Key Points
-
Melioidosis is caused by the aerobic, Gram-negative soil-dwelling bacillus Burkholderia pseudomallei and is an important cause of severe sepsis in Southeast Asia and Northern Australia. The high associated mortality rate, wide availability in the environment in endemic areas, intrinsic resistance to many antibiotics and the potential for aerosol spread has made this organism a potential bioterror agent.
-
The genome of B. pseudomallei consists of a large chromosome (4.07 Mb) carrying genes mainly associated with cell growth and metabolism, and a smaller chromosome (3.17 Mb) which has a greater proportion of genes encoding accessory functions such as adaptation and survival in different environments. Approximately 6% of the genome is made up of genomic islands that have probably been acquired by horizontal gene transfer.
-
Factors associated with disease acquisition in endemic regions include adverse weather conditions, the route and size of the inoculum and the integrity of the host immune system. The geographical incidence of B. pseudomallei and typical clinical features of the disease are reviewed.
-
No single B. pseudomallei determinant has been shown to have a role in virulence during human disease, persistence or latency. However, putative virulence factors include quorum sensing, a type III secretion system, capsular polysaccharide and, with less conclusive evidence, lipopolysaccharide and flagella.
-
B. pseudomallei is an intracellular pathogen that multiplies within macrophages. Recent data have shed light on the mechanisms that this bacterium uses to adapt to, and exploit, the intracellular environment. IFN-γ and TNF-α have an important role in early resistance against B. pseudomallei infection. Although more is becoming known about the pathogenesis of this bacterium in disease, the host–pathogen interactions are still ill-defined.
-
Potential new therapies that warrant further clinical evaluation include granulocyte colony-stimulating factor and CpG (bacterial DNA). Development of an effective human melioidosis vaccine is a research priority.
Abstract
Burkholderia pseudomallei is a potential bioterror agent and the causative agent of melioidosis, a severe disease that is endemic in areas of Southeast Asia and Northern Australia. Infection is often associated with bacterial dissemination to distant sites, and there are many possible disease manifestations, with melioidosis septic shock being the most severe. Eradication of the organism following infection is difficult, with a slow fever-clearance time, the need for prolonged antibiotic therapy and a high rate of relapse if therapy is not completed. Mortality from melioidosis septic shock remains high despite appropriate antimicrobial therapy. Prevention of disease and a reduction in mortality and the rate of relapse are priority areas for future research efforts. Studying how the disease is acquired and the host–pathogen interactions involved will underpin these efforts; this review presents an overview of current knowledge in these areas, highlighting key topics for evaluation.
Similar content being viewed by others
Main
Melioidosis is a serious disease caused by the aerobic, Gram-negative soil-dwelling bacillus Burkholderia pseudomallei and is most common in Southeast Asia and Northern Australia. Melioidosis is responsible for 20% of all community-acquired septicaemias and 40% of sepsis-related mortality in northeast Thailand. Reported cases are likely to represent 'the tip of the iceberg'1,2, as confirmation of disease depends on bacterial isolation, a technique that is not available in many of the affected areas. Melioidosis often affects individuals with one or more pre-existing conditions associated with an altered immune response, the most common being diabetes mellitus (50% of cases). The most severe clinical manifestation is melioidosis septic shock, which is often associated with pneumonia and bacterial dissemination to distant sites (Fig. 1).
The most severe clinical picture is melioidosis septic shock, which is often associated with bacterial dissemination to distant sites such as the lungs, liver and spleen. The lungs are the most commonly affected organ in adults, where there can be a localized or disseminated pulmonary infection, abscess formation or empyema. Chronic lung disease can also occur and can be difficult to distinguish from pulmonary tuberculosis. The clinical features of the disease in Thailand and Northern Australia (where most cases are reported) are largely shared, but there are some striking differences. Acute suppurative parotitis is the presenting feature in one-third of Thai paediatric cases but is uncommon in Australia; conversely, prostatic abscesses and brainstem encephalitis are more frequent in Australia2,4. Pictures courtesy of Dr Wirongrong Chierakul, Wellcome Trust, Faculty of Tropical Medicine, Mahidol University, Bangkok, Thailand. CNS, central nervous system.
Melioidosis can present with an array of clinical signs and symptoms and B. pseudomallei has been called 'the great mimicker'. There can be a prolonged period between exposure to the causative agent and the clinical manifestations of infection (the longest recorded incubation period documented is 62 years3). Furthermore, recurrence of infection is common despite adequate antimicrobial therapy2. B. pseudomallei is intrinsically resistant to many antibiotics (including penicillin, first- and second-generation cephalosporins, macrolides, rifamycins, colistin and aminoglycosides), but is usually susceptible to amoxicillin-clavulanate, chloramphenicol, doxycycline, trimethoprim-sulphamethoxazole, ureidopenicillins, ceftazidime and carbapenems2,4. Treatment is required for 20 weeks and is divided into intravenous and oral phases2,4. Initial intravenous therapy is given for 10–14 days; ceftazidime or a carbapenem are the drugs of choice. The overall mortality for primary disease is 50% in northeast Thailand (35% in children) and ∼20% overall in the higher-technology setting of Northern Australia2,5. Interest in the pathogenesis of B. pseudomallei and the related bacterium Burkholderia mallei has increased following their classification as category B agents by the US Centers for Disease Control and Prevention. This review presents an overview of our current knowledge of disease acquisition and the host–pathogen interactions that follow.
Taxonomy and genomics
B. pseudomallei is an aerobic, motile, non-spore-forming bacillus (Fig. 2). The genus Burkholderia contains >30 species, the most pathogenic members of which are B. pseudomallei, B. mallei and, in certain clinical conditions such as cystic fibrosis, Burkholderia cepacia (Fig. 3). The genus also includes Burkholderia thailandensis , which coexists with B. pseudomallei in the soil in Thailand but rarely causes disease and is >105-fold less virulent than B. pseudomallei in Syrian hamsters or mice6. B. mallei causes glanders in horses and is potentially highly virulent in humans, but natural disease in any host is now extremely rare.
a | Typical colony morphology of B. pseudomallei on Ashdown's agar after incubation at 37°C in air for 3 days. b,c | Colony variation is commonly seen during culture of clinical isolates on Ashdown's agar. (c) shows the variable colony morphology that can be seen from a single sample; genotyping of these colonies showed that one clonal type was present. Colony variation can also be seen within a single colony, as shown in (b) in which the parental colony (pink) has given rise to a second morphotype (red). Pictures courtesy of Mrs Vanaporn Wuthiekanun and Mrs Narisara Chantratita, Wellcome Trust, Faculty of Tropical Medicine, Mahidol University, Bangkok, Thailand.
The phylogenetic tree is based on novel recA sequences. Most Burkholderia species are plant pathogens, but two groups can cause disease in humans. The Burkholderia cepacia complex is a group of important opportunistic pathogens for individuals with cystic fibrosis, with Burkholderia cenocepacia the cause of 70% of cases of Burkholderia cepacia complex infection101. Burkholderia pseudomallei and Burkholderia mallei have the potential to cause human disease, but the related Burkholderia thailandensis is rarely pathogenic. In this analysis, recA sequences from the related genera Ralstonia, Bordetella, Xanthomonas and Neisseria were included, and the tree was rooted with the Pseudomonas aeruginosa PAO1 recA gene. Reproduced with permission from Ref. 102 © (2005) American Society for Microbiology.
The genome of B. pseudomallei (strain K96243 from Thailand) has been sequenced and comprises two chromosomes of 4.07 Mb and 3.17 Mb7. The large chromosome carries many genes associated with core functions such as cell growth and metabolism, and the smaller chromosome carries more genes encoding accessory functions that could be associated with adaptation and survival in different environments. Approximately 6% of the genome is made up of putative genomic islands that have probably been acquired through horizontal gene transfer. These are mostly absent from the B. thailandensis genome (and are absent from the B. mallei genome8); it is unclear whether these regions have a role in disease pathogenesis. The Institute for Genomic Research (TIGR, Rockville, Maryland, USA) is currently sequencing nine further B. pseudomallei isolates and 25 B. pseudomallei bacteriophage genomes from various sources. In addition, shotgun sequencing of the B. thailandensis strain E264 genome is now complete. The molecular epidemiology of B. pseudomallei has been investigated using multilocus sequence typing (MLST), and the findings suggest a high rate of genetic recombination9.
