Key Points
-
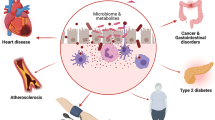
Microbial communities are associated with the human host, primarily in the intestinal tract, where they affect host metabolism and contribute to the pathogenesis of cardiovascular disease.
-
The susceptibility to atherosclerosis and thrombosis can be transmitted via gut microbial transplantation in mouse models.
-
Microbial-associated molecular patterns are sensed by host pattern recognition receptors and affect cardiovascular disease pathogenesis.
-
Microbial metabolism of common dietary nutrients produces both anti-atherogenic and pro-atherogenic metabolites that engage distinct host receptor systems and affect cardiovascular disease pathogenesis.
-
Plasma levels of the gut microbial metabolite trimethylamine-N-oxide are associated with incident development of cardiovascular disease and its adverse outcomes in humans.
-
Bacterially derived short-chain fatty acids (acetate, propionate and butyrate) can engage host receptor systems and potentially affect cardiovascular pathogenesis.
-
Bacterially derived secondary bile acids can modulate dietary fat absorption and signal through cell-surface and nuclear hormone receptors, potentially affecting cardiovascular disease pathogenesis.
-
Gut microorganism-targeted therapeutic strategies hold promise for the prevention and/or treatment of cardiovascular disease.
Abstract
Although diet has long been known to contribute to the pathogenesis of cardiovascular disease (CVD), research over the past decade has revealed an unexpected interplay between nutrient intake, gut microbial metabolism and the host to modify the risk of developing CVD. Microbial-associated molecular patterns are sensed by host pattern recognition receptors and have been suggested to drive CVD pathogenesis. In addition, the host microbiota produces various metabolites, such as trimethylamine-N-oxide, short-chain fatty acids and secondary bile acids, that affect CVD pathogenesis. These recent advances support the notion that targeting the interactions between the host and microorganisms may hold promise for the prevention or treatment of CVD. In this Review, we summarize our current knowledge of the gut microbial mechanisms that drive CVD, with special emphasis on therapeutic interventions, and we highlight the need to establish causal links between microbial pathways and CVD pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Mozaffarian, D. et al. Executive summary: heart disease and stroke statistics — 2016 update: a report from the American Heart Assocation. Circulation 133, 447–454 (2016).
Ardissino, D. et al. Influence of 9p21.3 genetic variants on clinical and angiographic outcomes in early-onset myocardial infarction. J. Am. Coll. Cardiol. 58, 426–434 (2011).
Ripatti, S. et al. A multilocus genetic risk score for coronary heart diease: case-control and prospective cohort analyses. Lancet 376, 1393–1400 (2010).
Yu, E. et al. Diet, lifestyle, biomarkers, genetic factors, and risk of cardiovascular disease in the Nurses' Health Study. Am. J. Publ. Health 106, 1616–1623 (2016).
Brown, J. M. & Hazen, S. L. The gut microbial endocrine organ: bacterially derived signals driving cardiometabolic diseases. Annu. Rev. Med. 66, 343–359 (2015).
Tang, W. H. & Hazen, S. L. The contributory role of gut microbiota in cardiovascular disease. J. Clin. Invest. 124, 4204–4211 (2014).
Gregory, J. C. et al. Transmission of atherosclerosis susceptibility with gut microbial transplantation. J. Biol. Chem. 290, 5647–5660 (2015). This study demonstrates for the first time that atherosclerosis susceptibility can be transmitted by gut microbial transplantation, fulfilling a key Koch postulate for microbial contribution to disease causation.
Zhu, W. et al. Gut microbial metabolite TMAO enhances platelet hyperreactivity and thrombosis risk. Cell 165, 111–124 (2016). This study shows the first cell autonomous effects of the gut microorganism-derived metabolite TMAO on platelet activation and thrombosis potential in vivo , that microbial transplantation transmits TMAO levels and thrombosis potential in vivo and that thrombotic event risk in subjects tracks with TMAO levels.
Kholy, K. E., Genco, R. J. & Van Dyke, T. E. Oral infections and cardiovascular disease. Trends Endocrinol. Metab. 26, 315–321 (2015).
Filardo, S. et al. Chlamydia pneumoniae-mediated inflammation in atherosclerosis: A meta-analysis. Mediators Inflamm. 2015, 378658 (2015).
Koren, O. et al. Human oral, gut, and plaque microbiota in patients with atherosclerosis. Proc. Natl Acad. Sci. USA 108 (Suppl. 1), 4592–4598 (2011). This study provides evidence that human atherosclerotic plaques contain bacteria.
Mitra, S. et al. In silico analyses of metagenomes from human atherosclerotic plaque samples. Microbiome 3, 38 (2015).
Caladrini, C. A. et al. Microbial composition of atherosclerotic plaques. Oral Dis. 20, e128–e134 (2014).
Karlsson, F. H. et al. Symptomatic atherosclerosis is associated with an altered gut metagenome. Nat. Commun. 3, 1245 (2012).
