Key Points
-
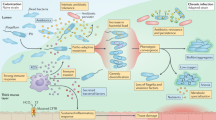
Cystic fibrosis (CF) is caused by a mutation in the CF transmembrane conductance regulator gene, which encodes a cyclic AMP-regulated chloride ion channel. The mutation results in defective ion transport across epithelial cell surfaces in the upper airways, interfering with the clearance of particles and microbial cells that are trapped in the overlying mucus and resulting in a greatly increased susceptibility to bacterial infection. As a consequence, the airways of patients with CF are nearly always infected with many different bacterial species, but Pseudomonas aeruginosa infection causes the greatest burden of morbidity and mortality.
-
P. aeruginosa infection of the airways of patients with CF begins with a period of recurrent, intermittent colonization from the environment or from a protected niche within the patient, such as the paranasal sinuses. These intermittent infections can be effectively controlled with aggressive antibiotic therapy, but eventually a chronic infection develops that can persist for the rest of the patient's lifetime.
-
Chronic infections of the CF airway by P. aeruginosa thus provide a valuable opportunity to study bacterial evolution in a complex natural environment. These infections are most often clonal, and regular sampling is possible from samples collected when the patients attend clinics. With the recent developments in DNA sequencing and other genome-wide analysis tools, it is now possible to produce a detailed characterization and time-resolved map of the evolutionary trajectories of the infecting bacteria.
-
In the CF lung, P. aeruginosa encounters stressful conditions, including oxidative, nitrosative and cell envelope stress, and antibiotics. The switches in P. aeruginosa gene expression that occur in response to these stresses and to antibiotics have been well characterized, and the global regulators responsible have been identified.
-
Recent tracking of genomic evolution in sequential P. aeruginosa isolates from individual patients has identified a set of genes that were repeatedly found to have acquired non-synonymous mutations. These genes include three global regulators that were known from in vitro work to be important in P. aeruginosa adaptation to the CF lung: mucA, which encodes an inner-membrane-associated anti-σ-factor and controls alginate synthesis; lasR, which encodes a transcriptional regulator of quorum sensing; and rpoN, which encodes an alternative RNA polymerase σ-factor involved in the regulation of virulence gene expression.
Abstract
The airways of patients with cystic fibrosis (CF) are nearly always infected with many different microorganisms. This environment offers warm, humid and nutrient-rich conditions, but is also stressful owing to frequent antibiotic therapy and the host immune response. Pseudomonas aeruginosa is commonly isolated from the airways of patients with CF, where it most often establishes chronic infections that usually persist for the rest of the lives of the patients. This bacterium is a major cause of mortality and morbidity and has therefore been studied intensely. Here, we discuss how P. aeruginosa evolves from a state of early, recurrent intermittent colonization of the airways of patients with CF to a chronic infection state, and how this process offers opportunities to study bacterial evolution in natural environments. We believe that such studies are valuable not only for our understanding of bacterial evolution but also for the future development of new therapeutic strategies to treat severe chronic infections.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Bye, M. R., Ewig, J. M. & Quittell, L. M. Cystic fibrosis. Lung 172, 251–270 (1994).
Boucher, R. C. Relationship of airway epithelial ion transport to chronic bronchitis. Proc. Am. Thorac. Soc. 1, 66–70 (2004).
Armstrong, D. S. et al. Lower respiratory infection and inflammation in infants with newly diagnosed cystic fibrosis. BMJ 310, 1571–1572 (1995).
Armstrong, D. S. et al. Lower airway inflammation in infants with cystic fibrosis detected by newborn screening. Pediatr. Pulmonol. 40, 500–510 (2005).
Hoiby, N., Frederiksen, B. & Pressler, T. Eradication of early Pseudomonas aeruginosa infection. J. Cyst. Fibros. 4, 49–54 (2005).
Cystic Fibrosis Foundation. Cystic Fibrosis Foundation patient registry 2010 annual data report (Cystic Fibrosis Foundation, 2011).
Bjarnsholt, T. et al. Pseudomonas aeruginosa biofilms in the respiratory tract of cystic fibrosis patients. Pediatr. Pulmonol. 44, 547–558 (2009).
Ulrich, M. et al. Alveolar inflammation in cystic fibrosis. J. Cyst. Fibros. 9, 217–227 (2010).
