Key Points
-
The evolution of seasonal influenza viruses is an important source of disease burden, as it allows for the reinfection of previously infected or vaccinated individuals
-
Given that 5–15% of the global human population is infected with seasonal influenza viruses each year, it is surprising that new antigenic variants arise only every 3–5 years for A/H3N2 viruses and less frequently for A/H1N1 and B viruses
-
The virus surface glycoprotein haemagglutinin is the primary target of the host immune response, and evolutionary selection pressure drives it to acquire mutations to escape immune recognition without eliminating its receptor binding function
-
Host immunity has a dual role in governing the pace of virus evolution: innate immunity acts as a constraint on the generation of new virus variants, whereas adaptive immunity selects for immune escape mutants
-
The acute nature of influenza virus infections and population-level epidemics provides limited opportunities for evolutionary selection, with most virus diversity being lost before selection can operate
-
Influenza virus vaccines can provide effective protection against infection when they are well matched to circulating viruses, but there remains scope for improving vaccine production and delivery to achieve better effectiveness
Abstract
Despite decades of surveillance and pharmaceutical and non-pharmaceutical interventions, seasonal influenza viruses continue to cause epidemics around the world each year. The key process underlying these recurrent epidemics is the evolution of the viruses to escape the immunity that is induced by prior infection or vaccination. Although we are beginning to understand the processes that underlie the evolutionary dynamics of seasonal influenza viruses, the timing and nature of emergence of new virus strains remain mostly unpredictable. In this Review, we discuss recent advances in understanding the molecular determinants of influenza virus immune escape, sources of evolutionary selection pressure, population dynamics of influenza viruses and prospects for better influenza virus control.
Similar content being viewed by others
Main
Seasonal influenza viruses infect 5–15% of the human population each year, resulting in ∼500,000 deaths worldwide1. The annual recurrence of seasonal epidemics is attributed to the continued evolution of seasonal influenza viruses, which enables them to escape the immunity that is induced by prior infections or vaccination, and to the ability of those viruses to be transmitted efficiently from human-to-human via respiratory droplets, direct contact and fomites. Influenza virus vaccines are effective in preventing the spread of seasonal influenza virus epidemics, but they must be updated regularly to keep pace with the evolution of the circulating viruses.
Influenza viruses belong to the Orthomyxoviridae family of viruses that have negative-sense, single-stranded, segmented RNA genomes. The vast majority of the seasonal influenza virus burden is associated with two types of influenza viruses: A and B (type C viruses also circulate in humans but cause less severe disease2). Influenza A viruses in humans originate from birds and swine3. Their introduction and subsequent adaptation to humans has caused global pandemics (for example, the 1918 'Spanish flu' and 2009 'Swine flu' pandemics) followed by their continued circulation in human populations as seasonal influenza viruses. Influenza A viruses are further classified into subtypes based on the combination of haemagglutinin (HA) and neuraminidase (NA) glycoproteins on their surfaces (Fig. 1a). There are currently 18 HA subtypes and 11 NA subtypes4, most of which circulate in wild birds, but only three combinations are known to have circulated widely in humans: A/H1N1, A/H2N2 and A/H3N2. Of these, A/H1N1 and A/H3N2 subtype viruses currently cause seasonal influenza virus epidemics. The influenza B viruses (Box 1) have no known animal reservoir and have circulated in humans since at least 1940 (Ref. 5) (when the first influenza B virus was isolated), though they have probably circulated in humans for much longer. The influenza B viruses fall into two major lineages (B/Victoria and B/Yamagata) that diverged from each other in the 1970s6,7.
