Key Points
-
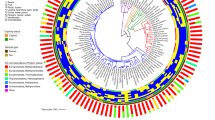
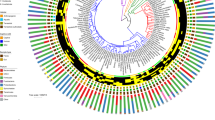
Comparative analyses between microbial communities in the human gut versus diverse other environments can help elucidate the environmental and evolutionary parameters that shape our intestinal microbiota. Here we use published 16S ribosomal RNA gene sequences to compare the bacterial assemblages that are associated with humans, other mammals, other metazoa and free-living microbial communities that span a range of environmental conditions.
-
Gut microorganisms and their animal hosts have co-evolved, and exploitation of a new diet niche is a powerful driver of the co-evolution of hosts and gut bacteria.
-
Gut microbial communities are distinct from those of other characterized habitats in the biosphere.
-
The vast dichotomy between vertebrate gut and free-living communities is evident from the bacterial phyla that comprise them, which indicates that strong selective forces differentiated gut-dwelling bacteria regardless of their lineage.
-
Most extant mammals are herbivores. The microbial 'solution' to herbivory in mammals with either a simple gut, hindgut or foregut seems to be similar, regardless of host phylogeny: that is, promiscuous gut microorganisms seem to have made it possible for 'unrelated' mammals with similar gut structures to assemble similar microbial communities to digest their polysaccharide-rich diets. This 'bottom-up' perspective of evolution implies that gut microorganisms are unindicted co-conspirators in the spectacular success of mammals.
-
Although few humans have been sampled to date, we seem to be typical omnivores that can be placed together with our omnivorous primate relatives. The recently initiated, international Human Microbiome Project should strive to include a wide representation of humans, as well as other mammals and environmental samples; comparative analyses of microbiotas and their microbiomes are a powerful and timely way to explore the evolutionary history of the biosphere.
Abstract
In this Analysis we use published 16S ribosomal RNA gene sequences to compare the bacterial assemblages that are associated with humans and other mammals, metazoa and free-living microbial communities that span a range of environments. The composition of the vertebrate gut microbiota is influenced by diet, host morphology and phylogeny, and in this respect the human gut bacterial community is typical of an omnivorous primate. However, the vertebrate gut microbiota is different from free-living communities that are not associated with animal body habitats. We propose that the recently initiated international Human Microbiome Project should strive to include a broad representation of humans, as well as other mammalian and environmental samples, as comparative analyses of microbiotas and their microbiomes are a powerful way to explore the evolutionary history of the biosphere.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Woese, C. R. On the evolution of cells. Proc. Natl Acad. Sci. USA 99, 8742–8747 (2002).
Allwood, A. C., Walter, M. R., Kamber, B. S., Marshall, C. P. & Burch, I. W. Stromatolite reef from the Early Archaean era of Australia. Nature 441, 714–718 (2006).
Martiny, J. B. et al. Microbial biogeography: putting microorganisms on the map. Nature Rev. Microbiol. 4, 102–112 (2006).
Lozupone, C. A., Hamady, M., Kelley, S. T. & Knight, R. Quantitative and qualitative b diversity measures lead to different insights into factors that structure microbial communities. Appl. Environ. Microbiol. 73, 1576–1585 (2007).
Lozupone, C. A. & Knight, R. Global patterns in bacterial diversity. Proc. Natl Acad. Sci. USA 104, 11436–11440 (2007). Combined data from hundreds of bacterial communities to show that phylogenetic tree-based measures of diversity can reveal large-scale trends that influence these communities.
Desnues, C. et al. Biodiversity and biogeography of phages in modern stromatolites and thrombolites. Nature 452, 340–343 (2008).
Butterfield, N. J., Knoll, A. H. & Swett, K. A bangiophyte red alga from the Proterozoic of arctic Canada. Science 250, 104–107 (1990).
Backhed, F., Ley, R. E., Sonnenburg, J. L., Peterson, D. A. & Gordon, J. I. Host–bacterial mutualism in the human intestine. Science 307, 1915–1920 (2005).
Robosky, L. C. et al. Metabonomic identification of two distinct phenotypes in Sprague–Dawley (Crl:CD(SD)) rats. Toxicol. Sci. 87, 277–284 (2005).
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006). The gut microbiomes of genetically obese mice were shown to have substantially different functional categories of genes compared with those of wild-type litter mates. This study revealed how comparative metagenomics can link host physiological states to the microbiome, as well as the power of testing predictions from such comparative studies with direct experimental tests that involve microbiota transplants into germ-free animals.
