 366, 768–778 (2007).
Sutherland, E., Coe, L. & Raleigh, E. A. McrBC: a multisubunit GTP-dependent restriction endonuclease. J. Mol. Biol. 225, 327–348 (1992).
Rifat, D., Wright, N. T., Varney, K. M., Weber, D. J. & Black, L. W. Restriction endonuclease inhibitor IPI* of bacteriophage T4: a novel structure for a dedicated target. J. Mol. Biol. 375, 720–734 (2008). This report describes the on-going evolutionary battle between phage T4 and the R–M systems of its hosts, in which the phage has evolved an inhibitor of the REase. IPI* is shown to protect T4 against the endonuclease, and the structure of IPI* is resolved.
Iida, S., Streiff, M. B., Bickle, T. A. & Arber, W. Two DNA antirestriction systems of bacteriophage P1, darA, and darB: characterization of darA− phages. Virology 157, 156–166 (1987).
Atanasiu, C., Su, T. J., Sturrock, S. S. & Dryden, D. T. Interaction of the ocr gene 0.3 protein of bacteriophage T7 with EcoKI restriction/modification enzyme. Nucleic Acids Res. 30, 3936–3944 (2002). The details of the interactions between Ocr (the best known antirestriction protein) and host Eco KI are described.
Walkinshaw, M. D. et al. Structure of Ocr from bacteriophage T7, a protein that mimics B-form DNA. Mol. Cell. 9, 187–194 (2002).
Wilson, G. G. & Murray, N. E. Restriction and modification systems. Annu. Rev. Genet. 25, 585–627 (1991).
Hill, C., Miller, L. A. & Klaenhammer, T. R. In vivo genetic exchange of a functional domain from a type II A methylase between lactococcal plasmid pTR2030 and a virulent bacteriophage. J. Bacteriol. 173, 4363–4370 (1991).
Drozdz, M., Piekarowicz, A., Bujnicki, J. M. & Radlinska, M. Novel non-specific DNA adenine methyltransferases. Nucleic Acids Res. 40, 2119–2130 (2012).
Kahmann, R. The mom gene of bacteriophage Mu. Curr. Top. Microbiol. Immunol. 108, 29–47 (1984).
Loenen, W. A. & Murray, N. E. Modification enhancement by the restriction alleviation protein (Ral) of bacteriophage λ. J. Mol. Biol. 190, 11–22 (1986).
Zabeau, M., Friedman, S., Van Montagu, M. & Schell, J. The ral gene of phage λ. I. Identification of a non-essential gene that modulates restriction and modification in E. coli. Mol. Gen. Genet. 179, 63–73 (1980).
King, G. & Murray, N. E. Restriction alleviation and modification enhancement by the Rac prophage of Escherichia coli K-12. Mol. Microbiol. 16, 769–777 (1995).
Penner, M., Morad, I., Snyder, L. & Kaufmann, G. Phage T4-coded Stp: double-edged effector of coupled DNA and tRNA-restriction systems. J. Mol. Biol. 249, 857–868 (1995).
Studier, F. W. & Movva, N. R. SAMase gene of bacteriophage T3 is responsible for overcoming host restriction. J. Virol. 19, 136–145 (1976).
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010).
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007).
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Sapranauskas, R. et al. The Streptococcus thermophilus CRISPR/Cas system provides immunity in Escherichia coli. Nucleic Acids Res. 39, 9275–9282 (2011).
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl Acad. Sci. USA 108, 10098–10103 (2011).
Datsenko, K. A. et al. Molecular memory of prior infections activates the CRISPR/Cas adaptive bacterial immunity system. Nature Commun. 3, 945 (2012).
Richter, C., Chang, J. T. & Fineran, P. C. Function and regulation of clustered regularly interspaced short palindromic repeats (CRISPR) / CRISPR Associated (Cas) systems. Viruses 4, 2291–2311 (2012).
