Key Points
-
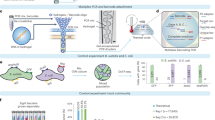
Genomic sequencing from a single cell became feasible with the introduction of the multiple displacement amplification (MDA) method, which generates micrograms of amplified DNA.
-
Methods have improved over the past 10 years to isolate single cells, amplify the DNA by MDA for use in sequencing, and assemble genomes from those single cells.
-
A vast range of novel microorganisms will now be amenable to genomic sequencing directly from single cells, eliminating the need to develop culture methods in order to obtain sufficient DNA template.
-
Single-cell sequencing is increasingly being used in combination with metagenomic sequencing to assemble individual genomes and analyse complex microbial communities.
Abstract
Sequencing DNA from single cells has opened new windows onto the microbial world. It is becoming routine to sequence bacterial species directly from environmental samples or clinical specimens without the need to develop cultivation methods. Recent technical improvements often allow nearly complete genome assembly from these otherwise inaccessible species. New bioinformatics methods are also improving genome assembly from single cells. The use of single-cell sequencing in combination with metagenomic analysis is also emerging as a powerful new strategy to analyse bacterial communities. Here, the technical developments that have enabled single-cell sequencing, as well as some of the most exciting applications of this approach from the past few years, are reviewed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Raghunathan, A. et al. Genomic DNA amplification from a single bacterium. Appl. Environ. Microbiol. 71, 3342–3347 (2005). The first paper to demonstrate DNA sequencing from a single cell.
Dean, F. B., Nelson, J. R., Giesler, T. L. & Lasken, R. S. Rapid amplification of plasmid and phage DNA using Phi29 DNA polymerase and multiply-primed rolling circle amplification. Genome Res. 11, 1095–1099 (2001).
Dean, F. B. et al. Comprehensive human genome amplification using multiple displacement amplification. Proc. Natl Acad. Sci. USA 99, 5261–5266 (2002). References 2 and 3 introduced the MDA method.
Lane, D. J. et al. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc. Natl Acad. Sci. USA 82, 6955–6959 (1985).
Olsen, G. J., Lane, D. J., Giovannoni, S. J., Pace, N. R. & Stahl, D. A. Microbial ecology & evolution: a ribosomal RNA approach. Annu. Rev. Microbiol. 40, 337–365 (1986).
Woese, C. R., Fox, G. E. & Zablen, L. Conservation of primary structure in 16S ribosomal RNA. Nature 254, 83–86 (1975).
Giovanonni, S. J., Britschgi, T. B., Moyer, C. L. & Field, K. G. Genetic diversity in Sargasso Sea bacterioplankton. Nature Biotech. 345, 60–63 (1990).
Schmidt, T. M., DeLong, E. F. & Pace, N. R. Analysis of a marine picoplankton community using 16S rRNA gene cloning and sequencing. J. Bacteriol. 173, 4371–4378 (1991).
Ward, D. M., Weller, R. & Bateson, M. M. 16S rRNA sequences reveal numerous uncultured microorganisms in a natural community. Nature Biotech. 345, 63–65 (1990).
Handelsman, J. Metagenomics: applications of genomics to uncultured microorganisms. Microbiol. Mol. Biol. Rev. 68, 669–685 (2004).
Tyson, G. W. et al. Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428, 37–43 (2004).
Venter, J. C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
Rusch, D. B. et al. The Sorcerer II Global Ocean Sampling expedition: Northwest Atlantic through Eastern Tropical Pacific. PLoS Biol. 5, e77 (2007).
Yooseph, S. et al. The Sorcerer II Global Ocean Sampling expedition: expanding the universe of protein families. PLoS Biol. 5, e16 (2007).
Gilbert, J. A. & Dupont, C. L. Microbial metagenomics: beyond the genome. Ann. Rev. Mar. Sci. 3, 347–371 (2011).
Rusch, D. B., Martiny, A. C., Dupont, C. L., Halpern, A. L. & Venter, J. C. Characterization of Prochlorococcus clades from iron-depleted oceanic regions. Proc. Natl Acad. Sci. USA 107, 16184–16189 (2010).
Telenius, H. et al. Degenerate oligonucleotide-primed PCR: general amplification of target DNA by a single degenerate primer. Genomics 13, 718–725 (1992).
Zhang, L. et al. Whole genome amplification from a single cell: implications for genetic analysis. Proc. Natl Acad. Sci. USA 89, 5847–5851 (1992).
Hartley, J. L. Amplification of nucleic acid sequences using oligonucleotides of random sequence as primers. US Patent 5043272 (1989).
Lizardi, P. M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nature Genet. 19, 225–232 (1998).