Rohde, C. M. et al. Metabonomic evaluation of Schaedler altered microflora rats. Chem. Res. Toxicol. 10, 1388–1392 (2007).
Rawls, J. F., Samuel, B. S. & Gordon, J. I. Gnotobiotic zebrafish reveal evolutionarily conserved responses to the gut microbiota. Proc. Natl Acad. Sci. USA 101, 4596–4601 (2004).
Turnbaugh, P. J. et al. The human microbiome project. Nature 449, 804–810 (2007).
Teaford, M. F. & Ungar, P. S. Diet and the evolution of the earliest human ancestors. Proc. Natl Acad. Sci. USA 97, 13506–13511 (2000).
O'Connell, J. F., Hawkes, K. & Blurton Jones, N. G. Grandmothering and the evolution of Homo erectus. J. Hum. Evol. 36, 461–485 (1999).
Ungar, P. S. Evolution of the Hominid Diet: The Known, the Unknown, and the Unknowable 39–55 (Oxford Univ. Press, 2006).
Ungar, P. S., Grine, F. E. & Teaford, M. F. Diet in early Homo: a review of the evidence and a new model of adaptive versatility. Annu. Rev. Anthropol. 35, 209–228 (2006).
Yeakel, J. D., Bennett, N. C., Koch, P. L. & Dominy, N. J. The isotopic ecology of African mole rats informs hypotheses on the evolution of human diet. Proc. R. Soc. Lond. B 274, 1723–1730 (2007).
Aiello, L. C. & Wheeler, P. The expensive-tissue hypothesis: the brain and the digestive system in human and primate evolution. Curr. Anthropol. 36, 199–221 (1995). The expensive-tissue hypothesis advanced in this classic paper proposes that the cost of a large brain is offset in humans by a reduction in gut size and that the evolution of a large brain must have been accompanied by a shift to higher-energy foods.
Wood, B. & Brooks, A. Human evolution: we are what we ate. Nature 400, 219–220 (1999).
Kehrer-Sawatzki, H. & Cooper, D. N. Understanding the recent evolution of the human genome: insights from human–chimpanzee genome comparisons. Hum. Mutat. 28, 99–130 (2007).
Samuelson, L. C., Wiebauer, K., Snow, C. M. & Meisler, M. H. Retroviral and pseudogene insertion sites reveal the lineage of human salivary and pancreatic amylase genes from a single gene during primate evolution. Mol. Cell. Biol. 10, 2513–2520 (1990).
Perry, G. H. et al. Diet and the evolution of human amylase gene copy number variation. Nature Genet. 39, 1256–1260 (2007).
Beja-Pereira, A. et al. Gene-culture co-evolution between cattle milk protein genes and human lactase genes. Nature Genet. 35, 311–313 (2003). Showed that genetic changes in both humans and cattle are associated with agriculture. This type of study emphasizes how changes in human culture, technology and cookery have shaped our microbial ecology and microbiomes.
Hollox, E. J. et al. Lactase haplotype diversity in the Old World. Am. J. Hum. Genet. 68, 160–172 (2001).
Falush, D. et al. Traces of human migrations in Helicobacter pylori populations. Science 299, 1582–1585 (2003).
Holmes, E. et al. Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 453, 396–400 (2008).
Li, M. et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proc. Natl Acad. Sci. USA 105, 2117–2123 (2008). This important paper provides a crucial link between human physiology and human microbial ecology by showing that the abundance of certain metabolites correlates with the abundance of particular kinds of gut bacteria, rather than with ancestry.
Suau, A. et al. Direct analysis of genes encoding 16S rRNA from complex communities reveals many novel molecular species within the human gut. Appl. Environ. Microbiol. 65, 4799–4807 (1999).
Eckburg, P. B. et al. Diversity of the human intestinal microbial flora. Science 308, 1635–1638 (2005).
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006).
Ley, R. et al. Evolution of mammals and their gut microbes. Science 320, 1647–1651 (2008). Provides a 16S rRNA-based survey of the gut bacterial communities in 60 species of mammals. The results show that diet and host phylogeny influence which bacteria live in which hosts.
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Hladik, C. M. & Pasquet, P. The human adaptations to meat eating: a reappraisal. Hum. Evol. 17, 199–206 (2002).
Collinson, M. E. et al. Fossil evidence of interactions between plants and plant-eating mammals. Philos. Trans. R. Soc. Lond. B 333, 197–208 (1991).
