Key Points
-
The toxin–antitoxin (TA) systems of bacteria are new-comers in microbiology and biochemistry, as most of the discoveries about these systems have been made in the past decade. To date, it has been revealed that most bacteria and many archaea contain a number of highly diverse TA systems in their genomes.
-
In the type II TA systems, a toxin is constitutively co-expressed with its cognate antitoxin from a TA operon to form a stable complex in normally growing cells. As the antitoxin is less stable than the toxin in the cell, antitoxin has to be constantly produced to neutralize the cognate toxin. Under stress conditions, antitoxins are degraded by stress-induced proteases to release free toxins in the cells, resulting in cell growth arrest and eventual cell death.
-
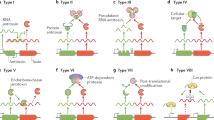
Cellular targets of TA systems are highly diverse, from DNA replication to mRNA stability, protein synthesis, ATP production and cell wall biosynthesis. Other cellular targets are likely to be identified as more TA systems are discovered. The most frequent cellular targets for the Escherichia coli TA systems are mRNAs, perhaps because the inhibition of mRNA function seems to be the mildest means of regulating cell growth. Out of 36 TA systems in E. coli, 11 are known to interfere with mRNA.
-
As toxins targeting mRNA cleave cellular mRNAs, they are termed mRNA interferases. These mRNA interferases are grouped into two distinct classes depending on how they cleave mRNAs: ribosome-independent mRNA interferases and ribosome-dependent mRNA interferases (which cleave mRNAs at the ribosomal A site).
-
It has been shown that some toxins are induced under several stress conditions, including amino acid starvation, glucose starvation, DNA damage, heat and the addition of antibiotics; these toxins regulate cell growth, triggering programmed cell death. It is also predicted that the induction of some toxins is able to cause the persistent or quasi-dormant state, making these cells resistant to antibiotics. The toxins may also play a part in eliminating damaged cells from their populations.
-
Elucidation of the functions of these toxins is essential for our understanding of their roles in bacterial physiology under various stress conditions and also their roles in bacterial pathogenicity. The study of TA systems opens the door into a new and exciting field in medical research, molecular biology and biotechnology, as various toxins from TA systems may be used as new therapeutic tools. TA systems may also provide novel technologies for gene regulation and protein production.
Abstract
Escherichia coli K-12 contains at least 36 toxin genes, the expression of which causes growth inhibition and eventual death. These toxins are usually co-expressed with their cognate antitoxins in operons called toxin–antitoxin (TA) modules. Under normal growth conditions, toxins and antitoxins form stable complexes. However, stress-induced proteases preferentially eliminate unstable antitoxins, releasing free toxins to inhibit various cellular functions. TA systems have important roles in the physiology of cells in their natural habitats, including functions in biofilm formation and multidrug resistance. In this Review, we describe these TA systems in light of their functions and roles in the regulation of cell growth and death.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gerdes, K., Christensen, S. K. & Lobner-Olesen, A. Prokaryotic toxin–antitoxin stress response loci. Nature Rev. Microbiol. 3, 371–382 (2005).
Hayes, F. Toxins-antitoxins: plasmid maintenance, programmed cell death, and cell cycle arrest. Science 301, 1496–1499 (2003).
Yamaguchi, Y. & Inouye, M. mRNA interferases, sequence-specific endoribonucleases from the toxin–antitoxin systems. Prog. Mol. Biol. Transl. Sci. 85, 467–500 (2009).
Sevin, E. W. & Barloy-Hubler, F. RASTA-Bacteria: a web-based tool for identifying toxin-antitoxin loci in prokaryotes. Genome Biol. 8, R155 (2007).
Shao, Y. et al. TADB: a web-based resource for Type 2 toxin–antitoxin loci in bacteria and archaea. Nucleic Acids Res. 39, D606–D611 (2011).
Bernard, P. & Couturier, M. Cell killing by the F plasmid CcdB protein involves poisoning of DNA-topoisomerase II complexes. J. Mol. Biol. 226, 735–745 (1992).