Whole-genome comparison between B. pseudomallei and B. mallei suggests that B. mallei has evolved through 'genomic downsizing' from a single clone of B. pseudomallei (this is consistent with a previous conclusion drawn from MLST9). A DNA microarray based on the whole-genome sequence of B. pseudomallei K96243 has been used to compare isolates of B. pseudomallei, B. mallei and B. thailandensis10. Deleted regions in B. mallei had significant genomic clustering compared with those in B. thailandensis, which were more uniformly dispersed. This indicates that the evolutionary processes that resulted in the divergence of these three species might have distinct mechanisms. Subtractive hybridization between B. pseudomallei and B. thailandensis has revealed several loci that are unique to the former11,12; publication of whole-genome comparisons between the two species is currently awaited.
Factors associated with disease acquisition
Melioidosis is found only in individuals who have been exposed to environments containing B. pseudomallei; infection is acquired through cutaneous inoculation, inhalation and aspiration. The factors associated with disease acquisition in endemic regions include environmental and host factors. There is no evidence that some isolates of B. pseudomallei are intrinsically more infectious than others.
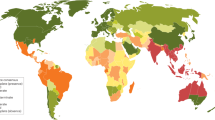
Environmental exposure. There is a positive association between disease incidence and the extent of environmental contamination with B. pseudomallei13,14,15. However, few environmental-sampling studies have been published, and a more complete picture of the geographical distribution of B. pseudomallei can be derived from the reported cases of melioidosis. This topic has recently been reviewed by Cheng and Currie4. The disease is endemic in parts of Thailand, Northern Australia, Malaysia, Singapore, Vietnam and Burma. Possible endemic areas include Southern India, Southern China, Hong Kong, Taiwan, Brunei, Laos and Cambodia. Sporadic cases and occasional clusters have been reported in large areas of Asia, the Americas (notably Brazil), the Caribbean, the Pacific, Africa and the Middle East.
Weather conditions, route of acquisition and inoculum. Melioidosis is seasonal in the tropics, where most cases occur during the rainy season. This can be explained by increased contact with the organism. Rice farmers plant at the start of the monsoon and work in flooded rice paddies until harvest. Thai farmers rarely wear protective footwear and their feet often show signs of repeated trauma and injuries. Extreme weather can be associated with a shift in the mode of acquisition of infection. Aerosols are created during heavy rain, and this can result in repeated inhalation of the organism. Heavy rainfall and winds consistently cause a shift towards more pneumonia in patients presenting with melioidosis in Northern Australia16. Severe or penetrating injury and near-drowning are known risk factors for melioidosis, as highlighted by a study of a cluster of melioidosis cases in Southern Thailand following the 2004 tsunami17.
Integrity of host immunity. In northeast Thailand, 80% of the population belongs to rice-farming families. Children have extensive contact with the organism, yet only one-fifth of all melioidosis cases here occur in children <14 years of age, with the disease incidence peaking later between the fourth and sixth decade of life. Most affected adults (>80%) have one or more underlying diseases (most commonly diabetes mellitus or renal failure). By contrast, children have an identifiable risk factor in <30% of cases (most commonly trauma). It is unclear whether affected children have a greater genetic susceptibility for disease. It is also possible that disease in childhood is caused by a subset of the bacterial population with increased pathogenic potential.
Immune response in the exposed but healthy population. Seroprevalence studies in northeast Thailand based on the indirect haemagglutination assay show that ∼80% of people have antibodies against B. pseudomallei by the age of 4 years18. It is not clear whether healthy individuals with high antibody titres are infected and have a quiescent focus (analogous to a quiescent tuberculosis infection), or whether repeated environmental exposure in a primed individual maintains high antibody levels. The strong seasonality in disease presentation, combined with a reported mean incubation period of 9 days19, suggests that primary disease occurs as a result of new infection rather than seasonal activation of a persistent focus. The possibility remains that the development of underlying risk factors causing immune dysfunction is seasonal, and this could be followed by seasonal disease breakthrough as bacteria escape from immune surveillance, but this is less biologically plausible. The lack of seasonality in documented relapse argues against this hypothesis. One episode of melioidosis does not protect susceptible individuals from further episodes of melioidosis due to re-infection20.
Putative virulence factors
There is a paucity of knowledge in this area compared with the knowledge for other Gram-negative bacteria. The factors described here are included on the basis of a known role in virulence for other pathogens, or virulence in experimental models, and have been grouped according to the strength of existing evidence.
Strong putative candidates: quorum sensing. Quorum sensing is a cell-density-dependent communication system in Gram-negative bacteria that uses N-acyl-homoserine lactones (AHLs) for the coordination of gene expression21,22. LuxI proteins are responsible for AHL biosynthesis, and LuxR transcriptional regulators, following association with their cognate AHL(s), mediate gene repression or expression21,22. The B. pseudomallei genome is reported to contain three LuxI and five LuxR quorum-sensing homologues22. Mass-spectrometry analysis of B. pseudomallei culture supernatants has demonstrated the presence of many signalling molecules, including N-decanoyl-homoserine-lactone and N-(3-oxotetradecanoyl)-L-homoserinelactone22. Disruption of the eight genes encoding luxIR quorum-sensing homologues led to a significant increase in the LD50 (the infectious dose that is lethal to 50% of the animals infected) in Syrian hamsters after intraperitoneal challenge, and increased the time to death and reduced organ colonization in aerosolized BALB/c mice22. A LuxI-LuxR homologue termed PmlI-PmlR, which directs the synthesis of N-decanoyl-homoserine-lactone and is involved in regulation of a metalloprotease, is essential for full virulence in a mouse model23. A homologue termed BpsI-BpsR is also required for optimal virulence and the secretion of exoproducts24.
Some quorum-sensing-controlled candidate virulence factors and processes such as siderophores, phospholipase C and biofilm formation are probably partially dependent on BpeAB-OprB, a multidrug efflux pump in B. pseudomallei that is also known to be responsible for conferring antimicrobial resistance to aminoglycosides and macrolides25. The bpeAB-oprB operon in turn might be regulated by quorum sensing, as N-decanoylhomoserine-lactone and N-octanoyl-homoserine lactone can induce bpeAB-oprB expression25. BpeAB mutants are also associated with attenuated cell invasion and cytotoxicity of human lung epithelial (A549) and human macrophage (THP-1) cells25.
Strong putative candidates: type III secretion system. B. pseudomallei contains three type III secretion system (TTSS) gene clusters, which contain between 16 and 18 reported genes7,26,27,28,29. One of these clusters (the TTSS3 cluster) shares homology with the inv/spa/prg TTSS of Salmonella enterica serovar Typhimurium (S. typhimurium) and the ipa/mxi/spa TTSS cluster of Shigella flexneri 26,28,29. This cluster encodes a secretion apparatus that functions like a molecular syringe. A subset of type-III-secreted proteins ('translocators') interact with the eukaryotic cell membrane and inoculate other type-III-secreted 'effector' proteins into the target-cell cytosol, where they subvert host-cell processes (reviewed by Cornelis and Van Gijsegem30). Mutations that disrupt the S. typhimurium Inv/Spa/Prg apparatus inhibit bacterial invasion and enteropathogenesis. The gene cluster in B. pseudomallei (termed bsa, Burkholderia secretion apparatus) encodes proteins that are very similar to the S. typhimurium and S. flexneri type-III-secreted proteins required for invasion, escape from endocytic vacuoles, intercellular spread and pathogenesis29.