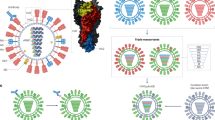
a | The influenza A and influenza B virus genomes are composed of eight viral ribonucleoprotein (vRNP) complexes, encoding at least ten proteins and multiple polypeptides. The surface of the virus is dominated by the haemagglutinin (HA) and neuraminidase (NA) glycoproteins. HA is required for cell attachment and entry and is the primary target of the anti-influenza antibody response; NA is responsible for cell egress and mucus penetration. HA is the primary target of anti-influenza antibodies, particularly the globular head of the HA1 domain. The virus surface protein (M2) acts as an ion channel and the nuclear export protein (NEP) mediates the nuclear export of vRNP complexes. b | HA is a trimeric protein with each monomer being composed of two domains: HA1 (orange) and HA2 (blue). HA1 makes up the majority of the globular head and contains the receptor binding site (RBS) (yellow) for cell attachment. HA2 makes up the majority of the protein stalk domain. c | The five classical antigenic sites of H3 HA (denoted A–E) identified by monoclonal antibody selection and protein crystal structure considerations are shown, all of which have been observed to evolve in nature34,35,36. d | Immune escape substitutions (red) responsible for the majority of antigenic change in the influenza A virus with H3 HA and NA subtype 2 glycoproteins (A/H3N2) from 1968 to present are shown37,38,39. e | Substitutions shown to facilitate the escape of the H3 subtype of HA from broadly neutralizing anti-HA antibodies within the protein stalk domain (amino acids 387 and 391) are highlighted in green48. M1, matrix protein 1.
The influenza A and B virus genomes comprise eight gene segments, including polymerase basic 1 (PB1), polymerase basic 2 (PB2) and polymerase acidic (PA), which encode the viral RNA-dependent RNA polymerase (RdRP) and other proteins that have been proposed to induce cell death (for example, PB1-F2 (Ref. 8)) and to modulate viral pathogenicity (for example, PA-X9); nucleoprotein (NP), which encodes the viral nucleoprotein; matrix (M), which encodes the virus matrix protein (M1) and a virus surface protein (M2) that acts as an ion channel; nonstructural (NS), which encodes both the nonstructural 1 (NS1) protein that is involved in immune evasion and the nuclear export protein (NEP; also known as NS2), which mediates the nuclear export of viral ribonucleoprotein complexes; and HA and NA, which encode the viral primary surface glycoproteins (Fig. 1a). HA is responsible for cell attachment and entry through the binding of sialic acid on the surface of sialylated cells and for subsequent membrane fusion. NA enables the release of new virions from infected cells by cleaving the bonds between HA and sialic acid and by facilitating virus movement through mucus10. Owing to the misincorporation of nucleotides by the viral RdRP during genome replication and the segmented nature of their genomes, influenza viruses evolve through the gradual accumulation of mutations and genome reassortment11, which are also known as antigenic drift and antigenic shift, respectively.
Within each virus subtype, the gradual accumulation of nucleotide mutations and amino acid substitutions in the HA and NA surface glycoproteins periodically results in the emergence of new antigenic variants (often referred to as strains, particularly in the context of vaccine design), a phenomenon called antigenic drift12,13,14. These punctuated antigenic changes, particularly in HA, result in escape from the immunity that was induced by prior infection or vaccination, thus allowing the virus to reinfect individuals who were once immune to the virus and necessitating reformulation of the seasonal influenza virus vaccine15,16,17. The punctuated nature of antigenic evolution makes graphical depictions of antigenic variation of the viral glycoprotein structures appear clustered and gives rise to the concept of antigenic clusters12.
One important consideration regarding the clustered appearance of antigenic evolution is that most antigenic variation is measured by haemagglutination inhibition (HAI) assays. These assays are unlikely to identify all forms of antigenic change, as they rely on antibodies that block sialic acid binding. Other mechanisms of antibody-mediated response (such as antibody-mediated or complement-dependent cellular cytotoxicity and antibody-dependent cellular phagocytosis) could have important roles during infection18,19,20,21,22, and the reliance on HAI assays might therefore occlude some forms of antigenic variation from being identified that could give antigenic evolution a less punctuated appearance. Indeed, even using HAI assays, there are occasional examples of modest, gradual antigenic changes that occur outside of periods marked by more substantial antigenic cluster changes, where a newly emerging antigenic variant replaces existing antigenic variants.