Blanco, L. & Salas, M. Characterization and purification of a phage φ29-encoded DNA polymerase required for the initiation of replication. Proc. Natl Acad. Sci. USA 81, 5325–5329 (1984).
Blanco, L. et al. Highly efficient DNA synthesis by the phage φ29 DNA polymerase. Symmetrical mode of DNA replication. J. Biol. Chem. 264, 8935–8940 (1989).
Lasken, R. S. in Whole Genome Amplification: Methods Express Series (eds Hughes, S. & Lasken, R.) 99–118 (Scion, 2005).
Hosono, S. et al. Unbiased whole-genome amplification directly from clinical samples. Genome Res. 13, 954–964 (2003).
Hawkins, T. L., Detter, J. C. & Richardson, P. M. Whole genome amplification — applications and advances. Curr. Opin. Biotechnol. 13, 65–67 (2002).
Detter, J. C. et al. Isothermal strand-displacement amplification applications for high-throughput genomics. Genomics 80, 691–698 (2002).
Nelson, J. R. et al. TempliPhi, φ29 DNA polymerase based rolling circle amplification of templates for DNA sequencing. Biotechniques 32 (Suppl. 1), 44–47 (2002).
Lasken, R. S. Single cell genomic sequencing using multiple displacement amplification. Curr. Opin. Microbiol. 10, 1–7 (2007).
Blainey, P. C. & Quake, S. R. Digital MDA for enumeration of total nucleic acid contamination. Nucleic Acids Res. 39, e19 (2011).
Rodrigue, S. et al. Whole genome amplification and de novo assembly of single bacterial cells. PLoS ONE 4, e6864 (2009). This paper demonstrates a comprehensive workflow to carry out single-cell sequencing and genome assembly.
Woyke, T. et al. One bacterial cell, one complete genome. PLoS ONE 5, e10314 (2010).
Zhang, K. et al. Sequencing genomes from single cells by polymerase cloning. Nature Biotech. 24, 680–686 (2006). The first attempt to complete a bacterial genome assembly from a single cell.
Hutchison, C. A., I. I. I., Smith, H. O., Pfannkoch, C. & Venter, J. C. Cell-free cloning using φ29 DNA polymerase. Proc. Natl Acad. Sci. USA 102, 17332–17336 (2005).
Marcy, Y. et al. Nanoliter reactors improve multiple displacement amplification of genomes from single cells. PLoS Genet. 3, 1702–1708 (2007).
Lasken, R. S. & Stockwell, T. B. Mechanism of chimera formation during the multiple displacement amplification reaction. BMC Biotechnol. 7, 19 (2007).
Lasken, R. S. et al. in Whole Genome Amplification: Methods Express Series (eds Hughes, S. & Lasken, R.) 119–147 (Scion, 2005).
Hongoh, Y. et al. Complete genome of the uncultured Termite Group 1 bacteria in a single host protist cell. Proc. Natl Acad. Sci. USA 105, 5555–5560 (2008). Sequencing of an intracellular symbiotic bacteria.
Yoon, H. S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011).
Woyke, T. et al. Assembling the marine metagenome, one cell at a time. PLoS ONE 4, e5299 (2009). This paper also demonstrates a comprehensive workflow to carry out single-cell sequencing and genome assembly.
Chitsaz, H. et al. Efficient de novo assembly of bacterial genomes from single cells. Nature Biotech. 29, 915–921 (2011). A bioinformatic approach designed to improve genome assembly from amplified DNA.
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Kvist, T., Ahring, B. K., Lasken, R. S. & Westermann, P. Specific single-cell isolation and genomic amplification of uncultured microorganisms. Appl. Microbiol. Biotechnol. 74, 926–935 (2007).
Blainey, P. C., Mosier, A. C., Potanina, A., Francis, C. A. & Quake, S. R. Genome of a low-salinity ammonia-oxidizing archaeon determined by single-cell and metagenomic analysis. PLoS ONE 6, e16626 (2011).
Podar, M. et al. Targeted access to the genomes of low abundance organisms in complex microbial communities. Appl. Environ. Microbiol. 73, 3205–3214 (2007).
Marcy, Y. et al. Dissecting biological “dark matter” with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc. Natl Acad. Sci. USA 104, 11889–118894 (2007). Reference 44 and 45 both sequenced TM7, a phylum lacking any sequenced members.
Mußmann, M. et al. Insights into the genome of large sulphur bacteria revealed by analysis of single filaments. PLoS Biol. 5, e230 (2007).
Winogradsky, S. Beiträge zur Morphologie undPphysiologie der Bakterien. Zur Morphologie und Physiologie der Schwefelbakterien Vol. 1, 1–120 (Arthur Felix,1888).
