Abstract
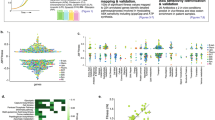
Large-scale, systems biology approaches now allow us to systematically map synergistic and antagonistic interactions between drugs. Consequently, drug antagonism is emerging as a powerful tool to study biological function and relatedness between cellular components as well as to uncover mechanisms of drug action. Furthermore, theoretical models and new experiments suggest that antagonistic interactions between antibiotics can counteract the evolution of drug resistance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hickman, M. & Cairns, J. The centenary of the one-gene one-enzyme hypothesis. Genetics 163, 839–841 (2003).
Loewe, S. Die quantitation probleme der pharmakologie. Ergeb. Physiol. 27, 47–187 (1928).
Bliss, C. I. The toxicity of poisons applied jointly. Ann. Appl. Biol. 26, 585–615 (1939).
Greco, W. R., Bravo, G. & Parsons, J. C. The search for synergy: a critical review from a response surface perspective. Pharmacol. Rev. 47, 331–385 (1995).
Frankel, W. N. & Schork, N. J. Who's afraid of epistasis? Nature Genet. 14, 371–373 (1996).
Phillips, P. C. The language of gene interaction. Genetics 149, 1167–1171 (1998).
Phillips, P. C., Otto, S. P. & Whitlock, M. C. in Epistasis and the Evolutionary Process (Oxford Univ. Press, New York, 2000).
Brodie, E. D. III. in Epistasis and the Evolutionary Process (Oxford Univ. Press, New York, 2000).
Mani, R., Onge, R. P. S., Hartman, J. L., Giaever, G. & Roth, F. P. Defining genetic interaction. Proc. Natl Acad. Sci. USA 105, 3461–3466 (2008).
Chait, R., Craney, A. & Kishony, R. Antibiotic interactions that select against resistance. Nature 446, 668–671 (2007).
Hegreness, M., Shoresh, N., Damian, D., Hartl, D. & Kishony, R. Accelerated evolution of resistance in multidrug environments. Proc. Natl Acad. Sci. USA 105, 13977–13981 (2008).
Michel, J.-B., Yeh, P., Chait, R., Moellering, R. C. & Kishony, R. Drug interactions modulate the potential for evolution of resistance. Proc. Natl Acad. Sci. USA 105, 14918–14923 (2008).
Tong, A. H. Y. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Tong, A. H. Y. et al. Global mapping of the yeast genetic interaction network. Science 303, 808–813 (2004).
Pan, X. W. et al. A DNA integrity network in the yeast Saccharomyces cerevisiae. Cell 124, 1069–1081 (2006).
Lehner, B., Crombie, C., Tischler, J., Fortunato, A. & Fraser, A. G. Systematic mapping of genetic interactions in Caenorhabditis elegans identifies common modifiers of diverse signaling pathways. Nature Genet. 38, 896–903 (2006).
Tischler, J., Lehner, B. & Fraser, A. G. Evolutionary plasticity of genetic interaction networks. Nature Genet. 40, 390–391 (2008).
Ye, P. et al. Gene function prediction from congruent synthetic lethal interactions in yeast. Mol. Syst. Biol. 1, 2005.0026 (2005).
Ooi, S. L. et al. Global synthetic-lethality analysis and yeast functional profiling. Trends Genet. 22, 56–63 (2006).
Meluh, P. B. et al. Analysis of genetic interactions on a genome-wide scale in budding yeast: diploid-based synthetic lethality analysis by microarray. Methods Mol. Biol. 416, 221–247 (2008).
Sanjuan, R., Moya, A. & Elena, S. F. The contribution of epistasis to the architecture of fitness in an RNA virus. Proc. Natl Acad. Sci. USA 101, 15376–15379 (2004).
Drees, B. L. et al. Derivation of genetic interaction networks from quantitative phenotype data. Genome Biol. 6, R38 (2005).
Schuldiner, M. et al. Exploration of the function and organization of the yeast early secretory pathway through an epistatic miniarray profile. Cell 123, 507–519 (2005).
Collins, S. R., Schuldiner, M., Krogan, N. J. & Weissman, J. S. A strategy for extracting and analyzing large-scale quantitative epistatic interaction data. Genome Biol. 7, R63 (2006).
St Onge, R. P. et al. Systematic pathway analysis using high-resolution fitness profiling of combinatorial gene deletions. Nature Genet. 39, 199–206 (2007).
Jasnos, L. & Korona, R. Epistatic buffering of fitness loss in yeast double deletion strains. Nature Genet. 39, 550–554 (2007).
Typas, A. et al. High-throughput, quantitative analyses of genetic interactions in E. coli. Nature Methods 5, 781–787 (2008).