Yang, L., Jelsbak, L. & Molin, S. Microbial ecology and adaptation in cystic fibrosis airways. Environ. Microbiol. 13, 1682–1689 (2011).
Hansen, C. R., Pressler, T. & Hoiby, N. Early aggressive eradication therapy for intermittent Pseudomonas aeruginosa airway colonization in cystic fibrosis patients: 15 years experience. J. Cyst. Fibros. 7, 523–530 (2008).
Rasmussen, J. et al. CT of the paranasal sinuses is not a valid indicator for sinus surgery in CF patients. J. Cyst. Fibros. 11, 93–99 (2012).
Harrison, F. Microbial ecology of the cystic fibrosis lung. Microbiology 153, 917–923 (2007).
Koch, C. Early infection and progression of cystic fibrosis lung disease. Pediatr. Pulmonol. 34, 232–236 (2002).
Hoiby, N. & Johansen, H. K. Isolation measures for prevention of infection with respiratory pathogens in cystic fibrosis: a systematic review? J. Hosp. Infect. 65, 374–375 (2007).
FitzSimmons, S. C. The changing epidemiology of cystic fibrosis. J. Pediatr. 122, 1–9 (1993).
Burns, J. L. et al. Longitudinal assessment of Pseudomonas aeruginosa in young children with cystic fibrosis. J. Infect. Dis. 183, 444–452 (2001).
Johansen, H. K. & Hoiby, N. Seasonal onset of initial colonisation and chronic infection with Pseudomonas aeruginosa in patients with cystic fibrosis in Denmark. Thorax 47, 109–111 (1992).
Doring, G., Taccetti, G., Campana, S., Festini, F. & Mascherini, M. Eradication of Pseudomonas aeruginosa in cystic fibrosis patients. Eur. Respir. J. 27, 653 (2006).
Gibson, R. L. et al. Significant microbiological effect of inhaled tobramycin in young children with cystic fibrosis. Am. J. Respir. Crit. Care Med. 167, 841–849 (2003).
Munck, A. et al. Genotypic characterization of Pseudomonas aeruginosa strains recovered from patients with cystic fibrosis after initial and subsequent colonization. Pediatr. Pulmonol. 32, 288–292 (2001).
Jelsbak, L. et al. Molecular epidemiology and dynamics of Pseudomonas aeruginosa populations in lungs of cystic fibrosis patients. Infect. Immun. 75, 2214–2224 (2007).
Doring, G. et al. Antibiotic therapy against Pseudomonas aeruginosa in cystic fibrosis: a European consensus. Eur. Respir. J. 16, 749–767 (2000).
Doring, G. & Hoiby, N. Early intervention and prevention of lung disease in cystic fibrosis: a European consensus. J. Cyst. Fibros. 3, 67–91 (2004).
Lee, T. W., Brownlee, K. G., Conway, S. P., Denton, M. & Littlewood, J. M. Evaluation of a new definition for chronic Pseudomonas aeruginosa infection in cystic fibrosis patients. J. Cyst. Fibros. 2, 29–34 (2003).
Nichols, D., Chmiel, J. & Berger, M. Chronic inflammation in the cystic fibrosis lung: alterations in inter- and intracellular signaling. Clin. Rev. Allergy Immunol. 34, 146–162 (2008).
Regamey, N., Jeffery, P. K., Alton, E. W., Bush, A. & Davies, J. C. Airway remodelling and its relationship to inflammation in cystic fibrosis. Thorax 66, 624–629 (2011).
Anderson, G. G., Moreau-Marquis, S., Stanton, B. A. & O'Toole, G. A. In vitro analysis of tobramycin-treated Pseudomonas aeruginosa biofilms on cystic fibrosis-derived airway epithelial cells. Infect. Immun. 76, 1423–1433 (2008).
Aspedon, A., Palmer, K. & Whiteley, M. Microarray analysis of the osmotic stress response in Pseudomonas aeruginosa. J. Bacteriol. 188, 2721–2725 (2006).
Bagge, N. et al. Pseudomonas aeruginosa biofilms exposed to imipenem exhibit changes in global gene expression and β-lactamase and alginate production. Antimicrob. Agents Chemother. 48, 1175–1187 (2004).
Cirz, R. T., O'Neill, B. M., Hammond, J. A., Head, S. R. & Romesberg, F. E. Defining the Pseudomonas aeruginosa SOS response and its role in the global response to the antibiotic ciprofloxacin. J. Bacteriol. 188, 7101–7110 (2006).