Following internalization, B. pseudomallei escapes from endocytic vacuoles into the cytoplasm of infected cells (Fig. 4). Induction of actin polymerization at one pole leads to the formation of membrane protrusions and cell-to-cell spread. B. pseudomallei mutants lacking components of the Bsa secretion and translocation apparatus have reduced replication in murine macrophage-like cells, an inability to escape from endocytic vacuoles and cannot form membrane protrusions and actin tails29. Inactivation of BopE, a TTSS protein that is encoded adjacent to the B. pseudomallei bsa locus and is homologous to Salmonella enterica SopE/SopE2, a guanine nucleotide-exchange factor, leads to impaired bacterial entry into HeLa cells, indicating that BopE facilitates invasion31. In addition, the B. pseudomallei bsa locus encodes homologues of the Salmonella Sip translocator proteins (BipB, BipC and BipD)29. Salmonella SipB, SipC and SipD proteins are required for injection of effector proteins and invasion of epithelial cells in vitro32; mutation of the B. pseudomallei bipD gene impairs invasion of epithelial cells in vitro31. B. pseudomallei BipD mutants that lack a component of the translocation apparatus are attenuated following intraperitoneal or intranasal challenge of BALB/c mice, and have impaired bacterial replication in liver and spleen32. B. pseudomallei BipB has been shown to mediate the formation of multinucleated giant cells, cell-to-cell spreading of bacteria and apoptosis of infected host cells33. bipB mutants are also associated with attenuation following intranasal challenge of BALB/c mice33.
a | Invasion of phagocytic and non-phagocytic cells. B. pseudomallei is capable of invading many cell types, including epithelial cells, and can survive and proliferate for prolonged periods within phagocytic cells. The Inv/Mxi/Spa-like type III secretion gene cluster, termed the Burkholderia secretion apparatus (bsa) system, encodes proteins that are required for invasion, escape from phagosomes and intercellular spread29. b | Endosome escape and intracellular proliferation. As early as 15 minutes after internalization, B. pseudomallei can escape from endocytic vacuoles into the cytoplasm of infected cells by lysing the endosome membrane. B. pseudomallei seems to be resistant to several host antimicrobial peptides (for example, protamine and certain defensins) and interferes with the synthesis of inducible nitric-oxide synthase (iNOS), which is known to have an important role in the killing of intracellular bacteria73,79. In addition, in certain circumstances B. pseudomallei can induce apoptosis in both phagocytic and non-phagocytic cells75. c | Cell-to-cell spread. Once inside the cytoplasm, B. pseudomallei induces the formation of actin-based membrane protrusions by continuous nucleation of actin at one pole of the bacterial cell29,75,81. The bacterial protein BimA is required for this process82. Cell-to-cell movement of B. pseudomallei occurs when a neighbouring cell phagocytoses this protrusion, thereby allowing the spread of B. pseudomallei without exposure to antibodies or immunoactive molecules. In the new cell, B. pseudomallei will subsequently escape from the secondary vacuoles and multiply intracellularly. d | Cell fusion. Uniquely among bacterial intracellular pathogens, B. pseudomallei can induce the formation of multinucleated giant cells by cell fusion75.
The TTSS1 gene cluster was first described in 1999 (Ref. 34). This cluster is homologous to a TTSS found in the plant pathogen Ralstonia solanacearum but is absent from B. mallei and B. thailandensis7,26,35. TTSS2 is present in R. solanacearum, B. pseudomallei, B. mallei and B. thailandensis. Whereas TTSS3 has been shown to be required for full virulence in a hamster model of infection27, the role of TTSS1 and TTSS2 in pathogenesis is not known.
Strong putative candidates: capsular polysaccharide. B. pseudomallei produces an extracellular capsular polysaccharide with the structure -3)-2-O-acetyl-6-deoxy-β-D-manno-heptopyranose-(1- (Refs 11,36). Previously characterized as a type I O-polysaccharide, it has more recently been considered to be a capsular polysaccharide based on its high molecular mass and genetic homology with group 3 capsular polysaccharides of other organisms. The capsular polysaccharide is required for B. pseudomallei virulence in experimental animal models11,37. Capsule expression is induced in the presence of serum, and the addition of purified B. pseudomallei capsule to serum bactericidal assays increases the survival of B. pseudomallei SLR5, a serum-sensitive strain, by 1,000-fold38. Phagocytosis is greater for a capsule-deficient mutant compared with the wild type in the presence of normal human serum38. Both observations can be explained by the finding that deposition of complement factor C3b on the bacterial cell surface is lower in the presence of capsule38. The capsule might act as a barrier, blocking access of the complement receptor-1 (CR1) on phagocytes to the C3b deposited on the bacterial surface38.
Other putative candidates: lipopolysaccharide. B. pseudomallei lipopolysaccharide (LPS) (formally termed type II O-antigenic polysaccharide) seems to differ in several respects from the LPS of other Gram-negative organisms. B. pseudomallei LPS exhibits weaker pyrogenic activity in rodents compared with enterobacterial LPS, but stronger mitogenic activity in murine splenocytes39. LPS-mediated activation of a mouse-macrophage cell line in vitro is slower for LPS from B. pseudomallei compared with LPS from Escherichia coli 40. Recognition of LPS by the host is of crucial importance for the initiation of a swift innate immune response to Gram-negative bacteria, primarily through activation of the pattern-recognition receptor Toll-like receptor (TLR)4 (see also further discussion)41. So, the fact that B. pseudomallei LPS is apparently less capable of activating immune cells might at least in part explain why TLR4 does not have a role in host defence against experimentally induced melioidosis in mice, whereas mice deficient for this receptor are highly susceptible to other Gram-negative infections41.
B. pseudomallei LPS seems to be largely conserved across this species. LPS profiling of >700 B. pseudomallei isolates using proteinase-K digestion and SDS-PAGE silver-stained gels, a technique that examines the O-side chain, demonstrated that most isolates had a 'typical' ladder pattern of extracted LPS, 3% had an 'atypical' pattern, and 0.1% did not exhibit a ladder appearance at all42. The different LPS preparations have similar endotoxic activity in the Limulus amoebocyte lysate assay. However, there seems to be a difference in the host immune response to these molecules, as there is a lack of immunological crossreactivity on western blot between typical and atypical LPS using patient sera infected with typical and atypical LPS isolates42.
The level of antibody to LPS on admission to hospital is higher in patients with melioidosis who survive compared with those who die, and in patients with non-septicaemic versus septicaemic melioidosis43. These antibodies might protect the host against death; alternatively, there could be an association between a raised anti-LPS antibody titre and a more efficient host immune response, including cell-mediated killing.
The LPS of B. pseudomallei (pathogenic) and B. thailandensis (non-pathogenic) have been compared. LPS profiling using proteinase-K digestion and SDS-PAGE silver-stained gel shows identical ladder patterns for most isolates of both species. The two species exhibit similar immunoblot profiles against pooled sera from patients with melioidosis, and with hyperimmune mouse sera44. LPS shedding profiles are also similar between the two species45. This has led to the suggestion that LPS is unlikely to be involved in the virulence and pathogenicity of B. pseudomallei44. Other possible explanations are that LPS from the two species is antigenically very similar but differs in biological activity or that LPS from both species has biological activity in vivo, but only B. pseudomallei has an additional complement of genes that promote successful invasion and bacterial dissemination within the host. Caution is required when interpreting the significance of LPS in B. pseudomallei virulence, as assays in which LPS is truncated could impair the insertion, stability and folding of other surface-anchored molecules.
Other putative candidates: flagella. B. pseudomallei is flagellated and motile. There is no difference in the ability of wild-type B. pseudomallei and an isogenic mutant defective in flagella expression to invade, and replicate in, human lung cells in vitro46. In one study, there was no difference between an isogenic mutant and wild-type B. pseudomallei in diabetic rat and Syrian hamster infection models47. In a second study, bacterial numbers were markedly reduced in the lung and spleen of BALB/c mice following intranasal infection of an aflagellate mutant compared with the wild type, and the mutant was less virulent following intraperitoneal infection of BALB/c mice, based on the LD50 (Ref. 46).
Other putative candidates: type IV pili-mediated adherence. Adherence is an important virulence mechanism mediated by carbohydrate molecules, pilus and non-pilus adhesins. Type IV pili are important for virulence in many Gram-negative bacteria. The B. pseudomallei K96243 genome contains multiple type IV pilin-associated loci, including one encoding a putative pilus structural protein (PilA)48. A pilA deletion mutant has reduced adherence to human epithelial cells and is less virulent in the nematode model of virulence and the murine model of melioidosis, suggesting a role for type IV pili in B. pseudomallei virulence.
