Key Points
-
Studies of the Archaea have had a substantial impact on the field of biology.
-
Carl Woese carried out several pioneering studies, examining the evolution of the genetic code, the translation apparatus and the cell as a whole, and the insight gained from his early work led to the discovery of the third domain.
-
Defining moments in the history of archaeal research include the construction of a universal tree of life and the rationalization of the phylogeny of small-subunit ribosomal RNA, aminoacyl-tRNA synthetases and other protein sequences, leading to the three-domain view of life.
-
Seminal advances were made through probing the biochemistry, physiology, genetics and evolution of methanogens, extreme halophiles and thermoacidophiles.
-
The nature of archaea as extremophiles can now be placed into context with the knowledge that archaea are abundant and ubiquitous throughout the Earth's biosphere, including in the vast cold reaches of the planet.
-
Archaea play a key part in maintaining important biogeochemical cycles, and several contemporary advances have been made in our understanding of the importance of archaea in global ecology.
-
Technological advances have greatly enhanced the output from the field; particularly noteworthy are the impact of DNA sequencing, and the dawning of the genomics era and application of metagenomics, as well as breakthroughs that have been made through the development of tractable genetic systems.
Abstract
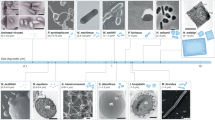
The Archaea evolved as one of the three primary lineages several billion years ago, but the first archaea to be discovered were described in the scientific literature about 130 years ago. Moreover, the Archaea were formally proposed as the third domain of life only 20 years ago. Over this very short period of investigative history, the scientific community has learned many remarkable things about the Archaea — their unique cellular components and pathways, their abundance and critical function in diverse natural environments, and their quintessential role in shaping the evolutionary path of life on Earth. This Review charts the 'archaea movement', from its genesis through to key findings that, when viewed together, illustrate just how strongly the field has built on new knowledge to advance our understanding not only of the Archaea, but of biology as a whole.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Cavicchioli, R. (ed) Archaea: Molecular and Cellular Biology (American Society for Microbiology Press, Washington DC, 2007). A strong collection of chapters from leaders in the field, including a testament about the discovery of the Archaea by Carl Woese.
Garrett, R. & Klenk, H.-P. Archaea: Evolution, Physiology and Molecular Biology (Blackwell, Oxford, UK, 2007).
Rothschild, L. J. & Mancinelli, R. L. Life in extreme environments. Nature 409, 1092–1101 (2001).
Kletzin, A. in Archaea: Molecular and Cellular Biology (ed. Cavicchioli, R.) 14–92 (American Society for Microbiology Press, Washington DC, 2007). A comprehensive overview of the microbiology, biochemistry and ecology of many types of archaea.
Makarova, K. S., Yutin, N., Bell, S. D. & Koonin, E. V. Evolution of diverse cell division and vesicle formation systems in Archaea. Nature Rev. Microbiol. 8, 731–741 (2010).
Gribaldo, S., Poole, A. M., Daubin, V., Forterre, P. & Brochier-Armanet, C. The origin of eukaryotes and their relationship with the Archaea: are we at a phylogenomic impasse? Nature Rev. Microbiol. 8, 743–752 (2010).
Walsby, A. E. A square bacterium. Nature 283, 69–71 (1980).
Burns, D. G. et al. Haloquadratum walsbyi gen. nov., sp. nov., the square haloarchaeon of Walsby, isolated from saltern crystallizers in Australia and Spain. Int. J. Syst. Evol. Microbiol. 57, 387–392 (2007).
Soppa, J. From genomes to function: haloarchaea as model organisms. Microbiology 152, 585–590 (2006).
Cavicchioli, R., Curmi, P. M. G., Saunders, N. & Thomas, T. Pathogenic archaea: do they exist? Bioessays 25, 1119–1128 (2003). A discussion about the concept of archaeal pathogens.
Lepp, P. W. et al. Methanogenic Archaea and human periodontal disease. Proc. Natl Acad. Sci. USA 101, 6176–6181 (2004).
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010).
Brulc, J. M. et al. Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc. Natl Acad. Sci. USA 106, 1948–1953 (2009).
Burkhardt, F. & Smith, S. (eds) The Correspondence of Charles Darwin:1856–1857 Vol. 6 (Cambridge Univ. Press, Cambridge, UK, 1990).
Woese, C. R. in Organization and Control in Prokaryotic and Eukaryotic Cells. 20th Symposium, Society for General Microbiology (eds Charles, H. P. & Knight, B. C. J. G.) 39–54 (Cambridge Univ. Press, Cambridge, UK (1970).