Roguev, A. et al. Conservation and rewiring of functional modules revealed by an epistasis map in fission yeast. Science 322, 405–410 (2008).
Collins, S. R. et al. Functional dissection of protein complexes involved in yeast chromosome biology using a genetic interaction map. Nature 446, 806–810 (2007).
Roguev, A., Wiren, M., Weissman, J. S. & Krogan, N. J. High-throughput genetic interaction mapping in the fission yeast Schizosaccharomyces pombe. Nature Methods 4, 861–866 (2007).
DeLuna, A. et al. Exposing the fitness contribution of duplicated genes. Nature Genet. 40, 676–681 (2008).
Breslow, D. K. et al. A comprehensive strategy enabling high-resolution functional analysis of the yeast genome. Nature Methods 5, 711–718 (2008).
Segre, D., DeLuna, A., Church, G. M. & Kishony, R. Modular epistasis in yeast metabolism. Nature Genet. 37, 77–83 (2005).
Bandyopadhyay, S., Kelley, R., Krogan, N. J. & Ideker, T. Functional maps of protein complexes from quantitative genetic interaction data. PloS Comput. Biol. 4, e1000065 (2008).
Yeh, P., Tschumi, A. I. & Kishony, R. Functional classification of drugs by properties of their pairwise interactions. Nature Genet. 38, 489–494 (2006).
Lehar, J. et al. Chemical combination effects predict connectivity in biological systems. Mol. Syst. Biol. 3, 80 (2007).
Yeh, P. & Kishony, R. Networks from drug–drug surfaces. Mol. Syst. Biol. 3, 85 (2007).
Pillai, S. K., Moellering, R. C. & Eliopoulos, G. M. in Antibiotics in Laboratory Medicine (ed. Lorian, V.) 365–440 (Lippincott Williams & Wilkins, Philadelphia, 2005).
Fraser, T. R. The antagonism between the actions of active substances. Br. Med. J. 2, 485–487 (1872).
Eagle, H. & Musselman, A. D. The rate of bactericidal action of penicillin in vitro as a function of its concentration, and its paradoxically reduced activity at high concentrations against certain organisms. J. Exp. Med. 88, 99–131 (1948).
Smith, J. T. The mode of action of 4-quinolones and possible mechanisms of resistance. J. Antimicrob. Chemother. 18, 21–29 (1986).
Lewin, C. S., Morrissey, I. & Smith, J. T. The mode of action of quinolones: the paradox in activity of low and high concentrations and activity in the anaerobic environment. Eur. J. Clin. Microbiol. Infect. Dis. 10, 240–248 (1991).
Baquero, F. Resistance to quinolones in gram-negative micororganisms: mechanisms and prevention. Eur. Urol. 17, 3–12 (1990).
Baquero, F. & Negri, M. C. Strategies to minimize the development of antibiotic resistance. J. Chemother. 9, 29–37 (1997).
Dong, Y. Z., Zhao, X. L., Domagala, J. & Drlica, K. Effect of fluoroquinolone concentration on selection of resistant mutants of Mycobacterium bovis BCG and Staphylococcus aureus. Antimicrob. Agents. Chemother. 43, 1756–1758 (1999).
Drlica, K. The mutant selection window and antimicrobial resistance. J. Antimicrob. Chemother. 52, 11–17 (2003).
Dong, Y. Z., Zhao, X. L., Kreiswirth, B. N. & Drlica, K. Mutant prevention concentration as a measure of antibiotic potency: studies with clinical isolates of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 44, 2581–2584 (2000).
Blondeau, J. M., Zhao, X. L., Hansen, G. & Drlica, K. Mutant prevention concentrations of fluoroquinolones for clinical isolates of Streptococcus pneumoniae. Antimicrob. Agents Chemother. 45, 433–438 (2001).
Zhao, X. L. & Drlica, K. Restricting the selection of antibiotic-resistant mutant bacteria: measurement and potential use of the mutant selection window. J. Infect. Dis. 185, 561–565 (2002).
Randall, L. P., Cooles, S. W., Piddock, L. J. V. & Woodward, M. J. Mutant prevention concentrations of ciprofloxacin and enrofloxacin for Salmonella enterica. J. Antimicrob. Chemother. 54, 688–691 (2004).
Metzler, K. et al. Comparison of minimal inhibitory and mutant prevention drug concentrations of 4 fluoroquinolones against clinical isolates of methicillin-susceptible and -resistant Staphylococcus aureus. Int. J. Antimicrob. Agents 24, 161–167 (2004).
Linde, H. J. & Lehn, N. Mutant prevention concentration of nalidixic acid, ciprofloxacin, clinafloxacin, levofloxacin, norfloxacin, ofloxacin, sparfloxacin or trovafloxacin for Escherichia coli under different growth conditions. J. Antimicrob. Chemother. 53, 252–257 (2004).