Jones, A. K. et al. Activation of the Pseudomonas aeruginosa AlgU regulon through mucA mutation inhibits cyclic AMP/Vfr signaling. J. Bacteriol. 192, 5709–5717 (2010).
Wu, W., Badrane, H., Arora, S., Baker, H. V. & Jin, S. MucA-mediated coordination of type III secretion and alginate synthesis in Pseudomonas aeruginosa. J. Bacteriol. 186, 7575–7585 (2004).
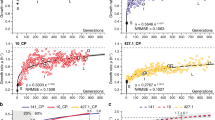
Huse, H. K. et al. Parallel evolution in Pseudomonas aeruginosa over 39,000 generations in vivo. mBio 1, e00199–10 (2010).
Rau, M. H. et al. Early adaptive developments of Pseudomonas aeruginosa after the transition from life in the environment to persistent colonization in the airways of human cystic fibrosis hosts. Environ. Microbiol. 12, 1643–1658 (2010).
Yang, L. et al. Evolutionary dynamics of bacteria in a human host environment. Proc. Natl Acad. Sci. USA 108, 7481–7486 (2011). This report outlines the evolutionary dynamics of a transmissible P. aeruginosa clone (DK2) during 35 years of growth in a CF airway.
Hull, J., Vervaart, P., Grimwood, K. & Phelan, P. Pulmonary oxidative stress response in young children with cystic fibrosis. Thorax 52, 557–560 (1997).
Hudson, A. L., Sotirchos, I. M., Davey, M. W. & Hudson, V. M. The activity and hydrogen peroxide sensitivity of the peroxiredoxins from the parasitic nematode Haemonchus contortus. Mol. Biochem. Parasitol. 176, 17–24 (2011).
Hudson, V. M. Rethinking cystic fibrosis pathology: the critical role of abnormal reduced glutathione (GSH) transport caused by CFTR mutation. Free Radic. Biol. Med. 30, 1440–1461 (2001).
Back, E. I. et al. Antioxidant deficiency in cystic fibrosis: when is the right time to take action? Am. J. Clin. Nutr. 80, 374–384 (2004).
Worlitzsch, D. et al. Effects of reduced mucus oxygen concentration in airway Pseudomonas infections of cystic fibrosis patients. J. Clin. Invest. 109, 317–325 (2002).
Kolpen, M. et al. Polymorphonuclear leucocytes consume oxygen in sputum from chronic Pseudomonas aeruginosa pneumonia in cystic fibrosis. Thorax 65, 57–62 (2010).
Hauser, A. R., Jain, M., Bar-Meir, M. & McColley, S. A. Clinical significance of microbial infection and adaptation in cystic fibrosis. Clin. Microbiol. Rev. 24, 29–70 (2011).
Johansen, H. K. et al. Antibody response to Pseudomonas aeruginosa in cystic fibrosis patients: a marker of therapeutic success? A 30-year cohort study of survival in Danish CF patients after onset of chronic P. aeruginosa lung infection. Pediatr. Pulmonol. 37, 427–432 (2004).
Lister, P. D., Wolter, D. J. & Hanson, N. D. Antibacterial-resistant Pseudomonas aeruginosa: clinical impact and complex regulation of chromosomally encoded resistance mechanisms. Clin. Microbiol. Rev. 22, 582–610 (2009).
Bert, F., Branger, C. & Lambert-Zechovsky, N. Identification of PSE and OXA β-lactamase genes in Pseudomonas aeruginosa using PCR-restriction fragment length polymorphism. J. Antimicrob. Chemother. 50, 11–18 (2002).
MacLeod, D. L. et al. Aminoglycoside-resistance mechanisms for cystic fibrosis Pseudomonas aeruginosa isolates are unchanged by long-term, intermittent, inhaled tobramycin treatment. J. Infect. Dis. 181, 1180–1184 (2000).
Poole, K. Aminoglycoside resistance in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 49, 479–487 (2005).
Smith, E. E. et al. Genetic adaptation by Pseudomonas aeruginosa to the airways of cystic fibrosis patients. Proc. Natl Acad. Sci. USA 103, 8487–8492 (2006). This paper describes the genomic analysis of longitudinal isolates of P. aeruginosa sampled from a young patient with CF, and provides a first insight into how this pathogen adapts to its human host.