Levin, B. R., Moineau, S., Bushman, M. & Barrangou, R. The population and evolutionary dynamics of phage and bacteria with CRISPR-mediated immunity. PLoS Genet. 9, e1003312 (2013).
Bondy-Denomy, J., Pawluk, A., Maxwell, K. L. & Davidson, A. R. Bacteriophage genes that inactivate the CRISPR/Cas bacterial immune system. Nature 493, 429–432 (2013). This article reports the first phage-encoded anti-CRISPR genes, which were discovered in P. aeruginosa lysogens.
Wiedenheft, B. In defense of phage: viral suppressors of CRISPR-mediated adaptive immunity in bacteria. RNA Biol. 10, 886–890 (2013).
Seed, K. D., Lazinski, D. W., Calderwood, S. B. & Camilli, A. A bacteriophage encodes its own CRISPR/Cas adaptive response to evade host innate immunity. Nature 494, 489–491 (2013). The first description of phages that encode their own CRISPR–Cas system to target a bacterial antiphage mechanism.
Villion, M. & Moineau, S. Virology: phages hijack a host's defence. Nature 494, 433–434 (2013).
Ram, G. et al. Staphylococcal pathogenicity island interference with helper phage reproduction is a paradigm of molecular parasitism. Proc. Natl Acad. Sci. USA 109, 16300–16305 (2012).
Molineux, I. J. Host-parasite interactions: recent developments in the genetics of abortive phage infections. New Biol. 3, 230–236 (1991).
Snyder, L. Phage-exclusion enzymes: a bonanza of biochemical and cell biology reagents? Mol. Microbiol. 15, 415–420 (1995).
Slavcev, R. A. & Hayes, S. Stationary phase-like properties of the bacteriophage lambda Rex exclusion phenotype. Mol. Genet. Genom. 269, 40–48 (2003).
Hinton, D. M. Transcriptional control in the prereplicative phase of T4 development. Virol. J. 7, 289 (2010).
Shinedling, S., Parma, D. & Gold, L. Wild-type bacteriophage T4 is restricted by the λ Rex genes. J. Virol. 61, 3790–3794 (1987).
Engelberg-Kulka, H. et al. rexB of bacteriophage λ is an anti-cell death gene. Proc. Natl Acad. Sci. USA 95, 15481–15486 (1998).
Bergsland, K. J., Kao, C., Yu, Y. T., Gulati, R. & Snyder, L. A site in the T4 bacteriophage major head protein gene that can promote the inhibition of all translation in Escherichia coli. J. Mol. Biol. 213, 477–494 (1990).
Bingham, R., Ekunwe, S. I., Falk, S., Snyder, L. & Kleanthous, C. The major head protein of bacteriophage T4 binds specifically to elongation factor Tu. J. Biol. Chem. 275, 23219–23226 (2000).
Yu, Y. T. & Snyder, L. Translation elongation factor Tu cleaved by a phage-exclusion system. Proc. Natl Acad. Sci. USA 91, 802–806 (1994).
Champness, W. C. & Snyder, L. The gol site: a cis-acting bacteriophage T4 regulatory region that can affect expression of all the T4 late genes. J. Mol. Biol. 155, 395–407 (1982).
Chopin, M. C., Chopin, A. & Bidnenko, E. Phage abortive infection in lactococci: variations on a theme. Curr. Opin. Microbiol. 8, 473–479 (2005).
Samson, J. E., Spinelli, S., Cambillau, C. & Moineau, S. Structure and activity of AbiQ, a lactococcal endoribonuclease belonging to the type III toxin–antitoxin system. Mol. Microbiol. 87, 756–768 (2013).
Bidnenko, E., Ehrlich, D. & Chopin, M. C. Phage operon involved in sensitivity to the Lactococcus lactis abortive infection mechanism AbiD1. J. Bacteriol. 177, 3824–3829 (1995).