Other putative candidates. Other putative candidiates include a siderophore for iron acquisition49 and secreted proteins such as haemolysin, lipases and proteases50. Bacterial colony morphology of clinical B. pseudomallei isolates can vary both within a given culture and in cultures from the same patient over time (Fig. 2). The relevance of this to human disease is the subject of intensive investigation in our laboratories.
Downregulation of virulence. An arabinose-assimilation operon that consists of nine genes is present in B. thailandensis and absent from the B. pseudomallei and B. mallei chromosomes. When this operon is cloned experimentally and re-introduced into a laboratory strain of B. pseudomallei, the mutant strain has a reduced ability to cause the death of Syrian hamsters compared with the parent strain51. On microarray analysis, several genes in the TTSS3 cluster are downregulated in the mutant when cells are grown in L-arabinose, suggesting a regulatory role for (a metabolite of) L-arabinose51. This could be one of many similar examples.
Host–pathogen interactions during melioidosis
Innate immune response. On the first encounter with a pathogen, cells of the innate immune system recognize conserved surface motifs termed 'pathogen-associated molecular patterns' or PAMPs through host-cell pattern-recognition receptors. The family of TLRs are important members of this surveillance system which initiate the innate immune response, and form a key link between innate and adaptive immunity52. B. pseudomallei expresses several PAMPs for which the corresponding TLR is known (Fig. 5). For example, in other Gram-negative species it has been shown that LPS activates the cells of the immune system through a receptor complex that consists of a ligand-binding molecule (CD14) and TLR4 as the signal transducer. Other candidate B. pseudomallei TLR ligands include peptidoglycan (TLR2), flagellin (TLR5) and bacterial DNA or CpG (TLR9). However, to date experimental data for a role for TLRs in melioidosis are lacking, although it is interesting to note that C3H/HeJ mice that carry a loss-of-function mutation in the tlr4 gene are resistant to extremely high doses of B. pseudomallei LPS (up to 10,000 ng per mouse)39.
Proposed scheme of the first encounter between B. pseudomallei and the immune system. Putative virulence factors on the bacterial cell surface include lipopolysaccharide (LPS), capsular polysaccharides and flagella. There might also be a role for biofilm formation. Bacterial gene regulation through N-acyl-homoserine lactones (AHLs) could be crucial for bacterial survival in vivo. Type III secretion systems (TTSSs) might facilitate invasion of the bacterium into the host cell, and enable escape from endocytic host-cell vesicles. Monocytes are probably the most important immune cells in early infection. Based on evidence from other Gram-negative pathogens, there might be a role for the Toll-like receptors (TLRs). TLR4, with its co-receptors MD2 and CD14, might recognize B. pseudomallei LPS, whereas TLR5 might recognize flagella, although to date there are no published data to support this. The complement receptors CR1, CR3 and possibly FcγR mediate opsonin-dependent phagocytosis. Recognition of B. pseudomallei will cause activation of pro-inflammatory genes through nuclear factor (NF)-κB and lead to the activation of the immune response through the release of pro-inflammatory cytokines.
The pro-inflammatory cytokine interferon (IFN)-γ has an important role in early resistance against B. pseudomallei infection. Inhibition of IFN-γ expression in mice lowered the LD50 from >5 × 105 to ∼2 colony-forming units (CFUs) and was associated with an 8,500- and 4,400-fold increase in bacterial load in liver and spleen, respectively53. Inhibition of interleukin (IL)- 12 or IL-18, the predominant endogenous inducers of IFN-γ production, resulted in increased mortality in the same model53,54. IFN-γ, IL-12 and IL-18 have a key role in the TH1 cell-mediated immune response. Recent data indicate that activation of suppressor of cytokine signalling 3 (SOCS3) and cytokine-inducible Src homology 2-containing protein (CIS) in B. pseudomallei-infected macrophages correlates with a decreased IFN-γ signalling response and facilitates bacterial escape55. Further evidence for the role of a TH1 response in protective immunity against melioidosis comes from studies with inbred mouse strains, in which TH1-response-prone C57BL/6 mice are relatively resistant to B. pseudomallei when compared with the TH2-response-prone BALB/c mice56,57. The pro-inflammatory cytokine tumour-necrosis factor (TNF)-α is also likely to be an important element of the early immune response, as passive immunization against this mainly macrophage-derived cytokine increased mortality in experimental murine melioidosis53. Serum IFN-γ, IL-12 and TNF-α concentrations are elevated in melioidosis patients58,59; the involvement of TNF-α in human disease is further suggested by a report that the −308 TNF-α promoter polymorphism, which is related to severity of disease for several other infectious diseases, was associated with both the occurrence and severity of melioidosis60.
Plasma or serum concentrations of several other pro-inflammatory mediators are elevated in patients with melioidosis, including IL-6, IL-15, IFN-γ-inducible protein (IP)-10 and monokine induced by IFN-γ (Mig)58,61, as well as the anti-inflammatory cytokine IL-10 (Ref. 62). This indicates that multiple inflammatory pathways become activated, including those involving cellular activation. For example, IP-10 and Mig are chemokines primarily induced by IFN-γ which share a common receptor (CXC chemokine receptor 3, CXCR3), expressed by activated T cells and natural killer (NK) cells. Cytotoxic T cells and NK cells are further implicated in the immune response to B. pseudomallei through the observation that circulating levels of granzymes A and B are elevated. Granzymes are secreted by cytotoxic T and NK cells and are important for the initiation of apoptosis in the target cell, although their precise role in bacterial infection remains to be established63. NK cells and CD8+ T cells harvested from the spleen of uninfected mice also release IFN-γ on stimulation with B. pseudomallei in vitro64, and during experimental melioidosis NK and T cells have been identified as the predominant IFN-γ producers54.
B. pseudomallei rapidly and efficiently activates complement, predominantly through the alternative activation pathway65. Complement activation results in the deposition of C3 on the bacterial surface and further opsonisation65. However, B. pseudomallei is resistant to the lytic action of complement65, a feature shared with several other pathogens including Borrelia spp., Salmonella spp., Neisseria spp. and streptococci66. In one study, adherence and phagocytosis of B. pseudomallei by granulocytes and macrophages was dependent on the presence of opsonins and mediated by the complement receptors CR1 and CR3 expressed on the macrophage membrane65. The capsular polysaccharide of B. pseudomallei contributes to resistance to phagocytosis by reducing the deposition of complement factor C3b (Ref. 38).
Adaptive immune response. In patients with melioidosis, the levels of IgG, IgA and IgM correlate positively with disease severity, with higher levels in those with invasive compared with localized disease. Melioidosis is positively associated with specific human leukocyte antigen (HLA) class II molecules in Thai patients; the HLA class II DRB1*1602 allele was positively associated with septicaemic melioidosis, whereas DQA1*03 was negatively associated67. Patients who recovered from melioidosis showed evidence for an antigen-specific cell-mediated immune response, as reflected by enhanced lymphocyte proliferation and IFN-γ production in response to B. pseudomallei antigens68. In addition, asymptomatic seropositive individuals showed a stronger cell-mediated adaptive immune response as measured by Burkholderia-specific lymphocyte reactions compared with subjects with a history of clinical melioidosis69, suggesting that a strong cell-mediated immune response might protect against disease progression. Conceivably, the production of IFN-γ by CD4+ T cells activates macrophages to become more bactericidal, a notion that is supported by the finding that IFN-γ increases the intracellular killing activity of macrophages in vitro70. Of considerable interest, a very recent study has suggested that B. pseudomallei-specific CD4+ T cells are important for late host resistance against murine melioidosis54. However, the importance of CD4+ T cells in the control of infection is open to debate, as there does not seem to be an association between HIV infection and melioidosis71.