Saikku, P. et al. Chronic chlamydia pneumoniae infection as a risk factor for coronary heart disease in the Helsinki Heart Study. Ann. Intern. Med. 15, 273–278 (1992).
Kol, A. et al. Chlamydia and human heat shock protein 60s activate human vascular endothelium, smooth muscle cells, and macrophages. J. Clin. Invest. 103, 571–577 (1999).
Chen, S. et al. Chlamydia pneumoniae-induced foam cell formation requires MyD88-dependent and -independent signaling and is reciprocally modulated by liver X receptor activation. J. Immunol. 181, 7186–7193 (2008).
Grayston, J. T. et al. Azithromycin for the secondary prevention of coronary events. N. Engl. J. Med. 352, 1637–1645 (2005).
Cannon, C. P. et al. Antibiotic treatment of Chlamydia pneumoniae after acute coronary syndrome. N. Engl. J. Med. 352, 1646–1654 (2005).
O'Connor, C. M. et al. Azithromycin for the secondary prevention of coronary heart disease events — The WIZARD Study: a randomized controlled trial. JAMA 290, 1459–1466 (2003).
Leung, C., Rivera, L., Furness, J. B. & Angus, P. W. The role of gut microbiota in NAFLD. Nat. Rev. Gastroenterol. Hepatol. 13, 412–425 (2016).
Munford, R. S. Endotoxemia — menace, marker, or mistake? J. Leukoc. Biol. 100, 687–698 (2016).
Caligiuri, G. et al. Chlamydia pneumoniae infection does not induce or modify atherosclerosis in mice. Circulation 103, 2834–2838 (2001).
Hussain, M., Stover, C. M. & Dupont, A. P. gingivalis in periodontal disease and atherosclerosis — scenes of action for antimicrobial peptides and complement. Front. Immunol. 6, 45 (2015).
Giacona, M. B. et al. Porphyromonas gingivalis induces its uptake by human macrophages and promotes foam cell formation in vitro. FEMS Microbiol. Lett. 241, 95–101 (2004).
Naito, M. et al. Porphyromonas gingivalis-induced platelet aggregation in plasma depends on Hgp44 adhesion but not Rgp proteinase. Mol. Microbiol. 59, 152–167 (2006).
Li, L. et al. Porphyromonas gingivalis infection accelerates the progression of atherosclerosis in a heterozygous apolipoprotein E-deficient murine model. Circulation 105, 861–867 (2002).
Brodala, N. et al. Porphyromonas gingivalis bacteremia induces coronary and aortic atherosclerosis in normocholesterolemic and hypercholesterolemic pigs. Arterioscler. Thromb. Vasc. Biol. 25, 1446–1451 (2005).
Lalla, E. et al. Oral infection with a periodontal pathogen accelerates early atherosclerosis in apolipoprotein E-null mice. Arterioscler. Thromb. Vasc. Biol. 23, 1405–1411 (2003).
Mach, F. et al. Influence of Helicobacter pylori infection during atherogenesis in vivo in mice. Circ. Res. 90, E1–E4 (2002).
Chen, X. H. et al. Helicobacter pylori infection enhances atherosclerosis in high-cholesterol diet fed C57BL/6 mice [Chinese]. Zhonghua Xin Xue Guan Bing Za Zhi 38, 259–263 (2010).
Liuba, P. et al. Co-infection with Clamydia pneumoniae and Helicobacter pylori results in vascular endothelial dysfunction and enhanced VCAM-1 expression in apoE-knockout mice. J. Vasc. Res. 40, 115–122 (2003).
El Moktari, N. E., Ott, S. J., Nebel, A., Simon, R. & Schreiber, S. A. functional variant in the CARD4 gene and risk of premature coronary heart disease. Int. J. Immunogenet. 33, 307–311 (2006).
Kanno, S. et al. Activation of an innate immune receptor, Nod1, accelerates atherogenesis in Apoe-/- mice. J. Immunol. 194, 773–780 (2015).
Schenk, M., Belisle, J. T. & Modlin, R. L. TLR2 looks at lipoproteins. Immunity 31, 847–849 (2009).
Mullick, A. E., Tobias, P. S. & Curtiss, L. K. Modulation of atherosclerosis in mice by Toll-like receptor 2. J. Clin. Invest. 115, 3149–3156 (2005).
Sale, M. L. et al. Toll-like receptor 6 Ser249Pro polymorphism is associated with lower left ventricular wall thickness and inflammatory response in hypertensive women. Am. J. Hypertens. 23, 649–654 (2010).
Hamann, L. et al. Association of a common TLR-6 polymorphism with coronary artery disease — implications for healthy ageing? Immun. Ageing 10, 43 (2013).
Lundberg, A. M. et al. Toll-like receptor 3 and 4 signaling through TRIF and TRAM adaptors in haematopoetic cells promotes atherosclerosis. Cardiovasc. Res. 99, 364–373 (2013).