Bidnenko, E., Ehrlich, S. D. & Chopin, M. C. Lactococcus lactis phage operon coding for an endonuclease homologous to RuvC. Mol. Microbiol. 28, 823–834 (1998).
Bidnenko, E., Chopin, A., Ehrlich, S. D. & Chopin, M. C. Activation of mRNA translation by phage protein and low temperature: the case of Lactococcus lactis abortive infection system AbiD1. BMC Mol. Biol. 10, 4 (2009).
Wang, C. et al. A reverse transcriptase-related protein mediates phage resistance and polymerizes untemplated DNA in vitro. Nucleic Acids Res. 39, 7620–7629 (2011).
Bouchard, J. D. & Moineau, S. Homologous recombination between a lactococcal bacteriophage and the chromosome of its host strain. Virology 270, 65–75 (2000).
Bouchard, J. D. & Moineau, S. Lactococcal phage genes involved in sensitivity to AbiK and their relation to single-strand annealing proteins. J. Bacteriol. 186, 3649–3652 (2004).
Labrie, S. J. & Moineau, S. Abortive infection mechanisms and prophage sequences significantly influence the genetic make-up of emerging lytic lactococcal phages. J. Bacteriol. 189, 1482–1487 (2007).
Scaltriti, E. et al. Deciphering the function of lactococcal phage ul36 Sak domains. J. Struct. Biol. 170, 462–469 (2010).
Scaltriti, E. et al. Lactococcal phage p2 ORF35-Sak3 is an ATPase involved in DNA recombination and AbiK mechanism. Mol. Microbiol. 80, 102–116 (2011).
Haaber, J., Rousseau, G. M., Hammer, K. & Moineau, S. Identification and characterization of the phage gene sav, involved in sensitivity to the lactococcal abortive infection mechanism AbiV. Appl. Environ. Microbiol. 75, 2484–2494 (2009).
Haaber, J. et al. Lactococcal abortive infection protein AbiV interacts directly with the phage protein SaV and prevents translation of phage proteins. Appl. Environ. Microbiol. 76, 7085–7092 (2010).
Labrie, S. J. et al. Involvement of the major capsid protein and two early-expressed phage genes in the activity of the lactococcal abortive infection mechanism AbiT. Appl. Environ. Microbiol. 78, 6890–6899 (2012).
Bouchard, J. D., Dion, E., Bissonnette, F. & Moineau, S. Characterization of the two-component abortive phage infection mechanism AbiT from Lactococcus lactis. J. Bacteriol. 184, 6325–6332 (2002).
Yamaguchi, Y., Park, J. H. & Inouye, M. Toxin-antitoxin systems in bacteria and archaea. Annu. Rev. Genet. 45, 61–79 (2011).
Masuda, H., Tan, Q., Awano, N., Wu, K. P. & Inouye, M. YeeU enhances the bundling of cytoskeletal polymers of MreB and FtsZ, antagonizing the CbtA (YeeV) toxicity in Escherichia coli. Mol. Microbiol. 84, 979–989 (2012).
Wang, X. et al. A new type V toxin-antitoxin system where mRNA for toxin GhoT is cleaved by antitoxin GhoS. Nature Chem. Biol. 8, 855–861 (2012).
Sberro, H. et al. Discovery of functional toxin/antitoxin systems in bacteria by shotgun cloning. Mol. Cell. 50, 136–148 (2013).
Koga, M., Otsuka, Y., Lemire, S. & Yonesaki, T. Escherichia coli rnlA and rnlB compose a novel toxin–antitoxin system. Genetics 187, 123–130 (2011).
Otsuka, Y. & Yonesaki, T. Dmd of bacteriophage T4 functions as an antitoxin against Escherichia coli LsoA and RnlA toxins. Mol. Microbiol. 83, 669–681 (2012). This work demonstrates that phage T4 encodes an antitoxin which can neutralize the effect of two bacterial toxins.