New antigenic variants of A/H3N2 viruses appear every 3–5 years, whereas new antigenic variants of A/H1N1 and influenza B viruses appear less frequently (2–5 years for A/H3N2 viruses compared with 3–8 years for A/H1N1 and influenza B viruses)12,23,24,25. Given that seasonal influenza viruses cause epidemics worldwide, infecting hundreds of millions of people each year1, and that each human is likely to be infected multiple times over their lifetime26,27, it is surprising that new antigenic variants appear so infrequently. In this Review, we explore why influenza viruses evolve so slowly, a concept that challenges the preconception that influenza viruses evolve rapidly. We discuss recent advances in our understanding of the processes that determine the rate of antigenic drift, including the molecular basis of antigenic evolution, within-host selection, between-host transmission and population-level epidemics. Throughout this Review, we focus on how different evolutionary and ecological mechanisms impede the generation of the genetic diversity that underlies virus evolution28, thus making the number of opportunities for evolutionary selection to act on new variants rare. We then discuss the prospects for improved control of influenza virus through vaccination and the potential to predict patterns of viral evolution.
Molecular basis of antigenic drift
The HA protein is the primary target of the human adaptive immune response to the influenza virus and the key component of influenza virus vaccines. HA is a homotrimeric protein, with each monomer containing a globular head (HA1) and a stalk (HA2) domain (Fig. 1b). The majority of the antibody-mediated responses to HA are directed against the globular head29,30, but antibodies that are directed against the stalk occasionally develop in older individuals31,32,33.
Identifying the precise molecular determinants of antigenic drift has been a research priority for decades. Seminal work in the 1980s used monoclonal antibodies and HA protein structures to identify five broadly defined antigenic sites, each composed of multiple amino acid residues on the globular head34,35,36 (Fig. 1c). These studies suggested that at least one amino acid substitution in each of these sites was required to give rise to a new antigenic variant. At the time, a separate study hypothesised that one of these sites could be immunodominant35, but only recently has a more precise definition of the molecular determinants of antigenic drift emerged.
A recent study showed that just seven amino acids around the HA receptor binding site (positions 145, 155, 156, 158, 159, 189 and 193 in H3) were the primary determinants of antigenic change in A/H3N2 viruses from 1968 to 2003, as measured by HAI assays, plaque assays and microneutralization assays37 (Fig. 1d). Single amino acid changes in one of these seven positions in HA can be sufficient to generate new antigenic variants, but in strains from some genetic backgrounds, two or three mutations are required to generate new antigenic variants. Antigenic variants of A/H3N2 viruses that have been identified since 2003 have been similarly associated with changes at these seven positions38,39. The importance of a small numbers of amino acid positions immediately adjacent to the receptor binding site in shaping antigenic evolution has also been shown for avian A/H5N1 viruses40, swine A/H3N2 viruses41,42 and equine A/H3N8 viruses43. However, substitutions that are distant from the receptor binding domain of HA, including substitutions that are not characterized in the most widely used antigenic assays, may have important roles in immune escape18. A recent study that used deep mutational scanning to identify substitutions in HA leading to immune escape found similar evidence for escape being determined by single amino acid substitutions44. Interestingly, deep mutational scanning also showed that the extent of escape from immune recognition by specific antibodies was dependent on the specific amino acid residue at each amino acid position.
Antibodies that target the stalk of the HA protein are much less prevalent than those that target the globular head and are more commonly isolated from older individuals31,32,33. Stalk epitopes are more inaccessible to antibodies than those on the globular head, which reduces their exposure to the immune system; thus, multiple infections are required for the generation of stalk antibodies33,45,46. These antibodies are of substantial interest, as the stalk region of HA tends to be conserved among subtypes, and antibodies that target the stalk could therefore potentially offer inter-subtype protection47. Antigenic escape from antibodies that target the stalk has not been widely reported, but this could be caused by a lack of selection pressure on the HA stalk due to the inaccessibility of the stalk region to antibodies and thus the rarity of antibodies to this region. However, in in vitro experiments, escape from stalk antibodies can be achieved through single amino acid substitutions (particularly through the Q387K substitution in A/H3N2 viruses)48 (Fig. 1e).
Antibodies are also raised against influenza virus NA during infection and have been shown to reduce viral replication and disease severity in mice49 and in humans50. However, NA antigenic variability has been studied to a lesser extent than that of HA owing to the immunodominance of HA, the greater neutralizing potential of anti-HA antibodies and the challenges that are associated with phenotypic assays of NA antigenicity. Similar to that of HA, the antigenic evolution of NA is clustered13,14, but the timing of emergence of new NA antigenic variants is asynchronous with antigenic drift in HA14. Influenza virus vaccines that are currently licensed include immunogenic quantities of NA, but the contribution of NA to vaccine efficacy is not known51, and the immunogenicity of the NA within vaccines could be compromised by the low thermostability of NA during vaccine production52.