Ballistreri, C. R. et al. TLR4 polymorphisms and ageing: implications for the pathophysiology of age-related diseases. J. Clin. Immunol. 29, 406–415 (2009).
Michelsen, K. S. et al. Lack of toll-like receptor 4 or myeloid differentiation factor 88 reduces atherosclerosis and alters plaque phenotype in mice deficient in apolipoprotein E. Proc. Natl Acad. Sci. USA 101, 10679–10684 (2004).
Stewart, C. R. et al. CD36 ligands promote sterile inflammation through assembly of a Toll-like receptor 4 and 6 heterodimer. Nat. Immunol. 11, 155–161 (2010).
Ding, Y. et al. Toll-like receptor 4 deficiency decreases atherosclerosis but does not protect against inflammation in obese low-density lipoprotein receptor-deficient mice. Arterioscler. Thromb. Vasc. Biol. 32, 1596–1604 (2012).
Kanellakis, P. et al. High-mobility group box protein 1 neutralization reduces development of diet-induced atherosclerosis in apolipoprotein e-deficient mice. Arterioscler. Thromb. Vasc. Biol. 31, 313–319 (2011).
Salagianni, M. et al. Toll-like receptor 7 protects from atherosclerosis by constraining “inflammatory” macrophage activation. Circulation 126, 952–962 (2012).
Tang, W. H. et al. Intestinal microbial metabolism of phosphatidylcholine and cardiovascular risk. N. Engl. J. Med. 368, 1575–1578 (2013).
Koulis, C. et al. Protective role for Toll-like receptor-9 in the development of atherosclerosis in apolipoprotein E-deficient mice. Arterioscler. Thromb. Vasc. Biol. 34, 516–525 (2014).
Lee, W. J. & Hase, K. Gut microbiota-generated metabolites in animal health and disease. Nat. Chem. Biol. 10, 416–424 (2014).
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Spanogiannopoulos, P. et al. The microbial pharmacists within us: a metagenomic view of xenobiotic metabolism. Nat. Rev. Microbiol. 14, 273–287 (2016).
Brown, J. M. & Hazen, S. L. Targeting of microbe-derived metabolites to improve human health: the next frontier for drug discovery. J. Biol. Chem. 292, 8560–8568 (2017).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–63 (2011). This paper is the first to directly demonstrate a pathogenic role for a gut microorganism-dependent metabolite, TMAO, in CVD pathogenesis with the use of both mouse models of disease and large-scale clinical studies.
Koeth, R. A. et al. Intestinal microbiota metabolism of L-carnitine, a nutrient in red meat, promotes atherosclerosis. Nat. Med. 19, 576–585 (2013). This study demonstrates a nutrient (carnitine)–gut microbial host metaorganismal pathway (the TMAO pathway), providing a mechanistic link between red meat consumption and CVD pathogenesis.
Wang, Z. et al. Prognostic value of choline and betaine depends on intestinal microbiota-generated metabolite trimethylamine-N-oxide. Eur. Heart J. 35, 904–910 (2014).
Koeth, R. A. et al. γ-Butyrobetaine is a proatherogenic intermediate in gut microbial metabolism of L-carnitine to TMAO. Cell Metab. 20, 799–812 (2014).
Senthong, V. et al. Plasma trimethylamine N-oxide, a gut microbe-generated phosphatidylcholine metabolite, is associated with atherosclerotic burden. J. Am. Coll. Cardiol. 67, 2620–2628 (2016).
Senthong, V. et al. Intestinal microbiota-generated metabolite trimethylamine N-oxide and 5-year mortality risk in stable coronary artery disease: the contributory role of intestinal microbiota in a COURAGE-like patient cohort. J. Am. Heart Assoc. 5, e002816 (2016).
Wang, Z. et al. Non-lethal inhibition of gut microbial trimethylamine production for the treatment of atherosclerosis. Cell 163, 1585–1595 (2015). This paper is the first to show that a nonlethal small-molecule inhibitor of a gut microbial enzyme (TMA lyase) can protect mice against diet-enhanced atherosclerosis development.
Zheng, Y. et al. Dietary phosphatidylcholine and risk of all-cause and cardiovascular-specific mortality among US women and men. Am. J. Clin. Nutr. 104, 173–180 (2016).
Randrianarisoa, E. et al. Relationship of serum trimethylamine N-oxide (TMAO) levels with early atherosclerosis in humans. Sci. Rep. 6, 26745 (2016).
Kim, R. B. et al. Advanced chronic kidney disease populations have elevated trimethylamine N-oxide levels associated with increased cardiovascular events. Kidney Int. 89, 1144–1152 (2016).
Skagen, K. et al. The carnitine-butyrobetaine-trimethylamine-oxide pathway and its association with cardiovascular mortality in patients with carotid atherosclerosis. Atherosclerosis 247, 64–69 (2016).
Missailidis, C. et al. Serum trimethylamine-N-oxide is strongly related to renal function and predicts outcome in chronic kidney disease. PLOS One 11, e0141738 (2016).