Gerdes, K. et al. Mechanism of postsegregational killing by the hok gene product of the parB system of plasmid R1 and its homology with the relF gene product of the E. coli relB operon. EMBO J. 5, 2023–2029 (1986).
Ogura, T. & Hiraga, S. Mini-F plasmid genes that couple host cell division to plasmid proliferation. Proc. Natl Acad. Sci. USA 80, 4784–4788 (1983).
Bernard, P. & Couturier, M. The 41 carboxy-terminal residues of the miniF plasmid CcdA protein are sufficient to antagonize the killer activity of the CcdB protein. Mol. Gen. Genet. 226, 297–304 (1991).
Thisted, T., Nielsen, A. K. & Gerdes, K. Mechanism of post-segregational killing: translation of Hok, SrnB and Pnd mRNAs of plasmids R1, F and R483 is activated by 3′-end processing. EMBO J. 13, 1950–1959 (1994).
Thisted, T., Sorensen, N. S., Wagner, E. G. & Gerdes, K. Mechanism of post-segregational killing: Sok antisense RNA interacts with Hok mRNA via its 5′-end single-stranded leader and competes with the 3′-end of Hok mRNA for binding to the mok translational initiation region. EMBO J. 13, 1960–1968 (1994).
Naito, T., Kusano, K. & Kobayashi, I. Selfish behavior of restriction-modification systems. Science 267, 897–899 (1995).
Yarmolinsky, M. B. Programmed cell death in bacterial populations. Science 267, 836–837 (1995).
Van Melderen, L. & Saavedra De Bast, M. Bacterial toxin–antitoxin systems: more than selfish entities? PLoS Genet. 5, e1000437 (2009).
Christensen-Dalsgaard, M., Jorgensen, M. G. & Gerdes, K. Three new RelE-homologous mRNA interferases of Escherichia coli differentially induced by environmental stresses. Mol. Microbiol. 75, 333–348 (2010).
Marianovsky, I., Aizenman, E., Engelberg-Kulka, H. & Glaser, G. The regulation of the Escherichia coli mazEF promoter involves an unusual alternating palindrome. J. Biol. Chem. 276, 5975–5984 (2001).
Yamaguchi, Y., Park, J. H. & Inouye, M. MqsR, a crucial regulator for quorum sensing and biofilm formation, is a GCU-specific mRNA interferase in Escherichia coli. J. Biol. Chem. 284, 28746–28753 (2009).
Gerdes, K. & Wagner, E. G. RNA antitoxins. Curr. Opin. Microbiol. 10, 117–124 (2007).
Fineran, P. C. et al. The phage abortive infection system, ToxIN, functions as a protein–RNA toxin–antitoxin pair. Proc. Natl Acad. Sci. USA 106, 894–899 (2009).
Zhang, Y., Zhang, J., Hara, H., Kato, I. & Inouye, M. Insights into the mRNA cleavage mechanism by MazF, an mRNA interferase. J. Biol. Chem. 280, 3143–3150 (2005).
Zhang, Y. et al. MazF cleaves cellular mRNAs specifically at ACA to block protein synthesis in Escherichia coli. Mol. Cell 12, 913–923 (2003).
Pedersen, K. et al. The bacterial toxin RelE displays codon-specific cleavage of mRNAs in the ribosomal A site. Cell 112, 131–140 (2003).
Takagi, H. et al. Crystal structure of archaeal toxin-antitoxin RelE–RelB complex with implications for toxin activity and antitoxin effects. Nature Struct. Mol. Biol. 12, 327–331 (2005).
Zhang, Y., Zhu, L., Zhang, J. & Inouye, M. Characterization of ChpBK, an mRNA interferase from Escherichia coli. J. Biol. Chem. 280, 26080–26088 (2005).
Motiejunaite, R., Armalyte, J., Markuckas, A. & Suziedeliene, E. Escherichia coli dinJ-yafQ genes act as a toxin–antitoxin module. FEMS Microbiol. Lett. 268, 112–119 (2007).