Costerton, J. W., Irvin, R. T. & Cheng, K. J. The bacterial glycocalyx in nature and disease. Annu. Rev. Microbiol. 35, 299–324 (1981).
Cabral, D. A., Loh, B. A. & Speert, D. P. Mucoid Pseudomonas aeruginosa resists nonopsonic phagocytosis by human neutrophils and macrophages. Pediatr. Res. 22, 429–431 (1987).
Meshulam, T., Obedeanu, N., Merzbach, D. & Sobel, J. D. Phagocytosis of mucoid and nonmucoid strains of Pseudomonas aeruginosa. Clin. Immunol. Immunopathol. 32, 151–165 (1984).
Meshulam, T., Verbrugh, H. A. & Verhoef, J. Opsonization and phagocytosis of mucoid and non-mucoid Pseudomonas aeruginosa strains. Eur. J. Clin. Microbiol. 1, 112–117 (1982).
Simpson, J. A., Smith, S. E. & Dean, R. T. Scavenging by alginate of free radicals released by macrophages. Free Radic. Biol. Med. 6, 347–353 (1989).
Pedersen, S. S., Kharazmi, A., Espersen, F. & Hoiby, N. Pseudomonas aeruginosa alginate in cystic fibrosis sputum and the inflammatory response. Infect. Immun. 58, 3363–3368 (1990).
Govan, J. R. & Deretic, V. Microbial pathogenesis in cystic fibrosis: mucoid Pseudomonas aeruginosa and Burkholderia cepacia. Microbiol. Rev. 60, 539–574 (1996).
Pedersen, S. S. Lung infection with alginate-producing, mucoid Pseudomonas aeruginosa in cystic fibrosis. APMIS Suppl. 28, 1–79 (1992).
Martin, D. W. et al. Mechanism of conversion to mucoidy in Pseudomonas aeruginosa infecting cystic fibrosis patients. Proc. Natl Acad. Sci. USA 90, 8377–8381 (1993).
Boucher, J. C., Yu, H., Mudd, M. H. & Deretic, V. Mucoid Pseudomonas aeruginosa in cystic fibrosis: characterization of muc mutations in clinical isolates and analysis of clearance in a mouse model of respiratory infection. Infect. Immun. 65, 3838–3846 (1997).
Wood, L. F. & Ohman, D. E. Identification of genes in the σ22 regulon of Pseudomonas aeruginosa required for cell envelope homeostasis in either the planktonic or the sessile mode of growth. mBio 3, e00094–12 (2012).
Ramsey, D. M. & Wozniak, D. J. Understanding the control of Pseudomonas aeruginosa alginate synthesis and the prospects for management of chronic infections in cystic fibrosis. Mol. Microbiol. 56, 309–322 (2005).
Deretic, V., Schurr, M. J., Boucher, J. C. & Martin, D. W. Conversion of Pseudomonas aeruginosa to mucoidy in cystic fibrosis: environmental stress and regulation of bacterial virulence by alternative sigma factors. J. Bacteriol. 176, 2773–2780 (1994).
Chitnis, C. E. & Ohman, D. E. Genetic analysis of the alginate biosynthetic gene cluster of Pseudomonas aeruginosa shows evidence of an operonic structure. Mol. Microbiol. 8, 583–593 (1993).
DeVries, C. A. & Ohman, D. E. Mucoid-to-nonmucoid conversion in alginate-producing Pseudomonas aeruginosa often results from spontaneous mutations in algT, encoding a putative alternate sigma factor, and shows evidence for autoregulation. J. Bacteriol. 176, 6677–6687 (1994).
Wood, L. F. & Ohman, D. E. Use of cell wall stress to characterize σ22 (AlgT/U) activation by regulated proteolysis and its regulon in Pseudomonas aeruginosa. Mol. Microbiol. 72, 183–201 (2009).
Wood, L. F., Leech, A. J. & Ohman, D. E. Cell wall-inhibitory antibiotics activate the alginate biosynthesis operon in Pseudomonas aeruginosa: roles of σ22 (AlgT) and the AlgW and Prc proteases. Mol. Microbiol. 62, 412–426 (2006).
Ciofu, O., Fussing, V., Bagge, N., Koch, C. & Hoiby, N. Characterization of paired mucoid/non-mucoid Pseudomonas aeruginosa isolates from Danish cystic fibrosis patients: antibiotic resistance, β-lactamase activity and RiboPrinting. J. Antimicrob. Chemother. 48, 391–396 (2001).