Woese, C. R., Sogin, M. L. & Sutton, L. A. Procaryote phylogeny. I. Concerning the relatedness of Aerobacter aerogenes to Escherichia coli. J. Mol. Evol. 3, 293–299 (1974).
Crick, F. H. C. The biological replication of macromolecules. Symp. Soc. Exp. Biol. 12, 138–163 (1958).
Zuckerkandl, E. & Pauling, L. Molecules as documents of evolutionary history. J. Theor. Biol. 8, 357–366 (1965).
Fox, G. E., Magrum, L. J., Balch, W. E., Wolfe, R. S. & Woese, C. R. Classification of methanogenic bacteria by 16S ribosomal RNA characterization. Proc. Natl Acad. Sci. USA 74, 4537–4541 (1977). A cornerstone of the discovery of the third domain: the first SSU rRNA sequences of methanogens.
Woese, C. R. & Fox, G. E. The phylogenetic structure of the procaryotic domain: the primary kingdoms. Proc. Natl Acad. Sci. USA 74, 5088–5090 (1977). The quintessential description of the third domain that started it all.
Woese, C. R. & Fox, G. E. The concept of cellular evolution. J. Mol. Evol. 10, 1–6 (1977). An important paper describing the importance of the genotype–phenotype relationship in the evolution of life.
Woese, C. R., Kandler, O. & Wheelis, M. L. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc. Natl Acad. Sci. USA 87, 4576–4579 (1990). The proposal for the formal description of the three domains — a must-read for all educational purposes.
Woese, C. R. in Archaea: Molecular and Cellular Biology (ed. Cavicchioli, R.) 1–13 (American Society for Microbiology Press, Washington DC, 2007). A provocative testament about the archaea movement.
Cavalier-Smith, T. The neomuran origin of archaebacteria, the negibacterial root of the universal tree and bacterial megaclassification. Int. J. Syst. Evol. Microbiol. 52, 7–76 (2002).
Simonson, A. B. et al. Decoding the genomic tree of life. Proc. Natl Acad. Sci. USA 102, 6608–6613 (2005).
Sapp, J. The prokaryote-eukaryote dichotomy: meanings and mythology. Microbiol. Mol. Biol. Rev. 69, 292–305 (2005). An extensive and revealing review about the archaea movement.
Friend, T. The Third Domain: the Untold Story of Archaea and the Future of Biotechnology. (Joseph Henry, Washington DC, 2007). An enjoyable, easy-to-read and insightful book about the history of the discovery of the Archaea.
Woese, C. R. & Goldenfeld, N. How the microbial world saved evolution from the Scylla of molecular biology and the Charybdis of the modern synthesis. Microbiol. Mol. Biol. Rev. 73, 14–21 (2009). A contemporary essay concerning how the evolutionary process has been studied in the twentieth century.
Iwabe, N., Kuma, K., Hasegawa, M., Osawa, S. & Miyata, T. Evolutionary relationship of archaebacteria, eubacteria, and eukaryotes inferred from phylogenetic trees of duplicated genes. Proc. Natl Acad. Sci. USA 86, 9355–9359 (1989).
Brown, J. R. & Doolittle, W. F. Archaea and the prokaryote-to-eukaryote transition. Microbiol. Mol. Biol. Rev. 61, 456–502 (1997). An extensive review of the evolution of the universal tree of life, with particular emphasis on protein phylogeny.
Woese, C. R. On the evolution of cells. Proc. Natl Acad. Sci. USA 99, 8742–8747 (2002).
Vetsigian, K., Woese, C. R. & Goldenfeld, N. Collective evolution and the genetic code. Proc. Natl Acad. Sci. USA 103, 10696–10701 (2006). A major advance in considering the role of Darwinian versus Lamarckian influences on the evolution of the genetic code, the translation process and cellular organization.
Woese, C. R., Olsen, G. J., Ibba, M. & Söll, D. Aminoacyl-tRNA synthetases, the genetic code, and the evolutionary process. Microbiol. Mol. Biol. Rev. 64, 202–236 (2000). A comprehensive review placing the evolution of aaRSs in the context of the rest of the translation apparatus.
Woese, C. R. A new biology for a new century. Microbiol. Mol. Biol. Rev. 68, 173–186 (2004). A reflection on the scientific process involved in revealing the Archaea to the scientific community.
Kavran, J. M. et al. Structure of pyrrolysyl-tRNA synthetase, an archaeal enzyme for genetic code innovation. Proc. Natl Acad. Sci. USA 104, 11268–11273 (2007).
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase-tRNA(Pyl) structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2009).
Fournier, G. P., Huang, J. & Gogarten, J. P. Horizontal gene transfer from extinct and extant lineages: biological innovation and the coral of life. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 364, 2229–2239 (2009). A provocative view of the evolution of life (in particular, concerning what might be inferred about extinct lineages).