Prysak, M. H. et al. Bacterial toxin YafQ is an endoribonuclease that associates with the ribosome and blocks translation elongation through sequence-specific and frame-dependent mRNA cleavage. Mol. Microbiol. 71, 1071–1087 (2009).
Kamada, K. & Hanaoka, F. Conformational change in the catalytic site of the ribonuclease YoeB toxin by YefM antitoxin. Mol. Cell 19, 497–509 (2005).
Zhang, Y. & Inouye, M. The inhibitory mechanism of protein synthesis by YoeB, an Escherichia coli toxin. J. Biol. Chem. 284, 6627–6638 (2009).
Keren, I., Shah, D., Spoering, A., Kaldalu, N. & Lewis, K. Specialized persister cells and the mechanism of multidrug tolerance in Escherichia coli. J. Bacteriol. 186, 8172–8180 (2004).
Korch, S. B., Henderson, T. A. & Hill, T. M. Characterization of the hipA7 allele of Escherichia coli and evidence that high persistence is governed by (p)ppGpp synthesis. Mol. Microbiol. 50, 1199–1213 (2003).
Brown, J. M. & Shaw, K. J. A novel family of Escherichia coli toxin-antitoxin gene pairs. J. Bacteriol. 185, 6600–6608 (2003).
Zhang, Y., Yamaguchi, Y. & Inouye, M. Characterization of YafO, an Escherichia coli toxin. J. Biol. Chem. 284, 25522–25531 (2009).
Brown, B. L. et al. Three dimensional structure of the MqsR:MqsA complex: a novel TA pair comprised of a toxin homologous to RelE and an antitoxin with unique properties. PLoS Pathog. 5, e1000706 (2009).
Pandey, D. P. & Gerdes, K. Toxin–antitoxin loci are highly abundant in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 33, 966–976 (2005).
Jaffe, A., Ogura, T. & Hiraga, S. Effects of the ccd function of the F plasmid on bacterial growth. J. Bacteriol. 163, 841–849 (1985).
Miki, T., Park, J. A., Nagao, K., Murayama, N. & Horiuchi, T. Control of segregation of chromosomal DNA by sex factor F in Escherichia coli. Mutants of DNA gyrase subunit A suppress letD (ccdB) product growth inhibition. J. Mol. Biol. 225, 39–52 (1992).
Roberts, R. C., Strom, A. R. & Helinski, D. R. The parDE operon of the broad-host-range plasmid RK2 specifies growth inhibition associated with plasmid loss. J. Mol. Biol. 237, 35–51 (1994).
Garcia-Contreras, R., Zhang, X. S., Kim, Y. & Wood, T. K. Protein translation and cell death: the role of rare tRNAs in biofilm formation and in activating dormant phage killer genes. PLoS ONE 3, e2394 (2008).
Neubauer, C. et al. The structural basis for mRNA recognition and cleavage by the ribosome-dependent endonuclease RelE. Cell 139, 1084–1095 (2009).
Zhang, Y. & Inouye, M. RatA (YfjG), an Escherichia coli toxin, inhibits 70S ribosome association to block translation. 79, 1418–1429 (2011).
Tan, Q., Awano, N. & Inouye, M. YeeV is an Escherichia coli toxin that inhibits cell division by targeting the cytoskeleton proteins, FtsZ and MreB. Mol. Microbiol. 79, 109–118 (2011).
Mutschler, H., Gebhardt, M., Shoeman, R. L. & Meinhart, A. A novel mechanism of programmed cell death in bacteria by toxin–antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 9, e1001033 (2011).
Inouye, M. The discovery of mRNA interferases: implication in bacterial physiology and application to biotechnology. J. Cell Physiol. 209, 670–676 (2006).
Jorgensen, M. G., Pandey, D. P., Jaskolska, M. & Gerdes, K. HicA of Escherichia coli defines a novel family of translation-independent mRNA interferases in bacteria and archaea. J. Bacteriol. 191, 1191–1199 (2009).
Schmidt, O. et al. prlF and yhaV encode a new toxin–antitoxin system in Escherichia coli. J. Mol. Biol. 372, 894–905 (2007).