Intracellular survival of B. pseudomallei. In vitro models indicate that B. pseudomallei survives and replicates within neutrophils and monocytes72,73,74,75, and uses multiple mechanisms to escape macrophage killing and evade host immunity. These include resistance to human defensins72 and inhibition of DNA and protein synthesis within the host cell76. Ultrastructural studies suggest that B. pseudomallei might reside within membrane-bound compartments73, particularly phagolysosomes65, making use of its ability to survive and grow in acidic environments77. Furthermore, B. pseudomallei can invade mouse macrophages without activating inducible nitric-oxide synthase (iNOS), an enzyme that is required for the generation of reactive nitrogen intermediates and is important for intracellular bacterial killing78. B. pseudomallei can escape from endocytic vesicles into the cytoplasm (Fig. 4) by lysing the endosome membranes as early as 15 minutes after internalization by phagocytic cells29,79,80. Moreover, intracellular B. pseudomallei can be propelled by inducing continuous polymerization of actin at one pole of the bacterial cell29,75,81, which results in the formation of membrane protrusions in host cells with the bacteria at the tip end. Such protrusions can project into an adjacent cell, facilitating the spread B. pseudomallei from one eukaryotic cell to another75(Fig. 4). A B. pseudomallei-specific protein termed BimA is required for the formation of actin tails. BimA is located at the pole of the bacterial cell at which actin polymerization occurs82. Mutation of the bimA gene in B. pseudomallei abolished actin-based motility of intracellular bacteria in a macrophage-like cell line. Actin-tail formation could be restored by inducible expression of the bimA gene on a plasmid, indicating the essential role for this gene in actin-tail formation. Of note, mutation of bimA does not influence the activity of the Bsa TTSS apparatus or bacterial escape from endosomes.
Multinucleated giant cells are occasionally seen in human tissue infected with B. pseudomallei and might represent giant macrophages83. It is possible that macrophages are a site of intracellular survival, given their relatively long half-life and reduced microbicidal capacity in comparison to neutrophils and monocytes65. A recent report indicated that B. pseudomallei might evade killing by macrophages through the induction of caspase-1-dependent host-cell death; this process requires a functional bsa TTSS84. Cell death was accompanied by the release of IL-1β and IL-18 (Ref. 84). Survival within microcolonies encapsulated in a protective biofilm is an alternative explanation for prolonged quiescent survival within the host. In vitro, B. pseudomallei can survive for years in distilled water and can also enter and survive within free-living amoebae belonging to the genus Acanthamoeba; it is possible that this might enhance bacterial survival in soil and aquatic environments85,86.
Interactions with human epithelial cells in vitro. B. pseudomallei adheres to cultured human epithelial cell lines derived from alveolar, bronchial, laryngeal, oral, conjunctival and cervical tissues87. Adherence of B. pseudomallei (but not B. thailandensis) to cell lines in vitro was enhanced when incubated at a temperature of 30°C compared with 37°C (Ref. 87). B. pseudomallei is more efficient in invasion, adherence and induction of cellular damage of respiratory epithelial cells compared with B. thailandensis88. A bacterial mutant defective in pilA (a putative type IV pilus gene) has reduced adherence to human epithelial cells48. The relevance of these observations for human disease pathogenesis is unknown.
New therapeutic and vaccine options
Potential new therapies targeting the immune response. Several conditions including diabetes mellitus, renal failure and alcohol abuse are important risk factors for the development of melioidosis. A common link is that these are associated with impairment of neutrophil function. This had led to interest in the therapeutic role of granulocyte colony-stimulating factor (G-CSF), a cytokine that increases the circulating neutrophil count and stimulates neutrophil function. A retrospective study conducted in Australia reported a reduction in mortality of melioidosis patients after the introduction of G-CSF as an adjunctive treatment for patients with septic shock89. A mouse model of melioidosis in which outcome was compared between mice given ceftazidime alone or in combination with G-CSF failed to show any benefit from G-CSF90. A randomized clinical trial of G-CSF is currently ongoing in Thailand91.
Unmethylated CpG motifs in synthetic oligodeoxynucleotide enhance the uptake of bacteria by mouse macrophages in a concentration-dependent manner and induce nitric-oxide production by mouse macrophages92. CpG treatment one hour before bacterial inoculation offered protection in a murine model of B. pseudomallei infection, associated with the rapid induction of pro-inflammatory cytokines93.
Vaccine development. There is no effective vaccine available that protects against B. pseudomallei infection. Current approaches under evaluation include conjugate, DNA, attenuated and heterologous vaccines94. Attenuated mutants that are invasive but have a reduced ability to multiply in phagocytes have been identified using transposon mutagenesis, and these induce a high degree of protective immunity in mice (K. Breitbach, personal communication). A mutant of B. pseudomallei that is auxotrophic for branched-chain amino acids induced a protective response against a subsequent challenge with an otherwise lethal dose of wild-type B. pseudomallei in an animal model95. Mice inoculated with a B. pseudomallei bipD mutant were partially protected against subsequent challenge with wild-type B. pseudomallei, although immunization with purified BipD protein was not protective32. However, it seems unlikely that live attenuated vaccination will be feasible for use in humans. Antibodies raised against B. pseudomallei flagellin markedly reduced bacterial motility and provided passive protection against experimentally induced B. pseudomallei infection96. Evaluation of LPS and capsular polysaccharide as subunit vaccines against experimental melioidosis demonstrated partial protection in a mouse model97. Inoculation of guinea pigs with B. thailandensis provided partial protection from a subsequent challenge with virulent B. pseudomallei98.
In addition, it was recently postulated that the T-cell response to primary B. pseudomallei infection is biphasic, comprising an early cytokine — most notably IFN-γ induced — phase in which T cells seem to be functionally redundant for initial bacterial clearance, followed by a later antigen-induced phase in which B. pseudomallei-specific T cells (in particular CD4+ T cells) are important for host resistance55. As a consequence, because B. pseudomallei infection generates a rapid and potent IFN-γ response from natural killer T cells and conventional T cells, it is has been suggested that an effective subunit vaccine against B. pseudomallei should target the generation of IFN-γ-secreting T cells54. This was further underscored by a recent vaccine-driven report suggesting that any potential vaccine would need to stimulate both cell-mediated and humoral immunity99. In this study, dendritic cells were used as a vaccine-delivery vector to induce cell-mediated immune responses to B. pseudomallei. Purified dendritic cells were pulsed with heat-killed whole-cell B. pseudomallei and used to immunize syngeneic mice. Strong cellular immune responses were elicited, although antibody responses were low. Subsequently, booster immunizations of either a second dose of dendritic cells or heat-killed B. pseudomallei were administered to increase the immune response. Immunized animals were challenged with fully virulent B. pseudomallei, and protection was shown in those with strong humoral and cell-mediated immunity99. The possible utility of these observations for vaccine development awaits further investigation.
Conclusions and perspectives
The recent advent of molecular genetics and in vitro and in vivo infection models, together with availability of the complete genome sequence of B. pseudomallei, B. mallei and (in the near future) B. thailandensis, has advanced our understanding of the virulence mechanisms that govern the complex interaction between B. pseudomallei and host cells. Preserved mechanisms underlying the intracellular lifestyle of B. pseudomallei have been discovered by comparing the B. pseudomallei genome with the genomes of other intracellular microorganisms (for example, Salmonella and Shigella species)86. New techniques that allow integrative analyses at genomic, transcriptional and proteomic levels promise to provide further insights into the complex interaction between this pathogen and its environment100. The genetic diversity within the natural population will be characterized further using techniques such as MLST and microarray analysis, along with the identification of the B. pseudomallei secretome (all secreted proteins secreted) and immunome (identification of immunogenic proteins). Some virulence factors have now been characterized, and the study of host–pathogen interactions has shed some light on pathogenic mechanisms, but many key questions await investigation, and Box 1 summarizes some important questions for future research on B. pseudomallei pathogenesis.
References
Dance, D. A. Melioidosis: the tip of the iceberg? Clin. Microbiol. Rev. 4, 52–60 (1991).
White, N. J. Melioidosis. Lancet 361, 1715–1722 (2003).
Ngauy, V., Lemeshev, Y., Sadkowski, L. & Crawford, G. Cutaneous melioidosis in a man who was taken as a prisoner of war by the Japanese during World War II. J. Clin. Microbiol. 43, 970–972 (2005).
Cheng, A. C. & Currie, B. J. Melioidosis: epidemiology, pathophysiology, and management. Clin. Microbiol. Rev. 18, 383–416 (2005).
Cheng, A. C. et al. Melioidosis in northern Australia, 2001–02. Commun. Dis. Intell. 27, 272–277 (2003).
Brett, P. J., DeShazer, D. & Woods, D. E. Burkholderia thailandensis sp. nov., a Burkholderia pseudomallei-like species. Int. J. Syst. Bacteriol. 48, 317–320 (1998). First description of B. thailandensis.