Mafune, A. et al. Associations among serum trimethylamine-N-oxide (TMAO) levels, kidney function and infarcted coronary artery number in patients undergoing cardiovascular surgery: a cross-sectional study. Clin. Exp. Nephrol. 20, 731–739 (2016).
Zhu, W., Wang, Z., Tang, W. H. & Hazen, S. L. Gut microbe-generated trimethylamine N-oxide from dietary choline is prothrombotic in subjects. Circulation 135, 1671–1673 (2017).
Troseid, M. et al. Microbiota-dependent metabolite trimethylamine-N-oxide is associated with disease severity and survival of patients with chronic heart failure. J. Intern. Med. 277, 717–726 (2015).
Organ, C. L. et al. Choline diet and its gut microbe-derive metabolite, Trimethylamine N-oxide, exacerbate pressure overload-induced heart failure. Circ. Heart Fail. 9, e002314 (2016). This paper demonstrates a role for the gut microbial metabolite TMAO in pressure-overload-induced heart failure.
Rhee, E. P. et al. A combined epidemiologic and metabolomics approach improves CKD prediction. J. Am. Soc. Nephrol. 24, 1330–1338 (2013).
Bell, J. D. et al. Nuclear magnetic resonance studies of blood plasma and urine from subjects with chronic renal failure: identification of trimethylamine-N-oxide. Biochim. Biophys. Acta 1096, 101–107 (1991).
Tang, W. H. et al. Gut microbiota-dependent trimethylamine N-oxide (TMAO) pathway contributes to both development of renal insufficiency and mortality risk in chronic kidney disease. Circ. Res. 116, 448–455 (2015). This paper demonstrates that the gut microbial metabolite TMAO can induce renal fibrosis and functional impairment and contribute to adverse CVD outcomes in subjects with chronic kidney disease.
Dambrova, M. et al. Diabetes is associated with higher trimethylamine N-oxide plasma levels. Exp. Clin. Endocrinol. Diabetes 124, 251–256 (2016).
Lever, M. et al. Betaine and trimethylamine-N-oxide as predictors of cardiovascular outcomes show different patterns in diabetes mellitus: an observational study. PLOS One 9, e114969 (2014).
Tang, W. H. et al. Increased trimethylamine N-oxide portends high mortality risk independent of glycemic control in patients with type 2 diabetes mellitus. Clin. Chem. 63, 297–306 (2017).
Miao, J. et al. Flavin-containing monooxygenase 3 as a potential player in diabetes-associated atherosclerosis. Nat. Commun. 6, 6498 (2015). Through unbiased metabolomics screening, this paper demonstrates for the first time a role for the TMAO-producing enzyme flavin-containing monooxygenase 3 in insulin action.
Schugar, R. C. et al. The TMAO-producing enzyme flavin-containing monooxygenase 3 regulates obesity and the beiging of white adipose tissue. Cell Rep. 19, 2451–2461 (2017). This paper is the first to show that pharmacologic inhibition or genetic deletion of FMO3 protects mice from high-fat-diet-induced obesity.
Craciun, S. & Balskus, E. P. Microbial conversion of choline to trimethylamine requires a glycyl radical enzyme. Proc. Natl Acad. Sci. USA 109, 21307–21312 (2012).
Zhu, Y. et al. Carnitine metabolism to trimethylamine by an usual Rieske-type oxygenase from human microbiota. Proc. Natl Acad. Sci. USA 111, 4268–4273 (2014).
Yancey, P. H. et al. Living with water stress: evolution of osmolyte systems. Science 217, 1214–1222 (1982).
Lin, T. Y. & Timasheff, S. N. Why do some organisms use a urea-methylamine mixture as osmolyte? Thermodynamic compensation of urea and trimethylamine-N-oxide interactions with protein. Biochemistry 33, 12695–12701 (1994).
Seldin, M. M. et al. Trimethylamine N-oxide promotes vascular inflammation through signaling of mitogen-activated protein kinase and nuclear factor kB. J. Am. Heart Assoc. 5, e002767 (2016).
Li, Q. et al. Synchronous evolution of an odor biosynthesis pathway and behavioral response. Curr. Biol. 23, 11–20 (2013). This paper demonstrates that the gut microbial metabolite TMA can activate a G-protein-coupled receptor to regulate behavioural responses.
Wallrabenstein, I. et al. Human trace amine-associated receptor TAAR5 can be activated by trimethylamine. PLOS One 8, e54950 (2013).
Falony, G., Vleira-Silva, S. & Raes, J. Microbiology meets big data: The case of gut microbe-derived trimethylamine. Annu. Rev. Microbiol. 69, 305–321 (2015). This innovative paper describes the power of mining reference genomes for prediction of bacterial enzymatic activity.