Pace, N. R. A molecular view of microbial diversity and the biosphere. Science 276, 734–740 (1997). An insightful review capturing the excitement emerging from the use of DNA sequence data to reveal the extent of the previously unknown phylogenetic and functional diversity in the microbial world.
Pace, N. R. Mapping the tree of life: progress and prospects. Microbiol. Mol. Biol. Rev. 73, 565–576 (2009). An up-to-date review about tree construction using SSU rRNA sequences and inferences about the universal tree of life.
Huber, H. et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417, 63–67 (2002). Insight into an interesting new lifestyle as well as a new division within the Archaea.
Elkins, J. G. et al., A korarchaeal genome reveals insights into the evolution of the Archaea. Proc. Natl Acad. Sci. USA 105, 8102–8107 (2008). Insight into an interesting new division within the Archaea.
Brochier-Armanet, C., Boussau, B., Gribaldo, S. & Forterre, P. Mesophilic crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nature Rev. Microbiol. 6, 245–252 (2008).
Casanueva, A. et al. Nanoarchaeal 16S rRNA gene sequences are widely dispersed in hyperthermophilic and mesophilic halophilic environments. Extremophiles 12, 651–656 (2008).
Robertson, C. E., Harris, J. K., Spear, J. R. & Pace, N. R. Phylogenetic diversity and ecology of environmental Archaea. Curr. Opin. Microbiol. 8, 638–642 (2005).
Ferry, J. G. Methanogenesis: Ecology, Physiology, Biochemistry and Genetics. Chapman and Hall Microbiology Series (Chapman and Hall Inc., New York, 1993). A compendium describing essentially everything that was known about methanogens in 1993.
Sohngen, N. L. Ueber bakterien, welche methan als kohlenstoffnahrung and energiequelle gebrauchen. Zentralbl. Bakteriol. Parasitik. Abt. I. 15, 513–517 (1906) (in German).
Wolfe, R. S. The sixth A. J. Kluyver Memorial Lecture. Methanogens: a surprising microbial group. Antonie Van Leeuwenhoek 45, 353–364 (1979).
Dalton, H. The Leeuwenhoek Lecture 2000: the natural and unnatural history of methane-oxidizing bacteria. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 360, 1207–1222 (2005).
Barker, H. A. Studies upon the methane-producing bacteria. Arch. Mikrobiol. 7, 420–438 (1936).
Kluyver, A. J. & van Niel, C. B. Prospects for a natural system of classification of bacteria. Zentralbl. Bakterol. Parasitenkd. Infectinskr. Hyg. Abt. II 94, 369–403 (1936).
Barker, H. A. Studies upon the methane fermentation. IV. The isolation and culture of Methanobacterium omelianskii. Antonie van Leeuwenhoek J. Microbiol. Serol. 6, 201–220 (1940).
Kandler, O. & Hippe, H. Lack of peptidoglycan in the cell walls of Methanosarcina barkeri. Arch. Microbiol. 113, 57–60 (1977).
Murray, P. A. & Zinder, S. Z. Nitrogen fixation by a methanogenic archaebacterium. Nature 312, 284–286 (1984).
Belay, N., Sparkling, R. & Daniels, L. Dintrogen fixation by a thermophilic methanogenic bacterium. Nature 312, 286–288 (1984).
DiMarco, A. A., Bobik, T. A. & Wolfe, R. S. Unusual coenzymes of methanogenesis. Annu. Rev. Biochem. 59, 355–394 (1990).
Ferry, J. G. How to make a living by exhaling methane. Annu. Rev. Microbiol. 64, 453–473 (2010).
Scheller, S., Goenrich, M., Boecher, R., Thauer, R. K. & Jaun, B. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane. Nature 465, 606–608 (2010).
Bult, C. A. et al. Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science 273, 1058–1073 (1996). A historical, eye-opening article documenting the first genome sequence of an archaeon, and the third complete genome sequence of an organism.
Orphan, V. J., House, C. H., Hinrichs, K.-U., McKeegan, K. D. & DeLong, E. F. Multiple archaeal groups mediate methane oxidation in anoxic cold seep sediments. Proc. Natl Acad. Sci. USA 99, 7663–7668 (2002).
Knittel, K. & Boetius, A. Anaerobic oxidation of methane: progress with an unknown process. Annu. Rev. Microbiol. 63, 311–334 (2009). A comprehensive review of anaerobic methane oxidation.
Alperin, M. & Hoehler, T. The ongoing mystery of sea-floor methane. Science 329, 288–289 (2010).