Koga, M., Otsuka, Y., Lemire, S. & Yonesaki, T. Escherichia coli rnlA and rnlB compose a novel toxin–antitoxin system. Genetics 187, 123–130 (2011).
Kamada, K., Hanaoka, F. & Burley, S. K. Crystal structure of the MazE/MazF complex: molecular bases of antidote-toxin recognition. Mol. Cell 11, 875–884 (2003).
Li, G. Y. et al. Characterization of dual substrate binding sites in the homodimeric structure of Escherichia coli mRNA interferase MazF. J. Mol. Biol. 357, 139–150 (2006).
Metzger, S. et al. The nucleotide sequence and characterization of the relA gene of Escherichia coli. J. Biol. Chem. 263, 15699–15704 (1988).
Aizenman, E., Engelberg-Kulka, H. & Glaser, G. An Escherichia coli chromosomal “addiction module” regulated by guanosine 3',5′-bispyrophosphate: a model for programmed bacterial cell death. Proc. Natl Acad. Sci. USA 93, 6059–6063 (1996).
Christensen, S. K., Pedersen, K., Hansen, F. G. & Gerdes, K. Toxin–antitoxin loci as stress-response-elements: ChpAK/MazF and ChpBK cleave translated RNAs and are counteracted by tmRNA. J. Mol. Biol. 332, 809–819 (2003).
Zhang, J., Zhang, Y. & Inouye, M. Characterization of the interactions within the mazEF addiction module of Escherichia coli. J. Biol. Chem. 278, 32300–32306 (2003).
Zhu, L. et al. The mRNA interferases, MazF-mt3 and MazF-mt7 from Mycobacterium tuberculosis target unique pentad sequences in single-stranded RNA. Mol. Microbiol. 69, 559–569 (2008).
Zhu, L. et al. Characterization of mRNA interferases from Mycobacterium tuberculosis. J. Biol. Chem. 281, 18638–18643 (2006).
Fu, Z., Donegan, N. P., Memmi, G. & Cheung, A. L. Characterization of MazFSa, an endoribonuclease from Staphylococcus aureus. J. Bacteriol. 189, 8871–8879 (2007).
Zhu, L. et al. Staphylococcus aureus MazF specifically cleaves a pentad sequence, UACAU, which is unusually abundant in the mRNA for pathogenic adhesive factor SraP. J. Bacteriol. 191, 3248–3255 (2009).
Pellegrini, O., Mathy, N., Gogos, A., Shapiro, L. & Condon, C. The Bacillus subtilis ydcDE operon encodes an endoribonuclease of the MazF/PemK family and its inhibitor. Mol. Microbiol. 56, 1139–1148 (2005).
Park, J. H., Yamaguchi, Y. & Inouye, M. Bacillus subtilis MazF-bs (EndoA) is a UACAU-specific mRNA interferase. FEBS Lett. 585, 2526–2532 (2011).
Nariya, H. & Inouye, M. MazF, an mRNA interferase, mediates programmed cell death during multicellular Myxococcus development. Cell 132, 55–66 (2008).
Ren, D., Bedzyk, L. A., Thomas, S. M., Ye, R. W. & Wood, T. K. Gene expression in Escherichia coli biofilms. Appl. Microbiol Biotechnol. 64, 515–524 (2004).
Gonzalez Barrios, A. F. et al. Autoinducer 2 controls biofilm formation in Escherichia coli through a novel motility quorum-sensing regulator (MqsR, B3022). J. Bacteriol. 188, 305–316 (2006).
Surette, M. G., Miller, M. B. & Bassler, B. L. Quorum sensing in Escherichia coli, Salmonella typhimurium, and Vibrio harveyi: a new family of genes responsible for autoinducer production. Proc. Natl Acad. Sci. USA 96, 1639–1644 (1999).
Sperandio, V., Torres, A. G. & Kaper, J. B. Quorum sensing Escherichia coli regulators B and C (QseBC): a novel two-component regulatory system involved in the regulation of flagella and motility by quorum sensing in E. coli. Mol. Microbiol. 43, 809–821 (2002).