Mackie, R. I. Mutualistic fermentative digestion in the gastrointestinal tract: diversity and evolution 1. Integr. Comp. Biol. 42, 319–326 (2002). This helpful review outlines the main carbohydrate-fermentation pathways in the rumen and discusses how digestive-tract anatomy and diet affect rumen function, as mediated by symbiotic microorganisms.
Russell, J. B. & Rychlik, J. L. Factors that alter rumen microbial ecology. Science 292, 1119–1122 (2001).
Stevens, C. & Hume, I. Comparative Physiology of the Vertebrate Digestive System (Cambridge Univ. Press, New York, 2004).
Hackstein, J. H. P. & Stumm, C. K. Methane production in terrestrial arthropods. Proc. Natl Acad. Sci. USA 91, 5441–5445 (1994).
Hackstein, J. H. P. & van Alen, T. A. Fecal methanogens and vertebrate evolution. Evolution 50, 559–572 (1996). Investigated methanogenesis in a large and phylogenetically diverse assemblage of animals. Methanogenesis was absent from entire clades regardless of diet, which supports the hypothesis that hosts harbour methanogens owing to genetic factors.
Dierenfeld, E. S., Hintz, H. F., Robertson, J. B., Van Soest, P. J. & Oftedal, O. T. Utilization of bamboo by the giant panda. J. Nutr. 112, 636–641 (1982).
Hengstmann, U., Chin, K. J., Janssen, P. H. & Liesack, W. Comparative phylogenetic assignment of environmental sequences of genes encoding 16S rRNA and numerically abundant culturable bacteria from an anoxic rice paddy soil. Appl. Environ. Microbiol. 65, 5050–5058 (1999).
Egert, M., Wagner, B., Lemke, T., Brune, A. & Friedrich, M. W. Microbial community structure in midgut and hindgut of the humus-feeding larva of Pachnoda ephippiata (Coleoptera: Scarabaeidae). Appl. Environ. Microbiol. 69, 6659–6668 (2003).
Egert, M., Stingl, U., Bruun, L. D., Pommerenke, B., Brune, A. & Friedrich, M. W. Structure and topology of microbial communities in the major gut compartments of Melolontha melolontha larvae (Coleoptera: Scarabaeidae). Appl. Environ. Microbiol. 71, 4556–4566 (2005).
Schloss, P. D., Delaibera, I., Handelsman, J. & Raffa, K. F. Bacteria associated with the guts of two wood-boring beetles: Anoplophora glabripennis and Saperda vestita (Cerambycidae). Environ. Entomol. 35, 625–629 (2006).
Klappenbach, J. A., Dunbar, J. M. & Schmidt, T. M. rRNA operon copy number reflects ecological strategies of bacteria. Appl. Environ. Microbiol. 66, 1328–1333 (2000). Correlated the ability of bacteria to grow fast in response to a sudden increase in available energy sources, 'weediness' or r selection with a high copy number of rRNA operons in the genome.
Acinas, S. G., Marcelino, L. A., Klepac-Ceraj, V. & Polz, M. F. Divergence and redundancy of 16S rRNA sequences in genomes with multiple rrn operons. J. Bacteriol. 186, 2629–2635 (2004).
Tanner, M., Goebel, B. M., Dojka, M. A. & Pace, N. R. Specific rDNA sequences from diverse environmental settings correlate with experimental contaminants. Appl. Environ. Microbiol. 8, 3110–3113 (1998).
Dethlefsen, L., Eckburg, P. B., Bik, E. M. & Relman, D. A. Assembly of the human intestinal microbiota. Trends Ecol. Evol. 21, 517–523 (2006).
Ley, R. E., Peterson, D. A. & Gordon, J. I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Dethlefsen, L., McFall-Ngai, M. & Relman, D. A. An ecological and evolutionary perspective on human–microbe mutualism and disease. Nature 449, 811–818 (2007).
Sloan, W. T. et al. Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ. Microbiol. 8, 732–740 (2006).
Woodcock, S. et al. Neutral assembly of bacterial communities. FEMS Microbiol. Ecol. 62, 171–180 (2007). This theoretical paper contributed to our understanding of what would be expected by chance if bacterial communities are assembled according to neutral processes (those without selection) and showed that many microbial communities could fit this model.
Hubbell, S. P. in The Unified Neutral Theory of Biodiversity and Biogeography (Princeton Univ Press, 2001).