Hinrichs, K. U., Hayes, J. M., Sylva, S. P., Brewer, P. G. & DeLong, E. F. Methane-consuming archaebacteria in marine sediments. Nature 398, 802–805 (1999). An important breakthrough in realizing a role for archaea in anaerobic methane oxidation.
Orphan, V. J., House, C. H., Hinrichs, K.-U., McKeegan, K. D. & DeLong, E. F. Methane-consuming archaea revealed by directly coupled isotopic and phylogenetic analysis. Science 293, 484–487 (2001).
Niemann, H. et al. Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443, 854–858 (2006).
Zehnder, A. J. & Brock, T. D. Methane formation and methane oxidation by methanogenic bacteria. J. Bacteriol. 137, 420–432 (1979).
Hallam, S. J. et al. Reverse methanogenesis: testing the hypothesis with environmental genomics. Science 305, 1457–1462 (2004). An illustration of how metagenomics can be used to tease out a previously unknown pathway when there is evidence that the process (in this case, anaerobic oxidation of methane) occurs.
Dekas, A. E., Poretsky, R. S. & Orphan, V. J. Deep-sea archaea fix and share nitrogen in methane-consuming microbial consortia. Science 326, 422–426 (2009).
Michaelis, W. et al. Microbial reefs in the Black Sea fueled by anaerobic oxidation of methane. Science 297, 1013–1015 (2002).
Raghoebarsing, A. A. et al. A microbial consortium couples anaerobic methane oxidation to denitrification. Nature 440, 918–921 (2006).
Ettwig, K. F. et al. Denitrifying bacteria anaerobically oxidize methane in the absence of Archaea. Environ. Microbiol. 10, 3164–3173 (2008).
Ettwig, K. F. et al. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464, 543–548 (2010). A lesson in what can be learned beyond the Archaea, from the Archaea.
Hao, B. et al. A new UAG-encoded residue in the structure of a methanogen methyltransferase. Science 296, 1462–1466 (2002). An important description of pyrrolysine and the mechanism of its incorporation into proteins.
Soares, J. A. et al. The residue mass of L-pyrrolysine in three distinct methylamine methyltransferases. J. Biol. Chem. 280, 36962–36969 (2005).
Krzycki, J. A. Function of genetically encoded pyrrolysine in corrinoid-dependent methylamine methyltransferases. Curr. Opin. Chem. Biol. 8, 484–491 (2004).
Heinemann, I. U. et al. The appearance of pyrrolysine in tRNAHis guanylyltransferase by neutral evolution. Proc. Natl Acad. Sci. USA 106, 21103–21108 (2009). An interesting description of what a novel amino acid and its cognate aaRSs can explain about evolution.
Farlow, W. G. in United States Commission of Fish and Fisheries Part IV. Report for The Commissioner 1878 969–973 (Government Printing Office, Washington DC, 1880).
Kocur, M. & Hodgkiss, W. Taxonomic status of the genus Halococcus Schoop. Int. J. Syst. Bacteriol. 23, 151–156 (1973).
Sehgal, S. N., Kates, M. & Gibbons, N. E. Lipids of Halobacterium cutirubrum. Can. J. Biochem. Physiol. 40, 69–81 (1962).
Marshall, C. L. & Brown, A. D. The membrane lipids of Halobacterium halobium. Biochem. J. 110, 441–448 (1968).
Fox, G. E. et al. The phylogeny of prokaryotes. Science 209, 457–463 (1980).
Oesterhelt, D. & Stoeckenius, W. Rhodopsin-like protein from the purple membrane of Halobacterium hatobium. Nature New Biol. 233, 149–152 (1971).
Oesterhelt, D. & Stoeckenius, W. Functions of a new photoreceptor membrane. Proc. Natl Acad. Sci. USA 70, 2853–2857 (1973).
Henderson, R. & Unwin, P. N. T. Three-dimensional model of purple membrane obtained by electron microscopy. Nature 257, 28–32 (1975). An important breakthrough, not only for research into light-harvesting proteins (bacteriorhodopsin), but also for structural biology studies of proteins that contain multiple transmembrane domains.
Spudich, J. L. & Bogomolni, R. A. Mechanism of colour discrimination by a bacterial sensory rhodopsin. Nature 312, 509–513 (1984).
Lanyi, J. K. Bacteriorhodopsin as a model for proton pumps. Nature 375, 461–463 (1995).
Kolbe, M., Besir, H., Essen, L. O. & Oesterhelt, D. Structure of the light-driven chloride pump halorhodopsin at 1.8 Å resolution. Science 288, 1390–1396 (2000).