Chilcott, G. S. & Hughes, K. T. Coupling of flagellar gene expression to flagellar assembly in Salmonella enterica serovar typhimurium and Escherichia coli. Microbiol. Mol. Biol. Rev. 64, 694–708 (2000).
Hayes, C. S. & Sauer, R. T. Cleavage of the A site mRNA codon during ribosome pausing provides a mechanism for translational quality control. Mol. Cell 12, 903–911 (2003).
Makarova, K. S., Grishin, N. V. & Koonin, E. V. The HicAB cassette, a putative novel, RNA-targeting toxin-antitoxin system in archaea and bacteria. Bioinformatics 22, 2581–2584 (2006).
Button, J. E., Silhavy, T. J. & Ruiz, N. A suppressor of cell death caused by the loss of σE downregulates extracytoplasmic stress responses and outer membrane vesicle production in Escherichia coli. J. Bacteriol. 189, 1523–1530 (2007).
Hurley, J. M., Cruz, J. W., Ouyang, M. & Woychik, N. A. Bacterial toxin RelE mediates frequent codon-independent mRNA cleavage from the 5′ end of coding regions in vivo. J. Biol. Chem. 286, 14770–14778 (2011).
Christensen, S. K. & Gerdes, K. RelE toxins from Bacteria and Archaea cleave mRNAs on translating ribosomes, which are rescued by tmRNA. Mol. Microbiol. 48, 1389–1400 (2003).
Li, G. Y., Zhang, Y., Inouye, M. & Ikura, M. Structural mechanism of transcriptional autorepression of the Escherichia coli RelB/RelE antitoxin/toxin module. J. Mol. Biol. 380, 107–119 (2008).
Christensen, S. K. et al. Overproduction of the Lon protease triggers inhibition of translation in Escherichia coli: involvement of the yefM-yoeB toxin-antitoxin system. Mol. Microbiol. 51, 1705–1717 (2004).
Christensen-Dalsgaard, M., Overgaard, M., Winther, K. S. & Gerdes, K. RNA decay by messenger RNA interferases. Methods Enzymol. 447, 521–535 (2008).
Winther, K. S. & Gerdes, K. Ectopic production of VapCs from Enterobacteria inhibits translation and trans-activates YoeB mRNA interferase. Mol. Microbiol. 72, 918–930 (2009).
McKenzie, G. J., Magner, D. B., Lee, P. L. & Rosenberg, S. M. The dinB operon and spontaneous mutation in Escherichia coli. J. Bacteriol. 185, 3972–3977 (2003).
Brotcorne-Lannoye, A. & Maenhaut-Michel, G. Role of RecA protein in untargeted UV mutagenesis of bacteriophage λ: evidence for the requirement for the dinB gene. Proc. Natl Acad. Sci. USA 83, 3904–3908 (1986).
Courcelle, J., Khodursky, A., Peter, B., Brown, P. O. & Hanawalt, P. C. Comparative gene expression profiles following UV exposure in wild-type and SOS-deficient Escherichia coli. Genetics 158, 41–64 (2001).
Kolodkin-Gal, I., Verdiger, R., Shlosberg-Fedida, A. & Engelberg-Kulka, H. A differential effect of E. coli toxin-antitoxin systems on cell death in liquid media and biofilm formation. PLoS ONE 4, e6785 (2009).
Lewis, L. K., Harlow, G. R., Gregg-Jolly, L. A. & Mount, D. W. Identification of high affinity binding sites for LexA which define new DNA damage-inducible genes in Escherichia coli. J. Mol. Biol. 241, 507–523 (1994).
Fernandez De Henestrosa, A. R. et al. Identification of additional genes belonging to the LexA regulon in Escherichia coli. Mol. Microbiol. 35, 1560–1572 (2000).
Sat, B. et al. Programmed cell death in Escherichia coli: some antibiotics can trigger mazEF lethality. J. Bacteriol. 183, 2041–2045 (2001).