Constraints on viral evolution
Given that only a small number of amino acid substitutions are required to generate new influenza virus antigenic variants, it is surprising that new variants are observed only every few years. Over the past decade, a number of hypotheses that concern host–virus interactions have been proposed to explain the frequency of emergence of new antigenic variants53,54,55,56,57,58,59,60.
The identification of the seven amino acid positions that are responsible for antigenic change in A/H3N2 viruses and their close proximity to the receptor binding site37 gives rise to a hypothesis about a key constraint on immune escape. Escape from antibodies that target the receptor binding site may require a delicate balancing act: mutations in these positions must cause antigenic change without disrupting receptor binding function. Thus, at any one time, only a minor subset of possible amino acid changes in each position can retain receptor binding function while causing antigenic change. This hypothesis could explain an interesting evolutionary event in the history of seasonal A/H1N1 viruses where between 2006 and 2007, viruses in three antigenically similar but geographically segregated HA genetic lineages each acquired the same amino acid substitution to become the same new antigenic variant24. This substitution, K140E, is immediately adjacent to the receptor binding site and structurally equivalent to the sites that were identified for A/H3N2 viruses (Fig. 1c).
Receptor binding avidity for sialic acid could also have an important role61 in limiting the rate of antigenic evolution, particularly if the extent of the antigenic similarity between old and new variants is associated with the extent of receptor binding disruption caused by the acquisition of antigenicity-altering substitutions around the receptor binding site. As sialic acid binding is essential for viral entry, HA subtypes with lower binding avidity for sialic acid could be less able to tolerate amino acid substitutions near the receptor binding site than viruses with higher receptor binding avidity for sialic acid. This balance of receptor binding and immune escape might effectively limit antigenic drift, thus explaining the relative uniformity of antigenic differences between successively circulating antigenic clusters12.
The genetic strain in which an amino acid substitution appears is likely to be a major determinant of the tolerance of that substitution. Experimental work on NP62 and NA63 has shown that even closely related viruses can vary in their ability to tolerate particular amino acid substitutions. A theoretical study that used dynamical models of influenza epidemiology and evolution53 showed that the rate of antigenic change could be constrained by the need to acquire multiple genetic mutations in order to acquire those substitutions that alter the antigenicity of the protein with little or no impact on the fitness of the virus. A more recent theoretical modelling study proposed that mutations that had a deleterious effect on viral fitness could limit antigenic change by requiring that substitutions that alter the antigenicity of HA confer a benefit to the fitness of the virus that offsets the cost of the accumulation of deleterious mutations54.
Even in the absence of these constraints, the opportunities for immune selection to affect an escape variant are likely to be rare, perhaps because antigenic drift variants need to reach a sufficient density for selection to operate. A hypothetical assessment of the factors shaping within-host virus evolution suggests that the acute nature of influenza virus infections combined with the probabilities of mutant virus emergence and the restriction of the innate immune response (see section on 'Selection pressures') should make the emergence of antigenic variants driven by within-host selection a rare event64,65,66 (though more probable in prolonged infections of immunocompromised individuals67). For example, the mutation rate of the influenza A virus RdRP is in the range of ∼2.0 × 10−6 to ∼2.0 × 10−4 mutations per site per round of genome replication68,69,70,71; thus, the probability of generating a specific antigenic drift variant through a single nucleotide mutation is ∼2/105 (the inverse of the mean of the two rates listed above) per round of replication. In an infected cell that produces ∼104 virions72, even if the variant arises in the first round of replication, it will be vastly outnumbered by all the other progeny that are generated in that cell. After exiting the cell, virus progeny must survive the mucociliary clearance system before antibody selection can operate. This immune barrier is likely to substantially reduce the population of newly generated variants, with only a minority of virus progeny successfully infecting other cells (Fig. 2).