Spudich, J. L., Yang, C. S., Jung, K. H. & Spudich, E. N. Retinylidene proteins: structures and functions from Archaea to humans. Annu. Rev. Cell. Dev. Biol. 16, 365–392 (2000).
Rasmussen, S. G. et al. Crystal structure of the human β2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007).
Overington, J. P., Al-Lazikani, B. & Hopkins, A. L. How many drug targets are there? Nature Rev. Drug Discov. 5, 993–936 (2006).
Béjà, O. et al. Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science 289, 1902–1906 (2000). An important study describing a previously unknown phototrophic system for marine heterotrophs.
Béjà, O., Spudich, E. N., Spudich, J. L., Leclerc, M. & DeLong, E. F. Proteorhodopsin phototrophy in the ocean. Nature 411, 786–789 (2001).
Venter, J. C. et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science 304, 66–74 (2004).
Frigaard, N. U., Martinez, A., Mincer, T. J. & DeLong, E. F. Proteorhodopsin lateral gene transfer between marine planktonic Bacteria and Archaea. Nature 439, 847–850 (2006).
Moran, M. A. & Miller, W. L. Resourceful heterotrophs make the most of light in the coastal ocean. Nature Rev. Microbiol. 5, 792–800 (2007).
Gómez-Consarnau, L. et al. Proteorhodopsin phototrophy promotes survival of marine bacteria during starvation. PLoS Biol. 8, e1000358 (2010).
DeLong, E. F. & Béjà, O. The light-driven proton pump proteorhodopsin enhances bacterial survival during tough times. PLoS Biol. 8, e1000359 (2010).
Torsvik, T. & Dundas, I. D. Bacteriophage of Halobacterium salinarium. Nature 248, 680–681 (1974).
Dyall-Smith, M., Tang, S-L. & Bath, C. Haloarchaeal viruses: how diverse are they? Res. Microbiol. 154, 309–313 (2003).
Prangishvili, D., Forterre, P. & Garrett, R. A. Viruses of the Archaea: a unifying view. Nature Rev. Microbiol. 4, 837–848 (2006).
Ortmann, A. C., Wiedenheft, B., Douglas, T. & Young, M. Hot crenarchaeal viruses reveal deep evolutionary connections. Nature Rev. Microbiol. 4, 520–528 (2006). This article, along with reference 99, reveals the diversity and extent of archaeal viruses.
Sapienza, C. & Doolittle, W. F. Unusual physical organization of the Halobacterium genome. Nature 295, 384–389 (1982). Early insight into genome rearrangement that led to theories about the role of LGT in evolution.
Ng, W. V. et al. Genome sequence of Halobacterium species NRC-1. Proc. Natl Acad. Sci. USA 97, 12176–12181 (2000).
Papke, R. T., Koenig, J. E., Rodríguez-Valera, F. & Doolittle, W. F. Frequent recombination in a saltern population of Halorubrum. Science 306, 1928–1929 (2004).
Papke, R. T. et al. Searching for species in haloarchaea. Proc. Natl Acad. Sci. USA 104, 14092–14097 (2007).
Rodriguez-Valera, F. et al. Explaining microbial population genomics through phage predation. Nature Rev. Microbiol. 7, 828–836 (2009).
Baliga, N. S. et al. Coordinate regulation of energy transduction modules in Halobacterium sp. analyzed by a global systems approach. Proc. Natl Acad. Sci. USA 99, 14913–14918 (2002).
Facciotti, M. T. General transcription factor specified global gene regulation in archaea Proc. Natl Acad. Sci. USA 104, 4630–4635 (2007).
Koide, T. et al. Prevalence of transcription promoters within archaeal operons and coding sequences. Mol. Syst. Biol. 5, 285 (2009).
Bonneau, R. et al. A predictive model for transcriptional control of physiology in a free living cell. Cell 131, 1354–1365 (2007).
Bare, J. C., Koide, T., Reiss, D. J., Tenenbaum, D. & Baliga, N. S. Integration and visualization of systems biology data in context of the genome. BMC Bioinformatics 11, 382 (2010).
Rosenshine, I., Tchelet, R. & Mevarech, M. The mechanism of DNA transfer in the mating system of an archaebacterium. Science 245, 1387–1389 (1989).
Humbard, M. A. et al. Ubiquitin-like small archaeal modifier proteins (SAMPs) in Haloferax volcanii. Nature 463, 54–60 (2010).
Grogan, D. W. Exchange of genetic markers at extremely high temperatures in the archaeon Sulfolobus acidocaldarius. J. Bacteriol. 178, 3207–3211 (1996).
Bertani, G. & Baresi, L. Genetic transformation in the methanogen Methanococcus voltae PS. J. Bacteriol. 169, 2730–2738 (1987).