Hazan, R., Sat, B. & Engelberg-Kulka, H. Escherichia coli mazEF-mediated cell death is triggered by various stressful conditions. J. Bacteriol. 186, 3663–3669 (2004).
Belitsky, M. et al. The Escherichia coli extracellular death factor EDF induces the endoribonucleolytic activities of the toxins MazF and ChpBK. Mol. Cell 41, 625–635 (2011).
Kolodkin-Gal, I. & Engelberg-Kulka, H. The extracellular death factor: physiological and genetic factors influencing its production and response in Escherichia coli. J. Bacteriol. 190, 3169–3175 (2008).
Kolodkin-Gal, I., Hazan, R., Gaathon, A., Carmeli, S. & Engelberg-Kulka, H. A linear pentapeptide is a quorum-sensing factor required for mazEF-mediated cell death in Escherichia coli. Science 318, 652–655 (2007).
Kolodkin-Gal, I., Sat, B., Keshet, A. & Engelberg-Kulka, H. The communication factor EDF and the toxin–antitoxin module mazEF determine the mode of action of antibiotics. PLoS Biol. 6, e319 (2008).
Amitai, S., Kolodkin-Gal, I., Hananya-Meltabashi, M., Sacher, A. & Engelberg-Kulka, H. Escherichia coli MazF leads to the simultaneous selective synthesis of both “death proteins” and “survival proteins”. PLoS Genet. 5, e1000390 (2009).
Kohanski, M. A., Dwyer, D. J., Wierzbowski, J., Cottarel, G. & Collins, J. J. Mistranslation of membrane proteins and two-component system activation trigger antibiotic-mediated cell death. Cell 135, 679–690 (2008).
Hong, J., Ahn, J. M., Kim, B. C. & Gu, M. B. Construction of a functional network for common DNA damage responses in Escherichia coli. Genomics 93, 514–524 (2009).
Suzuki, M., Zhang, J., Liu, M., Woychik, N. A. & Inouye, M. Single protein production in living cells facilitated by an mRNA interferase. Mol. Cell 18, 253–261 (2005).
Shinagawa, H. SOS response as an adaptive response to DNA damage in prokaryotes. EXS 77, 221–235 (1996).
Butala, M., Zgur-Bertok, D. & Busby, S. J. The bacterial LexA transcriptional repressor. Cell. Mol. Life Sci. 66, 82–93 (2009).
Kawano, M., Aravind, L. & Storz, G. An antisense RNA controls synthesis of an SOS-induced toxin evolved from an antitoxin. Mol. Microbiol. 64, 738–754 (2007).
Pedersen, K. & Gerdes, K. Multiple hok genes on the chromosome of Escherichia coli. Mol. Microbiol. 32, 1090–1102 (1999).
Vogel, J., Argaman, L., Wagner, E. G. & Altuvia, S. The small RNA IstR inhibits synthesis of an SOS-induced toxic peptide. Curr. Biol. 14, 2271–2276 (2004).
Cairns, J. & Foster, P. L. Adaptive reversion of a frameshift mutation in Escherichia coli. Genetics 128, 695–701 (1991).
Harris, R. S., Longerich, S. & Rosenberg, S. M. Recombination in adaptive mutation. Science 264, 258–260 (1994).
Harris, R. S., Ross, K. J. & Rosenberg, S. M. Opposing roles of the holliday junction processing systems of Escherichia coli in recombination-dependent adaptive mutation. Genetics 142, 681–691 (1996).
Foster, P. L., Trimarchi, J. M. & Maurer, R. A. Two enzymes, both of which process recombination intermediates, have opposite effects on adaptive mutation in Escherichia coli. Genetics 142, 25–37 (1996).
Singletary, L. A. et al. An SOS-regulated type 2 toxin-antitoxin system. J. Bacteriol. 191, 7456–7465 (2009).
Opperman, T., Murli, S., Smith, B. T. & Walker, G. C. A model for a umuDC-dependent prokaryotic DNA damage checkpoint. Proc. Natl Acad. Sci. USA 96, 9218–9223 (1999).