Romano, K. A., Vivas, E. I., Amador-Noguez, D. & Rey, F. E. Instestinal microbiota composition modulates choline bioavailability from diet and accumulation of the proatherogenic metabolite trimethylamine-N-oxide. mBio 6, e02481 (2015). This paper identifies human commensal bacteria that possess TMA lyase activity and demonstrates that transplantation of these strains can alter host choline homeostasis.
Martinez- del Campo, A. et al. Characterization and detection of a widely distributed gene cluster that predicts anaerobic choline utilization by human gut bacteria. mBio 6, e00042–e00015 (2015).
Levin, B. J. et al. A prominent glycyl radical enzyme in human gut microbiomes metabolizes trans-4-hydroxy-l-proline. Science 355, eaai8386 (2017).
Rath, S. et al. Uncovering the trimethylamine-producing bacteria of the human gut microbiota. Microbiome 5, 54 (2017).
Mejean, V. et al. TMAO anaerobic respiration in Escherichia coli: involvement of the tor operon. Mol. Microbiol. 11, 1169–1179 (1994).
Lidbury, I., Murrell, J. C. & Chen, Y. Trimethylamine N-oxide metabolism by abundant marine heterotrophic bacteria. Proc. Natl Acad. Sci. USA 111, 2710–2715 (2014).
Lidbury, I. D., Murrell, J. C. & Chen, Y. Trimethylamine and trimethylamine N-oxide are supplementary energy sources for marine carbon and nitrogen cycling. ISME J. 9, 760–769 (2015).
Li, C. Y. et al. Mechanistic insight into trimethylamine N-oxide recognition by the marine bacterium Ruegeria pomeroyi DSS-3. J. Bacteriol. 197, 3378–3387 (2015).
Brugere, J. F. et al. Archaebiotics: proposed therapeutic use of archaea to prevent trimethylaminuria and cardiovascular disease. Gut Microbes 5, 5–10 (2014).
Collins, H. L. et al. L-Carnitine intake and high trimethylamine N-oxide levels correlate with low aortic lesions in ApoE(-/-) transgenic mice expressing CETP. Atherosclerosis 244, 29–37 (2016).
Nagata, C. et al. Choline and betaine intakes are not associated with cardiovascular disease mortality risk in Japanese men and women. J. Nutr. 145, 1787–1792 (2015).
Bidulescu, A. et al. Usual choline and betaine dietary intake and incident coronary heart disease: the Atherosclerosis Risk in Communities (ARIC) study. BMC Cardiovasc. Disord. 7, 20 (2007).
Dalmeijer, G. W. et al. Prospective study on dietary intakes of folate, betaine, and choline and cardiovascular disease risk in women. Eur. J. Clin. Nutr. 62, 386–394 (2008).
Heianza, Y. et al. Gut microbiota metabolites and risk of major adverse cardiovascular events and death: a systematic review and meta-analysis of prospective studies. J. Am. Heart Assoc. 6, e004947.
Mueller, D. M. et al. Plasma levels of trimethylamine-N-oxide are confounded by impaired kidney function and poor metabolic control. Atherosclerosis 243, 638–644 (2015).
Yin, J. et al. Dysbiosis of gut microbiota with reduced trimethylamine-N-oxide level in patients with large-artery atherosclerotic stroke or transient ischemic attack. J. Am. Heart Assoc. 4, e002699.
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Canfora, E. E., Jocken, J. W. & Blaak, E. E. Short-chain fatty acids in control of body weight and insulin sensitivity. Nat. Rev. Endocrinol. 11, 577–591 (2015).
Ramezani, A. et al. Role of the gut microbiome in uremia: a potential therapeutic target. Am. J. Kidney Dis. 67, 483–498 (2016).
Miyamoto, J. et al. The role of short-chain fatty acid on blood pressure regulation. Curr. Opin. Nephrol. Hypertens. 25, 379–383 (2016).
Rey, F. E. et al. Dissecting the in vivo metabolic potential of two human gut acetogens. J. Biol. Chem. 285, 22082–22090 (2010).
Scott, K. P., Martin, J. C., Campbell, G., Mayer, C.-D. & Flint, H. J. Whole-genome transcription profiling reveals genes up-regulated by growth on fucose in the human gut bacterium “Roseburia inulinivorans”. J. Bacteriol. 188, 4340–4349 (2006).
Duncan, S. H., Barcenilla, A., Stewart, C. S., Pryde, S. E. & Flint, H. J. Acetate utilization and butyryl coenzyme A (CoA):acetate-CoA transferase in butyrate-producing bacteria from the human large intestine. Appl. Environ. Microbiol. 68, 5186–5190 (2002).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Brown, A. J. et al. The orphan G protein-coupled receptors GPR41 and GPR43 are activated by propionate and other short chain carboxylic acids. J. Biol. Chem. 278, 11312–11319 (2003).
Thangaraju, M. et al. GPR109A is a G-protein-coupled receptor for the bacterial fermentation product butyrate and functions as a tumor suppressor in colon. Cancer Res. 69, 2826–2832 (2009).