Sato, T., Fukui, T., Atomi, H. & Imanaka, T. Targeted gene disruption by homologous recombination in the hyperthermophilic archaeon Thermococcus kodakaraensis KOD1. J. Bacteriol. 185, 210–220 (2003).
Charlebois, R. L., Lam, W. L., Cline, S. W. & Doolittle, W. F. Characterization of pHV2 from Halobacterium volcanii and its use in demonstrating transformation of an archaebacterium. Proc. Natl Acad. Sci. USA 84, 8530–8534 (1987).
Metcalf, W. W., Zhang, J. K., Apolinario, E., Sowers, K. R. & Wolfe, R. S. A genetic system for Archaea of the genus Methanosarcina: liposome-mediated transformation and construction of shuttle vectors. Proc. Natl Acad. Sci. USA 94, 2626–2631 (1997). A striking example of the development of a novel, high-efficiency gene transfer system.
Schleper, C., Kubo, K. & Zillig, W. The particle SSV1 from the extremely thermophilic archaeon Sulfolobus is a virus: demonstration of infectivity and of transfection with viral DNA. Proc. Natl Acad. Sci. USA 89, 7645–7649 (1992).
Allers, T. & Mevarech, M. Archaeal genetics — the third way. Nature Rev. Genet. 6, 58–73 (2005). A good summary of the genetic systems that have been developed for archaea.
Sowers, K. & Anderson, K. in Archaea: Molecular and Cellular Biology (ed. Cavicchioli, R.) 463–477 (American Society for Microbiology Press, Washington DC, 2007).
Seghal, S. N., Kates, M. & Gibbons, N. E. Lipids of Halobacterium cutirubrum. Can. J. Biochem. Physiol. 40, 69–81 (1962).
Darland, G., Brock, T. D., Samsonoff, W. & Conti, S. F. A thermophilic, acidophilic mycoplasma isolated from a coal refuse pile. Science 170, 1416–1418 (1970). The discovery of an unusual archaeon before the knowledge that a third domain existed.
Langworthy, T. A., Smith, M. E. & Mayberry, W. R. A new class of lipopolysaccharide from Thermoplasma acidophilum. J. Bacteriol. 119, 106–116 (1974).
Brock, T. D., Brock, K. M., Belly, R. T. & Weiss, R. L. Sulfolobus: a new genus of sulphur oxidizing bacteria living at low pH and high temperature. Arch. Microbiol. 84, 54–68 (1972).
Langworthy, T. A., Smith, M. E. & Mayberry, W. R. Long-chain glycerol diether and polyol dialkyl glycerol triether lipids of Sulfolobus acidocaldarius. J. Bacteriol. 112, 1193–1200 (1972).
Stetter, K. O. History of discovery of the first hyperthermophiles. Extremophiles 10, 357–262 (2006). A brief, but informative, review describing how archaeal hyperthermophiles were discovered.
Stetter, K. O. Hyperthermophiles in the history of life. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 361, 1837–1842 (2006).
Waters, E. et al. The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism. Proc. Natl Acad. Sci. USA 100, 12984–12988 (2003).
Podar, M. et al. A genomic analysis of the archaeal system Ignicoccus hospitalis-Nanoarchaeum equitans. Genome Biol. 9, R158 (2008).
Küper, U., Meyer, C., Müller, V., Rachel, R. & Huber, H. Energized outer membrane and spatial separation of metabolic processes in the hyperthermophilic Archaeon Ignicoccus hospitalis. Proc. Natl Acad. Sci. USA 107, 3152–3126 (2010). Insight into the unusual relationship between an archaeal host and archaeal symbiont or parasite, with possible implications for the evolution of a eukaryote-like cell type.
van Ooij, C. A road to eukaryotes? Nature Rev. Microbiol. 8, 246–247 (2010).
Fuhrman, J. A., McCallum, K. & Davis, A. A. Novel major archaebacterial group from marine plankton. Nature 356, 148–149 (1992).
DeLong, E. F. Archaea in coastal marine environments. Proc. Natl Acad. Sci. USA 89, 5685–5689 (1992). Along with reference 132, this paper describes the new insight that archaea are important members of marine environments.
Karner, M. B., DeLong, E. F. & Karl, D. M. Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature 409, 507–510 (2001). The sheer abundance of archaea in the marine biosphere is described.
Cavicchioli, R. Cold adapted Archaea. Nature Rev. Microbiol. 4, 331–343 (2006). Insight into the diversity, functional roles and adaptive mechanisms of archaea in the cold biosphere.
Franzmann, P. D. et al. Halobacterium lacusprofundi sp. nov., a halophilic bacterium isolated from Deep Lake, Antarctica. Syst. Appl. Microbiol. 11, 20–27 (1988).