Hubbell, S. P. Neutral theory and the evolution of ecological equivalence. Ecology 87, 1387–1398 (2006).
McFall-Ngai, M. Adaptive immunity: care for the community. Nature 445, 153 (2007).
Peterson, D. A., McNulty, N. P., Guruge, J. L. & Gordon, J. I. IgA response to symbiotic bacteria as a mediator of gut homeostasis. Cell Host Microbe 2, 328–339 (2007).
Fagarasan, S. et al. Critical roles of activation-induced cytidine deaminase in the homeostasis of gut flora. Science 298, 1424–1427 (2002).
Suzuki, K. et al. Aberrant expansion of segmented filamentous bacteria in IgA-deficient gut. Proc. Natl Acad. Sci. USA 101, 1981–1986 (2004).
Lanyon, C. V. et al. Murine scent mark microbial communities are genetically determined. FEMS Microbiol. Ecol. 59, 576–583 (2007). Detected a genetic influence of the host on microbial communities that alter the smell of urine, a mechanism that provides the host with a range of functions, including territory marking and signalling to potential mates. The study also described a specific effect of the MHC.
Lombardo, M. P. Access to mutualistic endosymbiotic microbes: an underappreciated benefit of group living. Behav. Ecol. Sociobiol. 1–19 (2008). This study integrates mammalian behaviour with microbial community ecology to propose that communal living allows beneficial microorganisms to be shared.
Ley, R. E. et al. Obesity alters gut microbial ecology. Proc. Natl Acad. Sci. USA 102, 11070–11075 (2005).
Lucas, F. S. & Heeb, P. Environmental factors shape cloacal bacterial assemblages in great tit Parus major and blue tit P. caeruleus nestlings. J. Avian Biol. 36, 510–516 (2005).
Zoetendal, E. G. et al. The host genotype affects the bacterial community in the human gastrointestinal tract. Microb. Ecol. Health Dis. 13, 129–134 (2001).
Stewart, J. A., Chadwick, V. S. & Murray, A. Investigations into the influence of host genetics on the predominant eubacteria in the faecal microflora of children. J. Med. Microbiol. 54, 1239–1242 (2005).
Palmer, C., Bik, E. M., Digiulio, D. B., Relman, D. A. & Brown, P. O. Development of the human infant intestinal microbiota. PLoS Biol. 5, e177 (2007). Culture-independent methods were used on samples from infants with different mothers to show that establishment of gut microbial communities during the postnatal period follows a highly varied course. Dizygotic twins had the most similar pattern of community assembly, suggesting that early environmental exposure is important.
Perez, P. F. et al. Bacterial imprinting of the neonatal immune system: lessons from maternal cells? Pediatrics 119, e724–e732 (2007).
Frankenfeld, C. L. et al. Familial correlations, segregation analysis, and nongenetic correlates of soy isoflavone-metabolizing phenotypes. Exp. Biol. Med. 229, 902–913 (2004).
Moran, N. A. Symbiosis. Curr. Biol. 16, 866–871 (2006).
Lederberg, J. Infectious history. Science 288, 287–293 (2000).
Nicholson, J. K., Holmes, E. & Wilson, I. D. Gut microorganisms, mammalian metabolism and personalized health care. Nature Rev. Microbiol. 3, 431–438 (2005).
Borneman, J. in Microbial Diversity and Bioprospecting (ed. Bull, A. T.) 421–428 (ASM, Virginia, 2004).
Allison, S. D. & Martiny, J. B. H. Resistance, resilience and redundancy in microbial communities. Proc. Natl Acad. Sci. USA 105, 11512–11519 (2008).
Fuerst, J. A. & Hugenholtz, P. Microorganisms should be high on DNA preservation list. Science 290, 1503 (2000).
Turrini, A., Avio, L., Bedini, S. & Giovannetti, M. In situ collection of endangered arbuscular mychorrhizal fungi in a Mediterranean UNESCO biosphere reserve. Biodivers. Conserv. 17, 643–657 (2008).
Staley, J. T. Biodiversity: are microbial species threatened? Curr. Opin. Biotech. 8, 340–345 (1997).
Ceballos, G., Ehrlich, P. R., Soberon, J., Salazar, I. & Fay, J. P. Global mammal conservation: what must we manage? Science 309, 603–607 (2005).
Blaser, M. J. Who are we? Indigenous microbes and the ecology of human diseases. EMBO Rep. 7, 956–960 (2006).