Fineran, P. C. et al. The phage abortive infection system, ToxIN, functions as a protein–RNA toxin–antitoxin pair. Proc. Natl Acad. Sci. USA 106, 894–899 (2009).
Blower, T. R. et al. A processed noncoding RNA regulates an altruistic bacterial antiviral system. Nature Struct. Mol. Biol. 18, 185–190 (2011).
Short, F. L. et al. Selectivity and self-assembly in the control of a bacterial toxin by an antitoxic noncoding RNA pseudoknot. Proc. Natl Acad. Sci. USA 110, E241–E249 (2013).
Blower, T. R., Evans, T. J., Przybilski, R., Fineran, P. C. & Salmond, G. P. Viral evasion of a bacterial suicide system by RNA-based molecular mimicry enables infectious altruism. PLoS Genet. 8, e1003023 (2012). This report provides evidence that, to circumvent ToxIN, P. atrosepticum phages hijack the host antitoxin or encode an expanded pseudo-antitoxin RNA molecule. This pseudo-antitoxin can neutralize the toxin during a phage infection, thereby assuring phage propagation.
Emond, E. et al. AbiQ, an abortive infection mechanism from Lactococcus lactis. Appl. Environ. Microbiol. 64, 4748–4756 (1998).
Samson, J. E., Belanger, M. & Moineau, S. Effect of the abortive infection mechanism and type III toxin/antitoxin system AbiQ on the lytic cycle of Lactococcus lactis phages. J. Bacteriol. http://dx.doi.org/10.1128/JB.00296-13 (2013).
Kallmeyer, J., Pockalny, R., Adhikari, R. R., Smith, D. C. & D'Hondt, S. Global distribution of microbial abundance and biomass in subseafloor sediment. Proc. Natl Acad. Sci. USA 109, 16213–16216 (2012).
Avrani, S., Schwartz, D. A. & Lindell, D. Virus-host swinging party in the oceans: incorporating biological complexity into paradigms of antagonistic coexistence. Mob. Genet. Elements 2, 88–95 (2012).
Buckling, A. & Brockhurst, M. Bacteria–virus coevolution. Adv. Exp. Med. Biol. 751, 347–370 (2012).
Bono, L. M., Gensel, C. L., Pfennig, D. W. & Burch, C. L. Competition and the origins of novelty: experimental evolution of niche-width expansion in a virus. Biol. Lett. 9, 20120616 (2013).
Stern, A. & Sorek, R. The phage-host arms race: shaping the evolution of microbes. Bioessays 33, 43–51 (2011).
Dennehy, J. J. What can phages tell us about host-pathogen coevolution? Int. J. Evol. Biol. 2012, 396165 (2012).
Abedon, S. T. Phage evolution and ecology. Adv. Appl. Microbiol. 67, 1–45 (2009).
Samson, J. E. & Moineau, S. Bacteriophages in food fermentations: new frontiers in a continuous arms race. Annu. Rev. Food Sci. Technol. 4, 347–368 (2013).
Makarova, K. S. et al. Evolution and classification of the CRISP–Cas systems. Nature Rev. Microbiol. 9, 467–477 (2011).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Magadan, A. H., Dupuis, M. E., Villion, M. & Moineau, S. Cleavage of phage DNA by the Streptococcus thermophilus CRISPR3-Cas system. PLoS ONE 7, e40913 (2012).
Shadrin, A., Sheppard, C., Severinov, K., Matthews, S. & Wigneshweraraj, S. Substitutions in the Escherichia coli RNA polymerase inhibitor T7 Gp2 that allow inhibition of transcription when the primary interaction interface between Gp2 and RNA polymerase becomes compromised. Microbiology 158, 2753–2764 (2012). This paper describes how the phage T7 protein Gp2 interacts with the host RNA polymerase to inhibit its activity and how naturally occurring mutations in both proteins affect this interaction.