Holden, M. T. et al. Genomic plasticity of the causative agent of melioidosis, Burkholderia pseudomallei. Proc. Natl Acad. Sci. USA 101, 14240–14245 (2004). Provided the complete genome sequence of B. pseudomallei and a description of the pathogenicity islands.
Nierman, W. C. et al. Structural flexibility in the Burkholderia mallei genome. Proc. Natl Acad. Sci. USA 101, 14246–14251 (2004).
Godoy, D. et al. Multilocus sequence typing and evolutionary relationships among the causative agents of melioidosis and glanders, Burkholderia pseudomallei and Burkholderia mallei. J. Clin. Microbiol. 41, 2068–20679 (2003). Description of the development of MLST for B. pseudomallei.
Ong, C. et al. Patterns of large-scale genomic variation in virulent and avirulent Burkholderia species. Genome Res. 14, 2295–2307 (2004).
Reckseidler, S. L., DeShazer, D., Sokol, P. A. & Woods, D. E. Detection of bacterial virulence genes by subtractive hybridization: identification of capsular polysaccharide of Burkholderia pseudomallei as a major virulence determinant. Infect. Immun. 69, 34–44 (2001).
Brown, N. F. & Beacham, I. R. Cloning and analysis of genomic differences unique to Burkholderia pseudomallei by comparison with B. thailandensis. J. Med. Microbiol. 49, 993–1001 (2000).
Vuddhakul, V. et al. Epidemiology of Burkholderia pseudomallei in Thailand. Am. J. Trop. Med. Hyg. 60, 458–461 (1999).
Parry, C. M. et al. Melioidosis in Southern Vietnam: clinical surveillance and environmental sampling. Clin. Infect. Dis. 29, 1323–1326 (1999).
Wuthiekanun, V. et al. Detection of Burkholderia pseudomallei in soil within the Lao People's Democratic Republic. J. Clin. Microbiol. 43, 923–924 (2005).
Currie, B. J. & Jacups, S. P. Intensity of rainfall and severity of melioidosis, Australia. Emerg. Infect. Dis. 9, 1538–1542 (2003). Showed that heavy rains and winds can cause a shift towards inhalation of B. pseudomallei.
Chierakul, W. et al. Melioidosis in 6 tsunami survivors in Southern Thailand. Clin. Infect. Dis. 41, 982–990 (2005).
Kanaphun, P. et al. Serology and carriage of Pseudomonas pseudomallei: a prospective study in 1000 hospitalized children in northeast Thailand. J. Infect. Dis. 167, 230–233 (1993).
Currie, B. J. et al. The epidemiology of melioidosis in Australia and Papua New Guinea. Acta Trop. 74, 121–127 (2000).
Maharjan, B. et al. Recurrent melioidosis in patients in northeast Thailand is frequently due to reinfection rather than relapse. J. Clin. Microbiol. 43, 6032–6034 (2005).
Lazdunski, A. M., Ventre, I. & Sturgis, J. N. Regulatory circuits and communication in Gram-negative bacteria. Nature Rev. Microbiol. 2, 581–592 (2004).
Ulrich, R. L. et al. Role of quorum sensing in the pathogenicity of Burkholderia pseudomallei. J. Med. Microbiol. 53, 1053–1064 (2004).
Valade, E. et al. The PmlI-PmlR quorum-sensing system in Burkholderia pseudomallei plays a key role in virulence and modulates production of the MprA protease. J. Bacteriol. 186, 2288–2294 (2004).
Song, Y. et al. The BpsIR quorum-sensing system of Burkholderia pseudomallei. J. Bacteriol. 187, 785–790 (2005).
Chan, Y. Y. & Chua, K. L. The Burkholderia pseudomallei BpeAB-OprB efflux pump: expression and impact on quorum sensing and virulence. J. Bacteriol. 187, 4707–4719 (2005).
Rainbow, L., Hart, C. A. & Winstanley, C. Distribution of type III secretion gene clusters in Burkholderia pseudomallei, B. thailandensis and B. mallei. J. Med. Microbiol. 51, 374–384 (2002).
Warawa, J. & Woods, D. E. Type III secretion system cluster 3 is required for maximal virulence of Burkholderia pseudomallei in a hamster infection model. FEMS Microbiol. Lett. 242, 101–108 (2005).
Attree, O. & Attree, I. A second type III secretion system in Burkholderia pseudomallei: who is the real culprit? Microbiology 147, 3197–3199 (2001).
Stevens, M. P. et al. An Inv/Mxi-Spa-like type III protein secretion system in Burkholderia pseudomallei modulates intracellular behaviour of the pathogen. Mol. Microbiol. 46, 649–659 (2002).
Cornelis, G. R. & Van Gijsegem, F. Assembly and function of type III secretory systems. Annu. Rev. Microbiol. 54, 735–774 (2000).
Stevens, M. P. et al. A Burkholderia pseudomallei type III secreted protein, BopE, facilitates bacterial invasion of epithelial cells and exhibits guanine nucleotide exchange factor activity. J. Bacteriol. 185, 4992–4996 (2003).
Stevens, M. P. et al. Attenuated virulence and protective efficacy of a Burkholderia pseudomallei bsa type III secretion mutant in murine models of melioidosis. Microbiology 150, 2669–2676 (2004).
Suparak, S. et al. Multinucleated giant cell formation and apoptosis in infected host cells is mediated by Burkholderia pseudomallei type III secretion protein BipB. J. Bacteriol. 187, 6556–6560 (2005).
Winstanley, C., Hales, B. A. & Hart, C. A. Evidence for the presence in Burkholderia pseudomallei of a type III secretion system-associated gene cluster. J. Med. Microbiol. 48, 649–656 (1999).
Thibault, F. M., Valade, E. & Vidal, D. R. Identification and discrimination of Burkholderia pseudomallei, B. mallei, and B. thailandensis by real-time PCR targeting type III secretion system genes. J. Clin. Microbiol. 42, 5871–5874 (2004).
Steinmetz, I., Rohde, M. & Brenneke, B. Purification and characterization of an exopolysaccharide of Burkholderia (Pseudomonas) pseudomallei. Infect. Immun. 63, 3959–3965 (1995).
Atkins, T. et al. Characterisation of an acapsular mutant of Burkholderia pseudomallei identified by signature tagged mutagenesis. J. Med. Microbiol. 51, 539–547 (2002).
Reckseidler-Zenteno, S. L., DeVinney, R. & Woods, D. E. The capsular polysaccharide of Burkholderia pseudomallei contributes to survival in serum by reducing complement factor C3b deposition. Infect. Immun. 73, 1106–1115 (2005).
Matsuura, M., Kawahara, K., Ezaki, T. & Nakano, M. Biological activities of lipopolysaccharide of Burkholderia (Pseudomonas) pseudomallei. FEMS Microbiol. Lett. 137, 79–83 (1996).
Utaisincharoen, P. et al. Kinetic studies of the production of nitric oxide (NO) and tumour necrosis factor-α (TNF-α) in macrophages stimulated with Burkholderia pseudomallei endotoxin. Clin. Exp. Immunol. 122, 324–329 (2000).
Beutler, B. & Rietschel, E. T. Innate immune sensing and its roots: the story of endotoxin. Nature Rev. Immunol. 3, 169–176 (2003).
Anuntagool, N. et al. Antigenic heterogeneity of lipopolysaccharide among Burkholderia pseudomallei clinical isolates. Southeast Asian J. Trop. Med. Public Health 31 (Suppl. 1), 146–152 (2000).
Charuchaimontri, C. et al. Antilipopolysaccharide II: an antibody protective against fatal melioidosis. Clin. Infect. Dis. 29, 813–818 (1999).
Anuntagool, N., Intachote, P., Wuthiekanun, V., White, N. J. & Sirisinha, S. Lipopolysaccharide from nonvirulent Ara+ Burkholderia pseudomallei isolates is immunologically indistinguishable from lipopolysaccharide from virulent Ara– clinical isolates. Clin. Diagn. Lab. Immunol. 5, 225–229 (1998).
Anuntagool, N., Panichakul, T., Aramsri, P. & Sirisinha, S. Shedding of lipopolysaccharide and 200-kDa surface antigen during the in vitro growth of virulent Ara− and avirulent Ara+ Burkholderia pseudomallei. Acta Trop. 74, 221–228 (2000).