Gilligan, P. H. Microbiology of airway disease in patients with cystic fibrosis. Clin. Microbiol. Rev. 4, 35–51 (1991).
Rainey, P. B. & Travisano, M. Adaptive radiation in a heterogeneous environment. Nature. 394, 69–72 (1998).
Mowat, E. et al. Pseudomonas aeruginosa population diversity and turnover in cystic fibrosis chronic infections. Am. J. Respir. Crit. Care Med. 183, 1674–1679 (2011).
Hansen, S. K. et al. Evolution and diversification of Pseudomonas aeruginosa in the paranasal sinuses of cystic fibrosis children have implications for chronic lung infection. ISME J. 6, 31–45 (2012).
Oliver, A., Canton, R., Campo, P., Baquero, F. & Blazquez, J. High frequency of hypermutable Pseudomonas aeruginosa in cystic fibrosis lung infection. Science 288, 1251–1254 (2000).
Hassett, D. J. et al. Pseudomonas aeruginosa biofilm infections in cystic fibrosis: insights into pathogenic processes and treatment strategies. Expert Opin. Ther. Targets 14, 117–130 (2010).
Fokkens, W. J. et al. EPOS 2012: European position paper on rhinosinusitis and nasal polyps 2012. A summary for otorhinolaryngologists. Rhinology 50, 1–12 (2012).
Johansen, H. K. et al. Colonisation and infection of the paranasal sinuses in cystic fibrosis patients is accompanied by a reduced PMN response. J. Cyst. Fibros. 15 May 2012 (doi:10.1016/j.jcf.2012.04.011).
Aanaes, K. et al. Decreased mucosal oxygen tension in the maxillary sinuses in patients with cystic fibrosis. J. Cyst. Fibros. 10, 114–120 (2011).
Robertson, J. M., Friedman, E. M. & Rubin, B. K. Nasal and sinus disease in cystic fibrosis. Paediatr. Respir. Rev. 9, 213–219 (2008).
Mainz, J. G. et al. Concordant genotype of upper and lower airways P.aeruginosa and S.aureus isolates in cystic fibrosis. Thorax 64, 535–540 (2009).
Bonestroo, H. J., de Winter-de Groot, K. M., van der Ent, C. K. & Arets, H. G. Upper and lower airway cultures in children with cystic fibrosis: do not neglect the upper airways. J. Cyst. Fibros. 9, 130–134 (2010).
West, S. A., Griffin, A. S., Gardner, A. & Diggle, S. P. Social evolution theory for microorganisms. Nature Rev. Microbiol. 4, 597–607 (2006).
Hoffman, L. R. et al. Pseudomonas aeruginosa lasR mutants are associated with cystic fibrosis lung disease progression. J. Cyst. Fibros. 8, 66–70 (2009).
D'Argenio, D. A. et al. Growth phenotypes of Pseudomonas aeruginosa lasR mutants adapted to the airways of cystic fibrosis patients. Mol. Microbiol. 64, 512–533 (2007).
Cramer, N. et al. Microevolution of the major common Pseudomonas aeruginosa clones C and PA14 in cystic fibrosis lungs. Environ. Microbiol. 13, 1690–1704 (2011).
Blount, Z. D., Borland, C. Z. & Lenski, R. E. Historical contingency and the evolution of a key innovation in an experimental population of Escherichia coli. Proc. Natl Acad. Sci. USA 105, 7899–7906 (2008).
Weinreich, D. M., Delaney, N. F., Depristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006).
Woods, R. J. et al. Second-order selection for evolvability in a large Escherichia coli population. Science 331, 1433–1436 (2011).
Mahenthiralingam, E., Campbell, M. E., Foster, J., Lam, J. S. & Speert, D. P. Random amplified polymorphic DNA typing of Pseudomonas aeruginosa isolates recovered from patients with cystic fibrosis. J. Clin. Microbiol. 34, 1129–1135 (1996).
Romling, U. et al. Epidemiology of chronic Pseudomonas aeruginosa infections in cystic fibrosis. J. Infect. Dis. 170, 1616–1621 (1994).
Jelsbak, L. et al. Molecular epidemiology and dynamics of Pseudomonas aeruginosa populations in lungs of cystic fibrosis patients. Infect. Immun. 75, 2214–2224 (2007).