Pluznick, J. L. et al. Olfactory receptor responding to gut microbiota-derived signals plays a role in renin secretion and blood pressure regulation. Proc. Natl Acad. Sci. USA 110, 4410–4415 (2013).
Wolever, T., Spadafora, P. & Eshuis, H. Interaction between colonic acetate and propionate in humans. Am. J. Clin. Nutr. 53, 681–687 (1991).
Laurent, C. et al. Effect of acetate and propionate on fasting hepatic glucose production in humans. Eur. J. Clin. Nutr. 49, 484–491 (1995).
Fernandes, J., Vogt, J. & Wolever, T. M. Intravenous acetate elicits a greater free fatty acid rebound in normal than hyperinsulinemic humans. Eur. J. Clin. Nutr. 66, 1029–1034 (2012).
Venter, C. S., Vorster, H. H. & Cummings, J. H. Effects of dietary propionate on carbohydrate and lipid metabolism in healthy volunteers. Am. J. Gastroenterol. 85, 549–553 (1990).
Robertson, M. D., Bickerton, A. S., Dennis, A. L., Vidal, H. & Frayn, K. N. Insulin-sensitizing effects of dietary resistant starch and effects on skeletal muscle and adipose tissue metabolism. Am. J. Clin. Nutr. 82, 559–567 (2005).
Cani, P. D. et al. Gut microbiota fermentation of prebiotics increases satieogenic and incretin gut peptide production with consequences for appetite sensation and glucose response after a meal. Am. J. Clin. Nutr. 90, 1236–1243 (2009).
Parnell, J. A. & Reimer, R. A. Weight loss during oligofructose supplementation is associated with decreased ghrelin and increased peptide YY in overweight and obese adults. Am. J. Clin. Nutr. 89, 1751–1759 (2009).
Dewulf, E. et al. Insight into the prebiotic concept: lessons from an exploratory, double blind intervention study with inulin-type fructans in obese women. Gut 62, 1112–1121 (2013).
Daud, N. M. et al. The impact of oligofructose on stimulation of gut hormones, appetite regulation and adiposity. Obesity 22, 1430–1438 (2014).
Vulvic, J., Juric, A., Tzortzis, G. & Gibson, G. R. A mixture of trans-galactooligosaccharides reduces markers of metabolic syndrome and modulates the fecal microbiota and immune function of overweight adults. J. Nutr. 143, 324–331 (2013).
Pedersen, C. et al. Gut hormone release and appetite regulation in healthy non-obese participants following oligofructose intake. A dose escalation study. Appetite 66, 44–53 (2013).
Rejinders, D. et al. Effects of gut microbiota manipulation by antibiotics on host metabolism in obese humans: a randomized double-blind placebo-controlled trial. Cell Metab. 24, 341 (2016). This study associates large alterations in short-chain fatty acid levels with insulin sensitivity and energy metabolism in humans.
Durgan, D. J. et al. Role of the gut microbiome in obstructive sleep apnea-induced hypertension. Hypertension 67, 469–474 (2016).
Cai, T. Q. et al. Role of GPR81 in lactate-mediated reduction of adipose lipolysis. Biophys. Res. Commun. 377, 987–991 (2008).
Shen, Z. et al. Inhibition of G protein-coupled receptor 81 (GPR81) protects against ischemic brain injury. CNS Neurosci. Ther. 21, 271–279 (2015).
He, W. et al. Citric acid cycle intermediates as ligands for ophan G-protein-coupled receptors. Nature 429, 188–193 (2004).
Rubic, T. et al. Triggering the succinate receptor GPR91 on dendritic cells enhances immunity. Nat. Immunol. 9, 1261–1269 (2008).
De Vadder, F. et al. Microbiota-produced succinate improves glucose homeostasis via intestinal gluconeogenesis. Cell Metab. 24, 151–157 (2016).
Lukasova, M. et al. Nicotinic acid inhibits progression of atherosclerosis in mice through its receptor GPR109A expressed in immune cells. J. Clin. Invest. 121, 1163–1173 (2011).
Russell, D. W. Fifty years of advances in bile acid synthesis and metabolism. J. Lipid Res. 50, S120–S125 (2009).
Hylemon, P. B. et al. Bile acids as regulatory molecules. J. Lipid Res. 50, 1509–1520 (2009).
Kuipers, F., Bloks, V. W. & Groen, A. K. Beyond intestinal soap-bile acids in metabolic control. Nat. Rev. Endocrinol. 10, 488–498 (2014).
Dawson, P. A. & Karpen, S. J. Intestinal transport & metabolism of bile acids. J. Lipid Res. 56, 1085–1099 (2015).
Ridlon, J. M., Kang, D. J., Hylemon, P. B. & Bajaj, J. S. Bile acids and the gut microbiome. Curr. Opin. Gastroenterol. 30, 332–338 (2014).
Wahlstrom, A., Sayin, S. I., Marschall, H. U. & Backhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Klaassen, C. D. & Cui, J. Y. Review: mechanisms of how the intestinal microbiota alters the effects of drugs and bile acids. Drug Metab. Dispos. 43, 1505–1521 (2015).