Kim, Y., Wang, X., Ma, Q., Zhang, X. S. & Wood, T. K. Toxin-antitoxin systems in Escherichia coli influence biofilm formation through YjgK (TabA) and fimbriae. J. Bacteriol. 191, 1258–1267 (2009).
Harrison, J. J. et al. The chromosomal toxin gene yafQ is a determinant of multidrug tolerance for Escherichia coli growing in a biofilm. Antimicrob. Agents Chemother. 53, 2253–2258 (2009).
Hoffman, L. R. et al. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 436, 1171–1175 (2005).
Mulvey, M. R. & Loewen, P. C. Nucleotide sequence of katF of Escherichia coli suggests KatF protein is a novel σ transcription factor. Nucleic Acids Res. 17, 9979–9991 (1989).
Tanaka, K., Takayanagi, Y., Fujita, N., Ishihama, A. & Takahashi, H. Heterogeneity of the principal σ factor in Escherichia coli: the rpoS gene product, σ38, is a second principal σ factor of RNA polymerase in stationary-phase Escherichia coli. Proc. Natl Acad. Sci. USA 90, 3511–3515 (1993).
Ahmad, S. I., Kirk, S. H. & Eisenstark, A. Thymine metabolism and thymineless death in prokaryotes and eukaryotes. Annu. Rev. Microbiol. 52, 591–625 (1998).
Sat, B., Reches, M. & Engelberg-Kulka, H. The Escherichia coli mazEF suicide module mediates thymineless death. J. Bacteriol. 185, 1803–1807 (2003).
Engelberg-Kulka, H., Sat, B., Reches, M., Amitai, S. & Hazan, R. Bacterial programmed cell death systems as targets for antibiotics. Trends Microbiol. 12, 66–71 (2004).
Fonville, N. C., Bates, D., Hastings, P. J., Hanawalt, P. C. & Rosenberg, S. M. Role of RecA and the SOS response in thymineless death in Escherichia coli. PLoS Genet. 6, e1000865 (2010).
Foti, J. J., Schienda, J., Sutera, V. A. Jr & Lovett, S. T. A bacterial G protein-mediated response to replication arrest. Mol. Cell 17, 549–560 (2005).
Godoy, V. G., Jarosz, D. F., Walker, F. L., Simmons, L. A. & Walker, G. C. Y-family DNA polymerases respond to DNA damage-independent inhibition of replication fork progression. EMBO J. 25, 868–879 (2006).
Imlay, J. A., Chin, S. M. & Linn, S. Toxic DNA damage by hydrogen peroxide through the Fenton reaction in vivo and in vitro. Science 240, 640–642 (1988).
Imlay, J. A. & Linn, S. Bimodal pattern of killing of DNA-repair-defective or anoxically grown Escherichia coli by hydrogen peroxide. J. Bacteriol. 166, 519–527 (1986).
Davies, B. W. et al. Hydroxyurea induces hydroxyl radical-mediated cell death in Escherichia coli. Mol. Cell 36, 845–860 (2009).
Dukan, S. et al. Protein oxidation in response to increased transcriptional or translational errors. Proc. Natl Acad. Sci. USA 97, 5746–5749 (2000).
Kohanski, M. A., Dwyer, D. J., Hayete, B., Lawrence, C. A. & Collins, J. J. A common mechanism of cellular death induced by bactericidal antibiotics. Cell 130, 797–810 (2007).
Schumacher, M. A. et al. Molecular mechanisms of HipA-mediated multidrug tolerance and its neutralization by HipB. Science 323, 396–401 (2009).
Kim, Y. & Wood, T. K. Toxins Hha and CspD and small RNA regulator Hfq are involved in persister cell formation through MqsR in Escherichia coli. Biochem. Biophys. Res. Commun. 391, 209–213 (2010).
Shah, D. et al. Persisters: a distinct physiological state of E. coli. BMC Microbiol. 6, 53 (2006).
Christensen, S. K., Mikkelsen, M., Pedersen, K. & Gerdes, K. RelE, a global inhibitor of translation, is activated during nutritional stress. Proc. Natl Acad. Sci. USA 98, 14328–14333 (2001).