Franzmann, P. D., Stringer, N., Ludwig, W., Conway de Macario, E. & Rohde, M. A methanogenic archaeon from Ace Lake, Antarctica: Methanococcoides burtonii sp. nov. System. Appl. Microbiol. 15, 573–581 (1992).
Franzmann, P. D. et al. Methanogenium frigidum sp. nov., a psychrophilic, H2-using methanogen from Ace Lake, Antarctica. Int. J. Syst. Bacteriol. 47, 1068–1072 (1997).
Allen, M. et al. The genome sequence of the psychrophilic archaeon, Methanococcoides burtonii: the role of genome evolution in cold-adaptation. ISME J. 3, 1012–1035 (2009).
Takai, K. et al. Cell proliferation at 122 °C and isotopically heavy CH4 production by a hyperthermophilic methanogen under high-pressure cultivation. Proc. Natl Acad. Sci. USA 105, 10949–10954 (2008).
Schleper, C. et al. Genomic analysis reveals chromosomal variation in natural populations of the uncultured psychrophilic archaeon Cenarchaeum symbiosum. J. Bacteriol. 180, 5003–5009 (1998).
Schleper, C., Jurgens, G. & Jonuscheit, M. Genomic studies of uncultivated archaea. Nature Rev. Microbiol. 3, 479–488 (2005).
Könneke, M. et al. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437, 543–546 (2005). Revealing insight into an elusive phylogenetic and phenotypic type of archaea: chemolithoautrophic ammonia oxidizers.
Walker, C. B. et al. Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine Crenarchaea. Proc. Natl Acad. Sci. USA 107, 8818–8823 (2010).
Berg, I. A. et al. Autotrophic carbon fixation in archaea. Nature Rev. Microbiol. 8, 447–460 (2008).
Buckley, D. H., Graber, J. R. & Schmidt, T. M. Phylogenetic analysis of nonthermophilic members of the kingdom Crenarchaeota and their diversity and abundance in soils. Appl. Environ. Microbiol. 64, 4333–4339 (1998).
Leininger, S. et al. Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442, 806–809 (2006).
Erguder, T. H., Boon, N., Wittebolle, L., Marzorati, M. & Verstraete, W. Environmental factors shaping the ecological niches of ammonia-oxidizing archaea. FEMS Microbiol. Rev. 33, 855–869 (2009).
Hirata, A., Klein, B. J. & Murakami, K. S. The X-ray crystal structure of RNA polymerase from Archaea. Nature 451, 851–854 (2008). A striking example of how visually and functionally similar key components of the information processing systems are in the Archaea and Eukarya.
Cavicchioli, R., Siddiqui, K. S., Andrews, D. & Sowers, K. R. Low-temperature extremophiles and their applications. Curr. Opin. Biotech. 13, 253–261 (2002).
Schiraldi, C., Giuliano, M. & De Rosa, M. Perspectives on biotechnological applications of archaea. Archaea 1, 75–86 (2002).
Auernik, K. S., Cooper, C. R. & Kelly, R. M. Life in hot acid: pathway analyses in extremely thermoacidophilic archaea. Curr. Opin. Biotechnol. 19, 445–453 (2008).
Krishnan, L. & Sprott, G. D. Archaeosome adjuvants: immunological capabilities and mechanism(s) of action. Vaccine 26, 2043–2055 (2008).
Cavicchioli, R. Extremophiles and the search for extra-terrestrial life. Astrobiology 2, 281–292 (2002).
Tung, H. C., Bramall, N. E. & Price, P. B. Microbial origin of excess methane in glacial ice and implications for life on Mars. Proc. Natl Acad. Sci. USA 102, 18292–18296 (2005).
Oremland, R. S. & Voytek, M. A. Acetylene as fast food: implications for development of life on anoxic primordial Earth and in the outer solar system. Astrobiology 8, 45–58 (2008).
Oremland, R. S. NO connection with methane. Nature 464, 500–501 (2010).
Stetter, K. O. Hyperthermophilic procaryotes. FEMS Microbiol. Rev. 18, 149–158 (1996).
Koonin, E. V. & Martin, W. On the origin of genomes and cells within inorganic compartments. Trends Genet. 21, 647–654 (2005).
Brazelton, W. J. et al. Archaea and bacteria with surprising microdiversity show shifts in dominance over 1,000-year time scales in hydrothermal chimneys. Proc. Natl Acad. Sci. USA 107, 1612–1617 (2010).
Glansdorff, N., Xu, Y. & Labedan, B. The last universal common ancestor: emergence, constitution and genetic legacy of an elusive forerunner. Biol. Direct 3, 29 (2008).