Ridlon, J. M., Harris, S. C., Bhowmik, S., Kang, D. J. & Hylemon, P. E. Consequences of bile salt biotransformations by intestinal bacteria. Gut Microbes 7, 22–39 (2016).
Hofmann, A. F., Hagey, L. R. & Krasowski, M. D. Bile salts of vertebrates: structural variation and possible evolutionary significance. J. Lipid Res. 51, 226–246 (2010).
Joyce, S. A. & Gahan, C. G. Disease-associated changes in bile acid profiles and links to altered gut microbiota. Dig. Dis. 35, 169–177 (2017).
Lin, J., Sahin, O., Michael, L. O. & Zhang, Q. Critical role of multidrug efflux pump CmeABC in bile resistance and in vivo colonization of Camphylobacter jejuni. Infect. Immunol. 71, 4250–4259 (2003).
Yokota, A., Veenstra, M., Kurdi, P., van Veen, H. W. & Konings, W. N. Cholate resistance in Lactococcus lactis is mediated by an ATP-dependent multispecific organic anion transporter. J. Bacteriol. 182, 5196–5201 (2000).
Fernandez Murga, M. L., Bernick, D., de Valdez, G. F. & Disalvo, A. E. Permeability and stability properties of membranes formed by lipids extracted from Lactobacillus acidophilus grown at different temperatures. Arch. Biochem. Biophys. 364, 115–121 (1999).
Kimoto, H. Ohmono, S. & Okamoto, T. Enhancement of bile tolerance in Lactococci by Tween 80. J. Appl. Micro. 92, 41–46 (2002).
Liu, Y. et al. Functional role of tlyC1 encoding a hemolysin-like protein from Bifidobacterium longum BBMN68 in bile tolerance. FEMS Microbiol. Lett 360, 167–173 (2014).
Ruiz, L. et al. The cell-envelope proteome of Bifidobacterium longum in an in vivo bile environment. Microbiology 155, 957–967 (2009).
Schubert, R., Jaroni, H., Schoelmerich, J. & Schmidt, K. H. Studies on the mechanism of bile salt-induced liposomal membrane damage. Digestion 28, 181–190 (1983).
Prouty, A. M., Schwesinger, W. H. & Gunn, J. S. Biofilm formation and interaction with surfaces of gallstones by Salmonella spp. Infect. Immunol. 70, 2640–2649 (2002).
Lepercq, P. et al. Epimerization of chenodeoxycholic acid to ursodeoxycholic acid by Clostridium baratii isolated from human feces. FEMS Microbiol. Lett 235, 65–72 (2004).
Lepercq, P. et al. Isolates from normal human intestinal flora but not lactic acid bacteria exhibit 7α- and 7β-hydroxysteroid dehydrogenase activities. Microb. Ecol. Health Dis. 16, 195–201 (2004).
Kelsey, M. I., Molina, J. E., Huang, S. K., Hwang, K. K. The identification of microbial metabolites of sulfolithocholic acid. J. Lipid Res 21, 751–756 (1980).
Benson, G. M. et al. Polydeoxycholate in human and hamster feces: a major product of cholate metabolism. J. Lipid Res. 34, 2121–2134 (1993).
Tazuke, Y., Matsuda, K., Adachi, K. & Tsukada, Y. Purification and properties of a novel sulfatase from Pseudomonas testosteroni that hydrolyzed 3 β-hydroxy-5-cholenoic acid 3-sulfate. Biosci. Biotechnol. Biochem. 62, 1739–1744 (1998).
Makishima, M. et al. Identification of a nuclear receptor for bile acids. Science 284, 1362–1365 (1999). This is a landmark paper identifying the first-ever nuclear receptor for bile acids.
Heni, M. et al. Genetic variation in NR1H4 encoding the bile acid receptor FXR determines fasting glucose and free fatty acid levels in humans. J. Clin. Endocrinol. Metab. 98, E1224–E1229 (2013).
Zhang, Y. et al. FXR deficiency causes reduced atherosclerosis in Ldlr-/- mice. Arterioscler. Thromb. Vasc. Biol. 26, 2316–2321 (2006).
Wahlström, A., Kovatcheva-Datchary, P., Stahlman, M., Bäckhed, F. & Marschall, H. U. Crosstalk between bile acids and gut microbiota and its impact on farnesoid X receptor signaling. Dig. Dis. 35, 246–250 (2017).
Wahlström, A. et al. Induction of farnesoid X receptor signaling in germ-free mice colonized with a human microbiota. J. Lipid Res. 58, 412–419 (2017).
Watanabe, M. et al. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature 439, 484–489 (2006). This paper is the first to describe the role of the cell-surface bile acid receptor TGR5 in linking postprandial bile acid production to control of energy metabolism.
Pols, T. W. et al. TGR5 activation inhibits atherosclerosis by reducing macrophage inflammation and lipid loading. Cell Metab. 14, 747–757 (2011).