Stepanauskas, R. & Sieracki, M. E. Matching phylogeny and metabolism in the uncultured marine bacteria, one cell at a time. Proc. Natl Acad. Sci. USA 104, 9052–9057 (2007).
Giovannoni, S. J. et al. Genome streamlining in a cosmopolitan oceanic bacterium. Science 309, 1242–1245 (2005).
Martinez-Garcia, M. et al. High-throughput single-cell sequencing identifies photoheterotrophs and chemoautotrophs in freshwater bacterioplankton. ISME J. 6, 113–123 (2012).
Swan, B. K. et al. Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the dark ocean. Science 333, 1296–1300 (2011).
Chang, H. W. et al. Development of microbial genome-probing microarrays using digital multiple displacement amplification of uncultivated microbial single cells. Environ. Sci. Technol. 42, 6058–6064 (2008).
Fleming, E. J. et al. What's new is old: resolving the identity of Leptothrix ochracea using single cell genomics, pyrosequencing and FISH. PLoS ONE 6, e17769 (2011).
Ottesen, E. A., Hong, J. W., Quake, S. R. & Leadbetter, J. R. Microfluidic digital PCR enables multigene analysis of individual environmental bacteria. Science 314, 1464–1467 (2006).
Vogelstein, B. & Kinzler, K. W. Digital PCR. Proc. Natl Acad. Sci. USA 96, 9236–9241 (1999).
Worden, A. Z., Dupont, C. & Allen, A. E. Genomes of uncultured eukaryotes: sorting FACS from fiction. Genome Biol. 12, 117 (2011).
Cuveliera, M. L. et al. Targeted metagenomics and ecology of globally important uncultured eukaryotic phytoplankton. Proc. Natl Acad. Sci. USA 107, 14679–14684 (2010).
Heywood, J. L., Sieracki, M. E., Bellows, W., Poulton, N. J. & Stepanauskas, R. Capturing diversity of marine heterotrophic protists: one cell at a time. ISME J. 5, 674–684 (2011).
Martinez-Garcia, M. et al. Unveiling in situ interactions between marine protists and bacteria through single cell sequencing. ISME J. 6, 703–707 (2012).
Not, F. et al. Picobiliphytes: a marine picoplanktonic algal group with unknown affinities to other eukaryotes. Science 315, 253–255 (2007).
Grindberg, R. V. et al. Single cell genome amplification accelerates identification of the apratoxin biosynthetic pathway from a complex microbial assemblage. PLoS ONE 6, e1856 (2011). The use of single-cell sequencing to discover the gene cluster required for synthesis of a therapeutic agent.
Siegl, A. et al. Single-cell genomics reveals the lifestyle of Poribacteria, a candidate phylum symbiotically associated with marine sponges. ISME J. 5, 61–70 (2011).
Ehrlich, G. D. The distributed genome hypothesis as a rubric for understanding evolution in situ during chronic bacterial biofilm infectious processes. FEMS Immunol. Med. Microbiol. 59, 269–279 (2010).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2004).
Narasingarao, P. et al. De novo metagenomic assembly reveals abundant novel major lineage of Archaea in hypersaline microbial communities. ISME J. 6, 81–93 (2011).
Medini, D., Donati, C., Tettelin, H., Masignani, V. & Rappuoli, R. The microbial pan-genome. Curr. Opin. Genet. Dev. 15, 589–594 (2005).
Dupont, C. L. et al. Genomic insights to SAR86, an abundant and uncultivated marine bacterial lineage. ISME J. 6, 1186–1199 (2011).
Hess, M. et al. Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331, 463–467 (2011).
Eloe, E. A. et al. Going deeper: metagenome of a hadopelagic microbial community. PLoS ONE 6, e20388 (2011).
Edwards, R. A. & Rohwer, F. Viral metagenomics. Nature Rev. Microbiol. 3, 504–510 (2005).
Rappe, M. S. & Giovannoni, S. J. The uncultured microbial majority. Annu. Rev. Microbiol. 57, 369–394 (2003).
Allen, L. Z. et al. Single virus genomics: a new tool for virus discovery. PLoS ONE 23, e17722 (2011).
Martínez Martínez, J., Poulton, N. J., Stepanauskas, R., Sieracki, M. E. & Wilson, W. H. Targeted sorting of single virus-infected cells of the coccolithophore Emiliania huxleyi. PLoS ONE 6, e22520 (2011).
Woyke, T. et al. Decontamination of MDA reagents for single cell whole genome amplification. PLoS ONE 6, e26161 (2011).