(No authors listed). A sequence of changes. Nature Rev. Microbiol. 8, 85 (2010).
Delsuc, F., Brinkmann, H. & Philippe, H. Phylogenomics and the reconstruction of the tree of life. Nature Rev. Genet. 6, 361–375 (2005).
Doolittle, W. F. & Zhaxybayeva, O. On the origin of prokaryotic species. Genome Res. 19, 744–756 (2009).
Theobald, D. L. A formal test of the theory of universal common ancestry. Nature 465, 219–222 (2010).
Pace, N. R. Time for a change. Nature 441, 289 (2006). An important and axiomatic essay about learning from what we have learned.
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Acknowledgements
I dedicate this Review to Carl Woese, who pioneered, led and continues to inspire the field with his brilliance. I am indebted to C. Robertson, who generously constructed the phylogenetic tree, and M. DeMaere, who processed the RefSeq data. I also thank my many colleagues who informally and formally commented on the manuscript, in particular N. Pace, T. Kolesnikow, F. Lauro, H. Ertan and M. Dyall-Smith. This Review could not be exhaustive and I regret not being able to cite all of the relevant literature. Research in my laboratory is supported by the Australian Research Council.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- Domain
-
The highest level of taxonomic division; the three domains are the Archaea, Bacteria and Eukarya. In descending order, the other levels include: kingdom, phylum, class, order, family, genus and species.
- Extremophile
-
An organism that requires extreme environments for growth, such as extremes of temperature, salinity or pH, or a combination of these.
- Methanogen
-
An anaerobic organism that generates methane by the reduction of carbon dioxide, acetic acid, or various one-carbon compounds such as methylamines or methanol.
- Halophile
-
An organism that requires high concentrations of salt (typically greater than 1M NaCl) for growth.
- Thermoacidophile
-
An organism that requires high temperatures (typically greater than 60 °C) and a low pH (typically less than pH 3) for growth.
- Heterotroph
-
An organism that uses organic compounds as nutrients to produce energy for growth.
- Phototrophic
-
Pertaining to the growth of an organism: able to use sunlight to generate energy for growth.
- Hyperthermophile
-
An organism that requires extremely high temperatures (typically greater than 80 °C) for growth.
- Autotroph
-
An organism that can grow on carbon dioxide as a sole source of carbon.
- Small-subunit rRNA
-
The ribosome is the core biological machine of the translation apparatus and is essential for converting the genetic code described in DNA and mRNA into protein. Ribosomal RNA (rRNA) is the RNA component of the ribosome and forms two subunits, the small subunit (SSU) and the large subunit. SSU rRNA is highly conserved in all cellular forms of life and is commonly used for describing the phylogeny of organisms.
- Lateral gene transfer
-
Horizontal transfer of genes between unrelated species, as opposed to vertical inheritance within a species.
- Bootstrap value
-
A computationally derived measure of confidence about tree topology: the closer the bootstrap value is to 100, the more confidence we can have in the topology of the tree.
- Monophyletic
-
Pertaining to a natural taxonomic group or clade: consisting of individuals that share a common ancestor.
- Reverse methanogenesis
-
The methanogenesis pathway functioning in reverse to consume methane and produce cellular carbon and energy; this process leads to the anaerobic oxidation of methane.
- Chemolithoautotrophically
-
Pertaining to an organism: able to derive energy from a chemical reaction (chemotroph) using inorganic substrates as electron donors (lithotroph) and CO2 as a carbon source (autotroph).
- Ammonia-oxidizing members of the Crenarchaeota
-
An archaeon with the ability to grow chemolithoautotrophically with near-stoichiometric conversion of ammonium cations (NH4+) to nitrite ions (NO2−) using carbonic acid (H2CO3) and ammonium (NH4) as the sole sources of carbon and nitrogen, respectively.
Rights and permissions
About this article
Cite this article
Cavicchioli, R. Archaea — timeline of the third domain. Nat Rev Microbiol 9, 51–61 (2011). https://doi.org/10.1038/nrmicro2482
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2482
This article is cited by
-
Experimental warming leads to convergent succession of grassland archaeal community
Nature Climate Change (2023)
-
Community structure and activity potentials of archaeal communities in hadal sediments of the Mariana and Mussau trenches
Marine Life Science & Technology (2022)
-
Microbial Community Characterizing Vermiculations from Karst Caves and Its Role in Their Formation
Microbial Ecology (2021)
-
Transitions in microbial communities along two sediment cores collected from the landward walls of the New Britain trench
Marine Biology (2020)
-
Archaeal communities in the deep-sea sediments of the South China Sea revealed by Illumina high-throughput sequencing
Annals of Microbiology (2019)