Yang, L. et al. In situ growth rates and biofilm development of Pseudomonas aeruginosa populations in chronic lung infections. J. Bacteriol. 190, 2767–2776 (2008).
Lieberman, T. D. et al. Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nature Genet. 43, 1275–1280 (2011). This article provides an example of genomic evolution in a CF-associated pathogen.
Cramer, N., Wiehlmann, L. & Tummler, B. Clonal epidemiology of Pseudomonas aeruginosa in cystic fibrosis. Int. J. Med. Microbiol. 300, 526–533 (2010).
Bragonzi, A. et al. Pseudomonas aeruginosa microevolution during cystic fibrosis lung infection establishes clones with adapted virulence. Am. J. Respir. Crit. Care Med. 180, 138–145 (2009).
Hoboth, C. et al. Dynamics of adaptive microevolution of hypermutable Pseudomonas aeruginosa during chronic pulmonary infection in patients with cystic fibrosis. J. Infect. Dis. 200, 118–130 (2009).
Hogardt, M. & Heesemann, J. Adaptation of Pseudomonas aeruginosa during persistence in the cystic fibrosis lung. Int. J. Med. Microbiol. 300, 557–562 (2010).
Barrick, J. E. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature 461, 1243–1247 (2009). This work uses laboratory evolution experiments combined with genomic analysis to investigate evolution in action.
Cooper, V. S. & Lenski, R. E. The population genetics of ecological specialization in evolving Escherichia coli populations. Nature 407, 736–739 (2000).
Silander, O. K., Tenaillon, O. & Chao, L. Understanding the evolutionary fate of finite populations: the dynamics of mutational effects. PLoS Biol. 5, e94 (2007).
Holder, K. K. & Bull, J. J. Profiles of adaptation in two similar viruses. Genetics 159, 1393–1404 (2001).
Hansen, S. K. et al. Characterization of a Pseudomonas putida rough variant evolved in a mixed-species biofilm with Acinetobacter sp. strain C6. J. Bacteriol. 189, 4932–4943 (2007).
Boles, B. R., Thoendel, M. & Singh, P. K. Self-generated diversity produces “insurance effects” in biofilm communities. Proc. Natl Acad. Sci. USA 101, 16630–16635 (2004).
Keymer, J. E., Galajda, P., Muldoon, C., Park, S. & Austin, R. H. Bacterial metapopulations in nanofabricated landscapes. Proc. Natl Acad. Sci. USA 103, 17290–17295 (2006).
Wilder, C. N., Allada, G. & Schuster, M. Instantaneous within-patient diversity of Pseudomonas aeruginosa quorum-sensing populations from cystic fibrosis lung infections. Infect. Immun. 77, 5631–5639 (2009).
Hunter, P. The great leap forward. Major evolutionary jumps might be caused by changes in gene regulation rather than the emergence of new genes. EMBO Rep. 9, 608–611 (2008).
Wang, L. et al. Divergence involving global regulatory gene mutations in an Escherichia coli population evolving under phosphate limitation. Genome Biol. Evol. 2, 478–487 (2010).
Zambrano, M. M., Siegele, D. A., Almiron, M., Tormo, A. & Kolter, R. Microbial competition: Escherichia coli mutants that take over stationary phase cultures. Science 259, 1757–1760 (1993).
Conrad, T. M., Lewis, N. E. & Palsson, B. O. Microbial laboratory evolution in the era of genome-scale science. Mol. Syst. Biol. 7, 509 (2011).
Sumby, P., Whitney, A. R., Graviss, E. A., DeLeo, F. R. & Musser, J. M. Genome-wide analysis of group a streptococci reveals a mutation that modulates global phenotype and disease specificity. PLoS Pathog. 2, e5 (2006).
Giraud, A. et al. Dissecting the genetic components of adaptation of Escherichia coli to the mouse gut. PLoS Genet. 4, e2 (2008).
Kresse, A. U., Dinesh, S. D., Larbig, K. & Romling, U. Impact of large chromosomal inversions on the adaptation and evolution of Pseudomonas aeruginosa chronically colonizing cystic fibrosis lungs. Mol. Microbiol. 47, 145–158 (2003).
Klockgether, J., Cramer, N., Wiehlmann, L., Davenport, C. F. & Tummler, B. Pseudomonas aeruginosa genomic structure and diversity. Front. Microbiol. 2, 150 (2011).