Mekler, V., Minakhin, L., Sheppard, C., Wigneshweraraj, S. & Severinov, K. Molecular mechanism of transcription inhibition by phage T7 gp2 protein. J. Mol. Biol. 413, 1016–1027 (2011).
Ang, D., Keppel, F., Klein, G., Richardson, A. & Georgopoulos, C. Genetic analysis of bacteriophage-encoded cochaperonins. Annu. Rev. Genet. 34, 439–456 (2000).
Agashe, V. R. & Hartl, F. U. Roles of molecular chaperones in cytoplasmic protein folding. Semin. Cell Dev. Biol. 11, 15–25 (2000).
Sternberg, N. A genetic analysis of bacteriophage λ head assembly. Virology 71, 568–582 (1976).
Acknowledgements
The authors thank B.-A. Conway for editorial assistance. This work was supported by the Natural Sciences and Engineering Research Council of Canada (the discovery and strategic programmes), the Canadian Institutes of Health Research (an emerging-team grant) and the Ministère de l'Enseignement Supérieur, de la Recherche, de la Science et de la Technologie du Québec (the PSR SIIRI programme). J.E.S is the recipient of a scholarship from the Fonds Québécois de la Recherche sur la Nature et les Technologies. S.M. holds a Tier 1 Canada Research Chair in Bacteriophages.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Phage therapy
-
The use of phages as therapeutic agents to treat or prevent bacterial infection.
- Receptor-binding protein
-
A phage protein that is located at the tip of the phage tail and specifically recognizes its cognate receptor on the bacterial surface.
- LamB
-
An outer-membrane porin that is used to transport maltose into bacterial cells. It is also the receptor of phage λ.
- Tail fibre
-
A structural phage fibre that participates in the binding of the phage to its bacterial receptor.
- Quorum sensing signal
-
A chemical molecule that is used by bacteria to communicate with each other. This signal usually causes a bacterial response, such as the modulation of gene expression or a change in the use of metabolic pathways.
- Phase variation
-
Variation in protein expression that results in a new phenotype, but without genetic modification. It is usually induced by environmental changes.
- His box
-
A GXHXH motif that is repeated six times in the receptor-binding protein encoded by phage T4 gp37; the motif is also widely represented in several other coliphages. Recombination between the His box repeats results in the generation of a range of host-recognition specificities.
- λ reverse
-
A hybrid phage λ that has recombined with the Rac prophage of Escherichia coli K-12.
- Temperate phages
-
Phages that can integrate their genome into the bacterial chromosome through a process called lysogeny. The bacterium is then a lysogen, and the integrated phage is in a prophage state.
- Phage-inducible chromosomal island-like element
-
An 18 kb chromosomal island that is found in Vibrio cholerae strains and resembles the phage-inducible chromosomal islands of Gram-positive bacteria, including the prototypical Staphylococcus aureus pathogenicity islands.
- Terminal transferase
-
An enzyme that catalyses the addition of nucleotides to the 3′ hydroxyl terminus of DNA without the need for a specific primer.
Rights and permissions
About this article
Cite this article
Samson, J., Magadán, A., Sabri, M. et al. Revenge of the phages: defeating bacterial defences. Nat Rev Microbiol 11, 675–687 (2013). https://doi.org/10.1038/nrmicro3096
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3096
This article is cited by
-
Viruses wrap up bacterial defence systems
Nature (2024)
-
Comprehensive analysis of the CRISPR-Cas systems in Streptococcus thermophilus strains isolated from traditional yogurts
Antonie van Leeuwenhoek (2024)
-
Variants of a putative baseplate wedge protein extend the host range of Pseudomonas phage K8
Microbiome (2023)
-
Genomic and transcriptomic insights into complex virus–prokaryote interactions in marine biofilms
The ISME Journal (2023)
-
The highly diverse antiphage defence systems of bacteria
Nature Reviews Microbiology (2023)