Chua, K. L., Chan, Y. Y. & Gan, Y. H. Flagella are virulence determinants of Burkholderia pseudomallei. Infect. Immun. 71, 1622–1629 (2003).
DeShazer, D., Brett, P. J., Carlyon, R. & Woods, D. E. Mutagenesis of Burkholderia pseudomallei with Tn5-OT182: isolation of motility mutants and molecular characterization of the flagellin structural gene. J. Bacteriol. 179, 2116–2125 (1997).
Essex-Lopresti, A. E. et al. A type IV pilin, PilA, contributes to adherence of Burkholderia pseudomallei and virulence in vivo. Infect. Immun. 73, 1260–1264 (2005).
Yang, H., Kooi, C. D. & Sokol, P. A. Ability of Pseudomonas pseudomallei malleobactin to acquire transferrin-bound, lactoferrin-bound, and cell-derived iron. Infect. Immun. 61, 656–662 (1993).
Ashdown, L. R. & Koehler, J. M. Production of hemolysin and other extracellular enzymes by clinical isolates of Pseudomonas pseudomallei. J. Clin. Microbiol. 28, 2331–2334 (1990).
Moore, R. A. et al. Contribution of gene loss to the pathogenic evolution of Burkholderia pseudomallei and Burkholderia mallei. Infect. Immun. 72, 4172–4187 (2004). Suggests that gene loss during the evolution of B. pseudomallei might have contributed to virulence.
Takeda, K., Kaisho, T. & Akira, S. Toll-like receptors. Annu. Rev. Immunol. 21, 335–376 (2003).
Santanirand, P., Harley, V. S., Dance, D. A., Drasar, B. S. & Bancroft, G. J. Obligatory role of γ interferon for host survival in a murine model of infection with Burkholderia pseudomallei. Infect. Immun. 67, 3593–3600 (1999). Highlights the essential role of IFN-γ in adequate host defence in melioidosis.
Haque, A. et al. Role of T cells in innate and adaptive immunity against murine Burkholderia pseudomallei infection. J. Infect. Dis. 193, 370–379 (2006).
Ekchariyawat, P. et al. Burkholderia pseudomallei-induced expression of suppressor of cytokine signaling 3 and cytokine-inducible Src homology 2-containing protein in mouse macrophages: a possible mechanism for suppression of the response to γ interferon stimulation. Infect. Immun. 73, 7332–7339 (2005).
Hoppe, I. et al. Characterization of a murine model of melioidosis: comparison of different strains of mice. Infect. Immun. 67, 2891–2900 (1999).
Leakey, A. K., Ulett, G. C. & Hirst, R. G. BALB/c and C57Bl/6 mice infected with virulent Burkholderia pseudomallei provide contrasting animal models for the acute and chronic forms of human melioidosis. Microb. Pathog. 24, 269–275 (1998).
Lauw, F. N. et al. Elevated plasma concentrations of interferon (IFN)-γ and the IFN-γ-inducing cytokines interleukin (IL)-18, IL-12, and IL-15 in severe melioidosis. J. Infect. Dis. 180, 1878–1885 (1999).
Brown, A. E. et al. Immune cell activation in melioidosis: increased serum levels of interferon-γ and soluble interleukin-2 receptors without change in soluble CD8 protein. J. Infect. Dis. 163, 1145–1148 (1991).
Nuntayanuwat, S., Dharakul, T., Chaowagul, W. & Songsivilai, S. Polymorphism in the promoter region of tumor necrosis factor-α gene is associated with severe meliodosis. Hum. Immunol. 60, 979–983 (1999).
Lauw, F. N. et al. The CXC chemokines γ interferon (IFN-γ)-inducible protein 10 and monokine induced by IFN-γ are released during severe melioidosis. Infect. Immun. 68, 3888–3893 (2000).
Simpson, A. J. et al. Prognostic value of cytokine concentrations (tumor necrosis factor-α, interleukin-6, and interleukin-10) and clinical parameters in severe melioidosis. J. Infect. Dis. 181, 621–625 (2000).
Lauw, F. N. et al. Soluble granzymes are released during human endotoxemia and in patients with severe infection due to Gram-negative bacteria. J. Infect. Dis. 182, 206–213 (2000).
Lertmemongkolchai, G., Cai, G., Hunter, C. A. & Bancroft, G. J. Bystander activation of CD8+ T cells contributes to the rapid production of IFN-γ in response to bacterial pathogens. J. Immunol. 166, 1097–1105 (2001).
Egan, A. M. & Gordon, D. L. Burkholderia pseudomallei activates complement and is ingested but not killed by polymorphonuclear leukocytes. Infect. Immun. 64, 4952–4959 (1996).
Frank, M. M., Joiner, K. & Hammer, C. The function of antibody and complement in the lysis of bacteria. Rev. Infect. Dis. 9 (Suppl. 5), S537–S545 (1987).
Dharakul, T. et al. HLA-DR and-DQ associations with melioidosis. Hum. Immunol. 59, 580–586 (1998).
Ketheesan, N. et al. Demonstration of a cell-mediated immune response in melioidosis. J. Infect. Dis. 186, 286–289 (2002).
Barnes, J. L. et al. Adaptive immunity in melioidosis: a possible role for T cells in determining outcome of infection with Burkholderia pseudomallei. Clin. Immunol. 113, 22–28 (2004).
Utaisincharoen, P. et al. Induction of iNOS expression and antimicrobial activity by interferon (IFN)-β is distinct from IFN-γ in Burkholderia pseudomallei-infected mouse macrophages. Clin. Exp. Immunol. 136, 277–283 (2004).
Chierakul, W. et al. Disease severity and outcome of melioidosis in HIV coinfected individuals. Am. J. Trop. Med. Hyg. 73, 1165–1166 (2005).
Jones, A. L., Beveridge, T. J. & Woods, D. E. Intracellular survival of Burkholderia pseudomallei. Infect. Immun. 64, 782–790 (1996).
Pruksachartvuthi S, A. N. & Thankerngpol, K. Survival of Pseudomonas pseudomallei in human phagocytes. J. Med. Microbiol. 31, 109–114 (1990).
Egan, A. M. & Gordon, D. L. Burkholderia pseudomallei activates complement and is ingested but not killed by polymorphonuclear leukocytes. Infect. Immun. 64, 4952–4959 (1996).
Kespichayawattana, W., Rattanachetkul, S., Wanun, T., Utaisincharoen, P. & Sirisinha, S. Burkholderia pseudomallei induces cell fusion and actin-associated membrane protrusion: a possible mechanism for cell-to-cell spreading. Infect. Immun. 68, 5377–5384 (2000).
Mohamed, R., Nathan, S., Embi, N., Razak, N. & Ismail, G. Inhibition of macromolecular synthesis in cultured macrophages by Pseudomonas pseudomallei exotoxin. Microbiol. Immunol. 33, 811–820 (1989).
Dejsirilert, S., Kondo, E., Chiewsilp, D. & Kanai, K. Growth and survival of Pseudomonas pseudomallei in acidic environments. Jpn J. Med. Sci. Biol. 44, 63–74 (1991).
Utaisincharoen, P., Tangthawornchaikul, N., Kespichayawattana, W., Chaisuriya, P. & Sirisinha, S. Burkholderia pseudomallei interferes with inducible nitric oxide synthase (iNOS) production: a possible mechanism of evading macrophage killing. Microbiol. Immunol. 45, 307–313 (2001).
Harley, V. S., Dance, D. A., Drasar, B. S. & Tovey, G. Effects of Burkholderia pseudomallei and other Burkholderia species on eukaryotic cells in tissue culture. Microbios 96, 71–93 (1998).
Harley, V. S., Dance, D. A., Tovey, G., McCrossan, M. V. & Drasar, B. S. An ultrastructural study of the phagocytosis of Burkholderia pseudomallei. Microbios 94, 35–45 (1998).
Breitbach, K. et al. Actin-based motility of Burkholderia pseudomallei involves the Arp 2/3 complex, but not N-WASP and Ena/VASP proteins. Cell. Microbiol. 5, 385–393 (2003). Excellent study that added to our understanding of the intracellular lifestyle of B. pseudomallei.