Sunohara, T., Jojima, K., Tagami, H., Inada, T. & Aiba, H. Ribosome stalling during translation elongation induces cleavage of mRNA being translated in Escherichia coli. J. Biol. Chem. 279, 15368–15375 (2004).
Sunohara, T., Jojima, K., Yamamoto, Y., Inada, T. & Aiba, H. Nascent-peptide-mediated ribosome stalling at a stop codon induces mRNA cleavage resulting in nonstop mRNA that is recognized by tmRNA. RNA 10, 378–386 (2004).
Li, X., Yagi, M., Morita, T. & Aiba, H. Cleavage of mRNAs and role of tmRNA system under amino acid starvation in Escherichia coli. Mol. Microbiol. 68, 462–473 (2008).
Zhu, L., Sharp, J. D., Kobayashi, H., Woychik, N. A. & Inouye, M. Noncognate Mycobacterium tuberculosis toxin-antitoxins can physically and functionally interact. J. Biol. Chem. 285, 39732–39738 (2010).
Kim, Y. et al. Escherichia coli toxin/antitoxin pair MqsR/MqsA regulate toxin CspD. Environ. Microbiol. 12, 1105–1121 (2010).
Mao, L. et al. Production of membrane proteins for NMR studies using the condensed single protein (cSPP) production system. J. Struct. Funct. Genomics 10, 281–289 (2009).
Shimazu, T. et al. NBK/BIK antagonizes MCL-1 and BCL-XL and activates BAK-mediated apoptosis in response to protein synthesis inhibition. Genes Dev. 21, 929–941 (2007).
Chono, H. et al. Acquisition of HIV-1 Resistance in T lymphocytes using an ACA-specific E. coli mRNA interferase. Hum. Gene Ther. 22, 35–43 (2011).
Goldman, E. & Jakubowski, H. Uncharged tRNA, protein synthesis, and the bacterial stringent response. Mol. Microbiol. 4, 2035–2040 (1990).
Kuroda, A. et al. Role of inorganic polyphosphate in promoting ribosomal protein degradation by the Lon protease in E. coli. Science 293, 705–708 (2001).
Acknowledgements
The authors are grateful to S. Phadtare and C. J. Mozdzierz for their critical reading of this article. This work was partially supported by a US National Institutes of Health grant (1RO1GM081567).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Antisense RNA
-
A complementary RNA sequence that binds to mRNA.
- Autoinducer
-
A signalling molecule that causes the regulation of specific genes.
- Outer-membrane vesicles
-
Enclosed compartments that are separated from the outer membrane in Gram-negative bacteria.
- Fruiting body
-
A specialized spore-producing structure.
- Stress-induced mutagenesis
-
The stress-induced reversible activation of error-prone DNA polymerases and downregulation of error-correcting enzymes, the result of which is an increased mutation rate in bacteria.
- Stringent response
-
The physiological changes that are caused by amino acid starvation.
Rights and permissions
About this article
Cite this article
Yamaguchi, Y., Inouye, M. Regulation of growth and death in Escherichia coli by toxin–antitoxin systems. Nat Rev Microbiol 9, 779–790 (2011). https://doi.org/10.1038/nrmicro2651
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2651
This article is cited by
-
Molecular determination of O25b/ST131 clone type among extended spectrum β-lactamases production Escherichia coli recovering from urinary tract infection isolates
Annals of Clinical Microbiology and Antimicrobials (2022)
-
Quantifying heterologous gene expression during ectopic MazF production in Escherichia coli
BMC Research Notes (2022)
-
The DarTG toxin-antitoxin system provides phage defence by ADP-ribosylating viral DNA
Nature Microbiology (2022)
-
Streptococcal bacterial components in cancer therapy
Cancer Gene Therapy (2022)
-
Fusion of the N-terminal 119 amino acids of RelA with the CTD domain render growth inhibitory effects of the latter, (p)ppGpp-dependent
Molecular Genetics and Genomics (2022)