Pirages, D. C. Nature, disease, and globalization: an evolutionary perspective. Int. Stud. Rev. 9, 616–628 (2007).
Beard, A. S. & Blaser, M. J. The ecology of height: the effect of microbial transmission on human height. Perspect. Biol. Med. 45, 475–498 (2002). This intriguing integration of human historical, demographical and physiological data was used to propose a connection between the microbiota of a host and host height.
Knight, R. PyCogent: a toolkit for making sense from sequence. Genome Biol. 8, R171 (2007).
DeSantis, T. Z. Jr et al. NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Res. 34, 394–399 (2006).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004).
Widmann, J., Hamady, M. & Knight, R. DivergentSet, a tool for picking non-redundant sequences from large sequence collections. Mol. Cell Proteomics 5, 1520–1532 (2006).
Acknowledgements
Work in the authors' laboratories is supported by grants from the National Institutes of Health (NIH), the National Science Foundation (NSF), the W.M. Keck Foundation and the Crohn's and Colitis Foundation of America. M.H. is supported by training grants from the NIH (grant number T32GM065103) and NSF (East Asia and Pacific Summer Institutes (EAPSI) fellowship number OISE0812861).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Related links
Related links
DATABASES
Entrez Genome Project
FURTHER INFORMATION
SUPPLEMENTARY INFORMATION
See online article
Glossary
- Microbiota
-
The complete set of microbial lineages that live in a particular environment.
- Co-evolution
-
A process in which the evolution of one taxon or trait reciprocally influences the evolution of another.
- Co-metabolite
-
A metabolite that is synthesized or degraded by both the host and one or more microbial species.
- Microbiome
-
The complete collection of genes in the genomes of microorganisms that live in a particular environment (that is, the set of genes contained in a microbiota).
- UniFrac
-
A phylogenetic analysis technique that measures the distance between two community samples in terms of the amount of sequence divergence on a phylogenetic tree that is unique to each of the samples. In practice, a 'master' phylogenetic tree is constructed using all 16S ribosomal RNA sequences from all biological samples characterized in a study. Pairwise comparisons of samples (communities) are then performed with similarity defined (UniFrac metric) based on the degree of branch length that they share on the tree.
- Operational taxonomic unit
-
(OTU). A group of organisms that is defined by its sequence similarity (for Bacteria and Archaea, typically using their 16S ribosomal RNA (rRNA) genes). For example, OTUs at the 97% level (all 16S rRNA gene sequences are at least 97% identical) are often considered to define a species.
- OTU-based analysis
-
An analysis that is based on the counts of each type of OTU in each sample. Unlike phylogenetic analyses, in this approach, all OTUs are considered to be equal, independent units. For example, two OTUs that are closely related to one another will be treated the same as two OTUs that are distantly related.
- Horizontal gene transfer
-
A process by which a gene moves between two genomes rather than being vertically transmitted from parent to offspring.
- Principal coordinate analysis
-
An analysis that is based on a matrix of distances between samples, and detects a few dimensions to explain as much of the variation in the samples as possible using linear algebra techniques.
- Co-diversification
-
A process in which two lineages speciate in concert with one another; for example, when pathogens or commensals speciate into specialist lineages at the same time as their host lineage speciates.
- Metagenomics
-
Culture-independent analyses of the composition and dynamic operations of microbial communities. This includes community characterization at the level of DNA (microbiome), RNA (metatranscriptome), proteins (metaproteome) and metabolic networks (metabolome).
Rights and permissions
About this article
Cite this article
Ley, R., Lozupone, C., Hamady, M. et al. Worlds within worlds: evolution of the vertebrate gut microbiota. Nat Rev Microbiol 6, 776–788 (2008). https://doi.org/10.1038/nrmicro1978
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1978
This article is cited by
-
Comparative Analysis of Gut Bacterial Communities in Fish and Shellfish of Great Himalayan River System
Indian Journal of Microbiology (2024)
-
Host species and habitat shape fish-associated bacterial communities: phylosymbiosis between fish and their microbiome
Microbiome (2023)
-
Realising respiratory microbiomic meta-analyses: time for a standardised framework
Microbiome (2023)
-
Analysis of gut microbiota in chinese donkey in different regions using metagenomic sequencing
BMC Genomics (2023)
-
Host individual and gut location are more important in gut microbiota community composition than temporal variation in the marine herbivorous fish Kyphosus sydneyanus
BMC Microbiology (2023)