Abulencia, C. B. et al. Environmental whole-genome amplification to access microbial populations in contaminated sediments. Appl. Environ. Microbiol. 72, 3291–3301 (2006).
Yilmaz, S., Allgaier, M. & Hugenholtz, P. Multiple displacement amplification compromises quantitative analysis of metagenomes. Nature Methods 7, 943 (2010).
Chen, C. H. et al. Specific sorting of single bacterial cells with microfabricated fluorescence-activated cell sorting and tyramide signal amplification fluorescence in situ hybridization. Anal. Chem. 83, 7269–7275 (2011).
Pamp, S. J. et al. Single-cell sequencing provides clues about the host interactions of segmented filamentous bacteria (SFB). Genome Res. 22, 1107–1119 (2012).
Leung, K. et al. A programmable droplet-based microfluidic device applied to multiparameter analysis of single microbes and microbial communities. Proc. Natl Acad. Sci. USA 109, 7665–7670 (2012).
Dichosa, A. E. K. et al. Artificial polyploidy improves bacterial single cell genome recovery. PLoS ONE 7, e37387 (2012).
Martinez-Garcia, M. et al. Capturing single cell genomes of active polysaccharide degraders: an unexpected contribution of Verrucomicrobia. PLoS ONE 7, e35314 (2012).
Mason, O. U. et al. Metagenome, metatranscriptome and single-cell sequencing reveal microbial response to Deepwater Horizon oil spill. ISME J. 21 Jun 2012 (doi: 10.1038/ismej.2012.59).
Hou, Y. et al. Single-cell exome sequencing and monoclonal evolution of a JAK2-negative myeloproliferative neoplasm. Cell 148, 873–885 (2012).
Peters, B. A. et al. Accurate whole-genome sequencing and haplotyping from 10 to 20 human cells. Nature 487, 190–195 (2012).
Gibson, D. G. et al. Creation of a bacterial cell controlled by a chemically synthesized genome. Science 329, 52–56 (2010).
Acknowledgements
The author gratefully acknowledges C. Dupont for suggestions on metagenomics and marine communities.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- Global Ocean Sampling study
-
(GOS study). A global circumnavigation aboard the sailing yacht, Sorcerer II, conducted by the J. Craig Venter Institute (La Jolla, USA), with the aim of providing a comprehensive genomic survey of microbial (bacterial, archaeal and viral) life in the world's oceans.
- Klenow fragment
-
The large fragment of Escherichia coli DNA polymerase I; this fragment has a processivity of ∼10 nucleotides.
- Bar-coded DNA libraries
-
Mixed DNA libraries that are created by the addition of short DNA sequences (bar-codes) onto the DNA templates. Multiple DNA sets (for example, from different bacteria) are each tagged with a unique bar-code and then pooled for use in a single sequencing run. The sequences obtained can be separated back into the individual sets based on the bar-codes.
- Chemolithoautotrophic
-
Pertaining to an organism: deriving energy from a chemical reaction (chemotrophic) using inorganic substrates such as nitrate, ferrous iron or sulphur as electron donors (lithotrophic), and using CO2 as the sole carbon source (autotrophic).
- Pelagic
-
Relating to or occurring in the water column.
- Mesopelagic
-
Typically between 200m and 1,000m below the ocean surface.
- CARD–FISH
-
(Catalysed reporter deposition–fluorescence in situ hybridization). A method for increasing the sensitivity of FISH, for example, by using horseradish peroxidase-labelled oligonucleotide probes. The horseradish peroxidase catalyses the deposition of tyramine molecules, which results in amplification of the fluorescent signal at the site of probe hybridization.
- Phycobiliproteins
-
Water-soluble proteins that are present in cyanobacteria and certain algae and capture light energy, which is passed on to chlorophylls during photosynthesis. Phycobiliproteins are complexes consisting of proteins and covalently bound phycobilins that act as chromophores (capturing light).
Rights and permissions
About this article
Cite this article
Lasken, R. Genomic sequencing of uncultured microorganisms from single cells. Nat Rev Microbiol 10, 631–640 (2012). https://doi.org/10.1038/nrmicro2857
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2857
This article is cited by
-
Batch effects removal for microbiome data via conditional quantile regression
Nature Communications (2022)
-
Dissecting the dominant hot spring microbial populations based on community-wide sampling at single-cell genomic resolution
The ISME Journal (2022)
-
Prokaryotic and eukaryotic diversity in hydrothermal continental systems
Archives of Microbiology (2021)
-
Single-virus genomics and beyond
Nature Reviews Microbiology (2020)
-
Genome and pan-genome analysis to classify emerging bacteria
Biology Direct (2019)