Staudinger, J. L. et al. The nuclear receptor PXR is a lithocholic acid sensor that protects against liver toxicity. Proc. Natl Acad. Sci. USA 98, 3369–3374 (2001).
Sui, Y., Xu, J., Rios-Pilier, J. & Zhou, C. Deficiency of PXR decreases atherosclerosis in apoE-deficient mice. J. Lipid Res. 52, 1652–1659 (2011).
Makishima, M. et al. Vitamin D receptor as an intestinal bile acid sensor. Science 296, 1313–1316 (2002).
Lu, S. et al. The associations between the polymorphisms of vitamin D receptor and coronary artery disease: a systematic review and meta-analysis. Medicine (Baltimore) 95, e3467 (2016).
Szeto, F. L. et al. Vitamin D receptor signaling inhibits atherosclerosis in mice. Mol. Endocrinol. 26, 1091–1101 (2012).
Raufman, J. P., Cheng, K. & Zimniak, P. Activation of muscarinic receptor signaling by bile acids: physiological and medical implications. Dig. Dis. Sci. 48, 1431–1444 (2003).
Hautala, A. J. et al. Acetylcholine receptor M2 gene variants, heart rate recovery, and risk of cardiac death after an acute myocardial infarction. Ann. Med. 41, 197–207 (2009).
Studer, E. et al. Conjugated bile acids activate the sphingosine-1-phosphate receptor 2 in primary rodent hepatocytes. Hepatology 55, 267–276 (2012).
Skoura, A. et al. Sphingosine-1-phosphate receptor-2 function in myeloid cells regulates vascular inflammation and atherosclerosis. Arterioscler. Thromb. Vasc. Biol. 31, 81–85 (2011).
De Magalhaes Filho, C. D., Downes, M. & Evans, R. Bile acid analog intercepts liver fibrosis. Cell 166, 789 (2016).
Acknowledgements
The authors are supported by grants from the US National Heart Lung and Blood Institute, the US Office of Dietary Supplements and the US National Institute on Alcohol Abuse and Alcoholism (grants R01HL122283 (J.M.B.), P50AA024333 (J.M.B.), R01HL103866 (S.L.H.), P01HL076491 (S.L.H.), R01HL126827 (S.L.H.) and R01DK106000 (S.L.H.)) as well as the Cleveland Clinic Liver Tumor Center of Excellence.
Author information
Authors and Affiliations
Contributions
J.M.B. and S.L.H. substantially contributed to the discussion of content and the review and editing of the manuscript before submission.
Corresponding authors
Ethics declarations
Competing interests
S.L.H. is named as inventor on pending and issued patents held by the Cleveland Clinic relating to cardiovascular diagnostics and therapeutics. He is also a paid consultant for P&G and has received research funds from Astra Zeneca, P&G, Pfizer Inc., Roche Diagnostics and Takeda. S.L.H. has also received royalty payments for inventions or discoveries related to cardiovascular diagnostics or therapeutics from Cleveland HeartLab, Esperion and Siemens. J.M.B. declares no competing interests.
Glossary
- Pattern recognition receptors
-
(PRRs). Host sensors that detect molecules typical for pathogens.
- Atherosclerosis
-
A disease process in which the wall of an artery becomes thickened and inflamed owing to the accumulation of inflammation cells and lipids.
- Thrombosis
-
The formation of a clot inside a vessel.
- Ischaemic stroke
-
A stroke that occurs when a blood vessel to the brain is blocked by a blood clot.
- Metabolome
-
The complete set of small-molecule chemicals found within a biological sample.
- Transient ischaemic attack
-
A brief episode of neurological dysfunction caused by lack of blood flow to the brain; also called a 'mini-stroke'.
- Glycaemia
-
The level of glucose in an individual's blood
- Detergents
-
A surfactant or mix of surfactants that has cleaning or membrane-disturbing properties.
- Taurine
-
A major sulfur-containing amino acid.
- Postprandial state
-
The state immediately following a meal.
Rights and permissions
About this article
Cite this article
Brown, J., Hazen, S. Microbial modulation of cardiovascular disease. Nat Rev Microbiol 16, 171–181 (2018). https://doi.org/10.1038/nrmicro.2017.149
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2017.149
This article is cited by
-
Quinic acid regulated TMA/TMAO-related lipid metabolism and vascular endothelial function through gut microbiota to inhibit atherosclerotic
Journal of Translational Medicine (2024)
-
The underlying molecular mechanisms and biomarkers between periodontitis and COVID-19
BMC Oral Health (2023)
-
Vascular Behçet syndrome: from pathogenesis to treatment
Nature Reviews Rheumatology (2023)
-
Contribution of the microbiome for better phenotyping of people living with obesity
Reviews in Endocrine and Metabolic Disorders (2023)
-
Implication of gut microbes and its metabolites in colorectal cancer
Journal of Cancer Research and Clinical Oncology (2023)