Li, X. Y., Mariano, N., Rahal, J. J., Urban, C. M. & Drlica, K. Quinolone-resistant Haemophilus influenzae: determination of mutant selection window for ciprofloxacin, garenoxacin, levofloxacin, and moxifloxacin. Antimicrob. Agents Chemother. 48, 4460–4462 (2004).
Marcusson, L. L., Olofsson, S. K., Lindgren, P. K., Cars, O. & Hughes, D. Mutant prevention concentrations of ciprofloxacin for urinary tract infection isolates of Escherichia coli. J. Antimicrob. Chemother. 55, 938–943 (2005).
Rodriguez-Martinez, J. M. et al. Mutant prevention concentrations of fluoroquinolones for Enterobacteriaceae expressing the plasmid-carried quinolone resistance determinant qnrA1. Antimicrob. Agents Chemother. 51, 2236–2239 (2007).
Firsov, A. A. et al. In vitro pharmacodynamic evaluation of the mutant selection window hypothesis using four fluoroquinolones against Staphylococcus aureus. Antimicrob. Agents Chemother. 47, 1604–1613 (2003).
Firsov, A. A., Lubenko, I. Y., Smirnova, M. V., Strukova, E. N. & Zinner, S. H. Enrichment of fluoroquinolone-resistant Staphylococcus aureus: oscillating ciprofloxacin concentrations simulated at the upper and lower portions of the mutant selection window. Antimicrob. Agents Chemother. 52, 1924–1928 (2008).
Zhanel, G. G., Mayer, M., Laing, N. & Adam, H. J. Mutant prevention concentrations of levofloxacin alone and in combination with azithromycin, ceftazidime, colistin (polymyxin E), meropenem, piperacillin-tazobactam, and tobramycin against Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 50, 2228–2230 (2006).
Deeks, S. G. Treatment of anti retroviral-drug-resistant HIV-1 infection. Lancet 362, 2002–2011 (2003).
Nosten, F. & Brasseur, P. Combination therapy for malaria: the way forward? Drugs 62, 1315–1329 (2002).
Klein, M. & Schorr, S. E. The role of bacterial resistance in antibiotic synergism and antagonism. J. Bacteriol. 65, 454–465 (1953).
Jawetz, E. Infectious diseases: problems of antimicrobial therapy. Ann. Rev. Med. 5, 1–26 (1954).
Lipsitch, M. & Levin, B. R. The population dynamics of antimicrobial chemotherapy. Antimicrob. Agents Chemother. 41, 363–373 (1997).
Lepper, M. H. & Dowling, H. F. Treatment of pneumococcic meningitis with penicillin compared with penicillin plus aureomycin; studies including observations on an apparent antagonism between penicillin and aureomycin. AMA Arch. Intern. Med. 88, 489–494 (1951).
Kishony, R. & Leibler, S. Environmental stresses can alleviate the average deleterious effect of mutations. J. Biol. 2, 14 (2003).
Blagosklonny, M. V. Drug-resistance enables selective killing of resistant leukemia cells: exploiting of drug resistance instead of reversal. Leukemia 13, 2031–2035 (1999).
Acknowledgements
We thank A. DeLuna, J.-B. Michel, R. Chait, N. Shoresh and T. Bollenbach for helpful discussion and comments on the manuscript. R. Chait contributed many of the figures for Box 1. This work was supported in part by a National Institutes of Health postdoctoral fellowship to P.J.Y., by National Science Foundation and National Defense Science and Engineering graduate fellowships to A.P.A. and by National Institutes of Health Grant R01GM081617 to R.K.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Yeh, P., Hegreness, M., Aiden, A. et al. Drug interactions and the evolution of antibiotic resistance. Nat Rev Microbiol 7, 460–466 (2009). https://doi.org/10.1038/nrmicro2133
Issue Date:
DOI: https://doi.org/10.1038/nrmicro2133
This article is cited by
-
Modulation of antibiotic effects on microbial communities by resource competition
Nature Communications (2023)
-
Anwendungsbereiche von künstlicher Intelligenz im Kontext von One Health mit Fokus auf antimikrobielle Resistenzen
Bundesgesundheitsblatt - Gesundheitsforschung - Gesundheitsschutz (2023)
-
An efflux-susceptible antibiotic-adjuvant with systemic efficacy against mouse infections
Scientific Reports (2022)
-
Antibiotic combinations reduce Staphylococcus aureus clearance
Nature (2022)
-
Horizontal gene transfer enables programmable gene stability in synthetic microbiota
Nature Chemical Biology (2022)