Stevens, M. P. et al. Identification of a bacterial factor required for actin-based motility of Burkholderia pseudomallei. Mol. Microbiol. 56, 40–53 (2005). Further dissects the role of protein–protein interaction leading to the initiation of actin polymerization by B. pseudomallei.
Wong, K. T., Puthucheary, S. D. & Vadivelu, J. The histopathology of human melioidosis. Histopathology 26, 51–55 (1995).
Sun, G. W., Lu, J., Pervaiz, S., Cao, W. P. & Gan, Y. H. Caspase-1 dependent macrophage death induced by Burkholderia pseudomallei. Cell. Microbiol. 7, 1447–1458 (2005).
Inglis, T. J. et al. Interaction between Burkholderia pseudomallei and Acanthamoeba species results in coiling phagocytosis, endamebic bacterial survival, and escape. Infect. Immun. 68, 1681–1686 (2000).
Stevens, M. P. & Galyov, E. E. Exploitation of host cells by Burkholderia pseudomallei. Int. J. Med. Microbiol. 293, 549–555 (2004).
Brown, N. F., Boddey, J. A., Flegg, C. P. & Beacham, I. R. Adherence of Burkholderia pseudomallei cells to cultured human epithelial cell lines is regulated by growth temperature. Infect. Immun. 70, 974–980 (2002).
Kespichayawattana, W., Intachote, P., Utaisincharoen, P. & Sirisinha, S. Virulent Burkholderia pseudomallei is more efficient than avirulent Burkholderia thailandensis in invasion of and adherence to cultured human epithelial cells. Microb. Pathog. 36, 287–292 (2004).
Cheng, A. C., Stephens, D. P., Anstey, N. M. & Currie, B. J. Adjunctive granulocyte colony-stimulating factor for treatment of septic shock due to melioidosis. Clin. Infect. Dis. 38, 32–37 (2004). A retrospective study showing a fall in mortality of septic melioidosis with adjunctive G-CSF treatment.
Powell, K., Ulett, G., Hirst, R. & Norton, R. G-CSF immunotherapy for treatment of acute disseminated murine melioidosis. FEMS Microbiol. Lett. 224, 315–318 (2003).
Cheng, A. C. Meliodosis: Epidemiology, Pathophysiology And Management. Ph.D. thesis, Flinders University of South Australia http://catalogue.flinders.edu.au/local/adt/uploads/approved/adt-SFU20051121.141305/public/03SectionB.pdf (2005).
Utaisincharoen, P. et al. CpG ODN enhances uptake of bacteria by mouse macrophages. Clin. Exp. Immunol. 132, 70–75 (2003).
Wongratanacheewin, S. et al. Immunostimulatory CpG oligodeoxynucleotide confers protection in a murine model of infection with Burkholderia pseudomallei. Infect. Immun. 72, 4494–4502 (2004). Evaluation of a potential new immunomodulatory treatment target in melioidosis.
Warawa, J. & Woods, D. E. Melioidosis vaccines. Expert Rev. Vaccines 1, 477–482 (2002).
Atkins, T. et al. A mutant of Burkholderia pseudomallei, auxotrophic in the branched chain amino acid biosynthetic pathway, is attenuated and protective in a murine model of melioidosis. Infect. Immun. 70, 5290–5294 (2002).
Brett, P. J., Mah, D. C. & Woods, D. E. Isolation and characterization of Pseudomonas pseudomallei flagellin proteins. Infect. Immun. 62, 1914–1919 (1994).
Nelson, M. et al. Evaluation of lipopolysaccharide and capsular polysaccharide as subunit vaccines against experimental melioidosis. J. Med. Microbiol. 53, 1177–1182 (2004).
Iliukhin, V. I. et al. Burkholderia thailandensis: biological properties, identification and taxonomy. Mol. Gen. Mikrobiol. Virusol. 7–11 (2002).
Healey, G. D., Elvin, S. J., Morton, M. & Williamson, E. D. Humoral and cell-mediated adaptive immune responses are required for protection against Burkholderia pseudomallei challenge and bacterial clearance postinfection. Infect. Immun. 73, 5945–5951 (2005).
Ou, K. et al. Integrative genomic, transcriptional, and proteomic diversity in natural isolates of the human pathogen Burkholderia pseudomallei. J. Bacteriol. 187, 4276–4285 (2005).
Mahenthiralingam, E., Urban, T. A. & Goldberg, J. B. The multifarious, multireplicon Burkholderia cepacia complex. Nature Rev. Microbiol. 3, 144–156 (2005).
Payne, G. W. et al. Development of a recA gene-based identification approach for the entire Burkholderia genus. Appl. Environ. Microbiol. 71, 3917–3927 (2005).
Acknowledgements
We thank members of the Wellcome Trust–Oxford University–Mahidol University Tropical Medicine Research Programme for their support, in particular V. Wuthiekanun and W. Chierakul. We thank the staff of Sappasithiprasong Hospital for many years of fruitful collaboration; special thanks go to W. Chaowagul. W.J.W. is supported by the Dutch Foundation for Tropical Research (WOTRO). S.J.P. is supported by a Wellcome Trust Career Development Award in Clinical Tropical Medicine.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
Salmonella enterica serovar Typhimurium
FURTHER INFORMATION
Wellcome Trust South-East Asia Programme in Thailand
B. pseudomallei genome sequence
The Institute for Genomic Research (TIGR)
CDC list of bioterrorism agents
BICHAT, the European Commission's Task Force on biological and chemical threats
Glossary
- Genomic islands
-
Clusters of genes that have been imported from unrelated bacterial taxa through horizontal gene transfer, and which might help the bacterium to acquire a new (possibly pathogenic) lifestyle.
- Shotgun sequencing
-
A genomic sequencing strategy that involves random fragmentation of large DNA segments. The fragments are sequenced, and programs with highly refined algorithms are used to reassemble the original DNA sequence.
- Multilocus sequence typing
-
(MLST). A method for the genotypic characterization of prokaryotes at the infraspecific level, using the allelic mismatches of a small number of housekeeping genes. Designed as a tool in molecular epidemiology and used for recognizing distinct strains within named species.
- Subtractive hybridization
-
A technique used to identify differentially expressed genes. The DNA species present in one sample are specifically enriched by hybridization with nucleic acids from another sample and by removing the associated double-stranded molecules.
- Quorum sensing
-
A system by which bacteria communicate. Signalling molecules — chemicals similar to pheromones that are produced by an individual bacterium — can affect the behaviour of surrounding bacteria.
- Complement
-
A part of the innate immune system comprising serum proteins that can protect against infection.
- Limulus amoebocytelysate assay
-
A chromogenic assay used to monitor endotoxin production.
- Natural killer (NK) cells
-
Lymphocytes that do not express the T-cell receptor or B-cell receptor and that mediate natural killing against prototypical NK-cell-sensitive targets.
- Human leukocyte antigen
-
(HLA). Also known as major histocompatibility complex (MHC), a glycoprotein that is found on the surface of antigen-presenting cells that presents antigen for recognition by TH cells.
Rights and permissions
About this article
Cite this article
Wiersinga, W., van der Poll, T., White, N. et al. Melioidosis: insights into the pathogenicity of Burkholderia pseudomallei. Nat Rev Microbiol 4, 272–282 (2006). https://doi.org/10.1038/nrmicro1385
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1385
This article is cited by
-
Impact of template denaturation prior to whole genome amplification on gene detection in high GC-content species, Burkholderia mallei and B. pseudomallei
BMC Research Notes (2024)
-
Development of a Novel Internally Controlled HrpB1 Gene-Based Real-Time qPCR Assay for Detection of Burkholderia pseudomallei
Molecular Diagnosis & Therapy (2024)
-
Nucleic Acid Amplification Free-QCM-DNA Biosensor for Burkholderia pseudomallei Detection
Current Microbiology (2023)
-
Paravertebral abscess and bloodstream infection caused by Burkholderia pseudomallei after acupuncture: a case report
BMC Complementary Medicine and Therapies (2022)
-
Molecular basis of specificity and deamidation of eIF4A by Burkholderia Lethal Factor 1
Communications Biology (2022)