a | An illustration depicting within-cell and within-host factors that reduce virus diversity, including the frequency of de novo mutant generation, innate and adaptive immune capture and subsequent infection of available host cells. Mutants are likely to be produced in each infected cell owing to the error-prone nature of the virus polymerase complex. Though most variants inside a donor cell are likely to be unfit, some will be fit. After exiting an infected cell, a substantial portion of free virus will be trapped in mucus. In previously infected individuals, free virus populations will be further reduced by the adaptive immune response. These virus population bottlenecks reduce both virus numbers and the newly generated mutant diversity that can propagate in recipient cells. b | The factors for between-host and population-level spread of influenza viruses include donor host excretion, recipient host infection and epidemic seeding. Most virus diversity generated within an infected host will be lost during transmission to a new host, as only a small number of virus particles are likely to establish the infection in the new host. This same process occurs at the population level, as long-term virus survival requires that the viruses from an epidemic manage to start epidemics in new locations. Much like the between-host transmission bottleneck, the between-epidemic bottleneck is likely to similarly reduce virus diversity. vRNP, viral ribonucleoprotein.
Selection pressures
Evolutionary selection pressure on influenza virus antigenicity primarily stems from immunity that is induced by prior infections or vaccinations. The extinction of old antigenic variants upon the emergence of new ones is evidence that immune selection on influenza virus occurs12; however, the mechanisms of within-host selection that account for this observation are not well understood. Indeed, a recent study65 showed that vaccination and prior infections had minimal effects on the emergence of new antigenic variants in the 166 individuals studied. Understanding how within-host immune selection could influence virus evolution requires careful consideration of the temporal and spatial features of influenza virus infection and of the respective roles of innate and adaptive immunity in naive and previously exposed individuals.
Innate immune selection. In individuals who have not been previously exposed to influenza virus antigens (naive individuals), infection is largely controlled by the innate immune response. Upon initial exposure to the virus, the mucosal barrier creates a layer of mucus rich in sialic acid, which acts as a decoy by binding to the viral HA protein and thus traps a substantial portion of viruses, effectively reducing the infectious dose10,73. After the virus has infected host cells, Toll-like receptor signalling and interferon-mediated responses are triggered, resulting in the rapid establishment of an antiviral state that limits the permissivity of nearby cells to virus replication74,75. The efficiency of innate immune selection is dependent on infectious dose76, virus immunogenicity that leads to a pro-inflammatory state77,78 and host genetic factors that affect the strength of innate immune reactivity79,80,81. Overall, the innate immune response constrains virus evolution by decreasing the availability of cells that are permissive to virus replication and by reducing the virus diversity on which selection can operate28, independent of the antigenic phenotype.
Adaptive immune selection. The recurrent nature of influenza virus epidemics and repeated vaccination results in a complex mosaic of immune selection pressures from antibodies and immune memory cells. For an individual, the strength of the immune selection depends on the history of virus or vaccine antigen exposure82, the similarity of the antigen from the infecting virus to antigens from previous exposures27 and the extent of immune waning, which can lead to reduced selection pressures despite previous exposures83.
In naive individuals, viruses that escape the innate response are able to replicate, leading to a peak in virus titre 24–72 hours after infection, depending on the infectious dose84 (Fig. 3a). However, the time that is required for the de novo generation of virus-specific antibodies is 7–10 days after infection85, leading to minimal within-host antigenic selection on the virus during primary infection. In addition to the low selection pressure on the virus during primary infection, the acute nature of influenza virus infections leads to transient exposure of virus antigens to the adaptive immune system, which requires the prolonged presence of an antigen to develop highly specific adaptive immunity86. Thus, naive individuals would likely require more than one infection or vaccination to generate highly specific antibodies that are able to confer protection87. This potential need for more than one exposure to generate immune selection pressure means that, despite the high prevalence, infections in young children could have a limited role in virus evolution.