Hindre, T., Knibbe, C., Beslon, G. & Schneider, D. New insights into bacterial adaptation through in vivo and in silico experimental evolution. Nature Rev. Microbiol. 10, 352–365 (2012). This review describes how experimental evolution studies have provided fundamental insight into bacterial adaptive mechanisms.
Willner, D. et al. Spatial distribution of microbial communities in the cystic fibrosis lung. ISME J. 6, 471–474 (2012).
Zhao, J. et al. Decade-long bacterial community dynamics in cystic fibrosis airways. Proc. Natl Acad. Sci. USA 109, 5809–5814 (2012).
Klepac-Ceraj, V. et al. Relationship between cystic fibrosis respiratory tract bacterial communities and age, genotype, antibiotics and Pseudomonas aeruginosa. Environ. Microbiol. 12, 1293–1303 (2010).
Kleinstreuer, C., Zhang, Z. and Donohue, J. F. Targeted drug-aerosol delivery in the human respiratory system. Annu. Rev. Biomed. Eng. 10, 195–220 (2008).
Winnie, G. B. & Cowan, R. G. Respiratory tract colonization with Pseudomonas aeruginosa in cystic fibrosis: correlations between anti-Pseudomonas aeruginosa antibody levels and pulmonary function. Pediatr. Pulmonol. 10, 92–100 (1991).
Martin, D. W., Schurr, M. J., Yu, H. & Deretic, V. Analysis of promoters controlled by the putative sigma factor AlgU regulating conversion to mucoidy in Pseudomonas aeruginosa: relationship to σE and stress response. J. Bacteriol. 176, 6688–6696 (1994).
Acknowledgements
The authors thank the Danish Agency for Science and Technology and the Lundbeck Foundation for generous support to the cystic fibrosis collaborative project during the past decade.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Periciliary fluid
-
The layer of fluid that is in contact with the surface epithelial cells in the airway.
- Polymorphonuclear leukocytes
-
A class of white blood cells that includes neutrophils, eosinophils and basophils, and is characterized by the possession of multilobed nuclei.
- Envelope stress response
-
A signal transduction pathway that senses and responds to extracytoplasmic stress.
- Complement
-
Part of the innate immune response. The complement system comprises more than 25 proteins that recognize foreign objects and target them for destruction or phagocytosis.
- Anti-σ-factor
-
A negative transcriptional regulator that binds to a target RNA polymerase σ-factor to prevent its activity.
- Hypermutability
-
A mutation rate that is high compared with the average mutation rate for the species.
- Chronic rhinosinusitis
-
In adults: inflammation of the nose and the paranasal sinuses, characterized by two or more symptoms, one of which should be nasal blockage, obstruction, congestion or discharge (anterior or posterior nasal drip). The other symptoms are one or more of the following: facial pain or pressure; a reduction or loss of smell; a cough; endoscopic signs of nasal polyps; mucopurulent discharge (primarily from the middle meatus); oedema or mucosal obstruction (primarily in the middle meatus); and CT changes.
- Clonal complex
-
A group of bacterial isolates, as derived from multilocus sequence typing (MLST) analysis. A clonal complex usually includes isolates that differ from each other at only one of the seven loci analysed by MLST.
Rights and permissions
About this article
Cite this article
Folkesson, A., Jelsbak, L., Yang, L. et al. Adaptation of Pseudomonas aeruginosa to the cystic fibrosis airway: an evolutionary perspective. Nat Rev Microbiol 10, 841–851 (2012). https://doi.org/10.1038/nrmicro2907
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2907
This article is cited by
-
Mortality and exacerbations associated with Stenotrophomonas maltophilia in chronic obstructive pulmonary disease. A regional cohort study of 22,689 outpatients
Respiratory Research (2023)
-
Quantifying the local adaptive landscape of a nascent bacterial community
Nature Communications (2023)
-
Mixed strain pathogen populations accelerate the evolution of antibiotic resistance in patients
Nature Communications (2023)
-
Characterization of the bacterial microbiome of Amblyomma scalpturatum and Amblyomma ovale collected from Tapirus terrestris and Amblyomma sabanerae collected from Chelonoidis denticulata, Madre de Dios- Peru
BMC Microbiology (2022)
-
Gut to lung translocation and antibiotic mediated selection shape the dynamics of Pseudomonas aeruginosa in an ICU patient
Nature Communications (2022)