The left y axis of each panel represents the relative contribution of each type of host immune response (innate response, mucosal response, recall adaptive response and novel adaptive response) to the magnitude of immune response against influenza virus. The red line corresponds to the right y axes, which represent the relative influenza virus titre. These panels were generated from basic immunological principles of primary and secondary infections assuming an average duration of influenza virus infection. However, dynamics could differ substantially in immunocompromised individuals165,166. a | Primary influenza virus infection in an individual who is immunologically naive to influenza virus is shown. Induction of interferon response correlates with influenza virus replication167,168 and dominates the early immune response169. This time course of generation of novel B cell response is consistent with dynamics of antigen-specific germinal centre reaction, which takes place approximately six days after antigen exposure170. A low degree of mucosal immune response can be present as a result of antigen-independent induction of natural immunoglobulin M antibodies171. b | In the case of secondary infection with a variant with partial cross reactivity to previous antigenic variants, immune selection is based on cross-reactive serological and memory B cell responses155, which have competitive advantages relative to naive B cell responses172. As in part a, the generation of specific adaptive immunity occurs after the peak of virus replication. However, the generation of antibodies to drifted strains is limited by the activity of cross-reactive antibodies, which bind to the HA head and reduce the amount of influenza antigen available to trigger a specific response94,95. c | In the case of secondary infection with a new antigenic variant with low cross reactivity to previous influenza viruses, neutralization of the virus by prior immunity is minimal, and the virus replicates to titres similar to those in primary infection (as in part a). This allows for sufficient antigen exposure to generate novel B cell responses specific to the new variant, but these typically occur 5–6 days after infection169. Thus, increasing antigenic distance between strains favours the generation of novel immune responses147.
After primary infection, subsequent exposures to previously encountered antigenic variants induce mucosal immune responses to previously encountered virus epitopes. Such mucosal responses are largely mediated by immunoglobulin A (IgA) antibodies in the upper respiratory tract, which are able to cross the epithelial barrier from the blood to the upper respiratory tract lumen in order to bind to and neutralize influenza viruses, thus preventing the infection of host cells88 (a process referred to as 'immune exclusion' (Ref. 89)). As long as highly specific antibodies to the infecting virus were generated after a previous exposure, an infection can be rapidly contained with limited additional immune activation, which explains the reduction in serological responses after each subsequent infection with homologous influenza virus strains26. However, this response is likely to strongly select for any new antigenic variants that are present in an infecting inoculum and to be a key source of selection pressure for virus evolution at the human population level.
Secondary infections with partially cross-reactive antigenic variants typically lead to incomplete early antibody neutralization and infection90,91. As most influenza virus infections result in the recall of some immune memory27, individuals typically have the highest cumulative antibody titres against viruses that were encountered earlier in life27,92,93,94. This immunological backboosting gives rise to so-called antigenic seniority of antigenic variants that were encountered in the first decade of life95 and is consistent with the hypothesis of 'original antigenic sin' (Ref. 96), where the recall of previously generated immunological memory compromises the generation of a highly specific antibody response to the new antigenic variant (reviewed in Ref. 19). During infections in which the immune response is dominated by a recall of immune memory (Fig. 3b), there is the possibility of immune selection pressure for new antigenic variants arising de novo, but this will be limited by the acute nature of typical infections, the predominance of existing antibodies with sub-neutralizing activities and the potentially reduced virus population size on which selection can exert its effects.
Infection with an antigenic variant that has a substantially reduced cross reactivity to previously encountered antigenic variants is primarily subject to restriction and clearance by the innate immune response independent of its antigenic profile, as it only weakly elicits a memory recall response and is unlikely to be neutralized by existing antibodies. Similar to naive individuals, selection that is mediated by virus-specific antibodies is likely to occur only upon secondary exposure owing to the asynchrony in the generation of novel adaptive immune response and peak virus titre (Fig. 3c).
Measuring immune selection. Anti-influenza virus antibody levels and cross reactivity with new viruses are routinely used to assess potential protection against new influenza virus infections97. When combined with characterizations of virus genomic diversity, serological assays can provide insight into immune-driven virus evolution40,98. However, because the majority of influenza infections are restricted to the upper respiratory tract, in order for antibodies to apply selection on newly infecting variants, they need to be present in the lumen of the upper respiratory tract at the point of infection. Not all antibodies from the serum have access to this site of infection owing to differences in their capabilities to cross epithelial barriers, an ability that is mostly limited to secreted IgA antibodies (and IgG antibodies in the lungs at a much lower rate). This means that high immune activation and elevation of serum antibody levels do not directly reflect the antibody-mediated selection applied to influenza viruses at the site of infection, as only a small subset of these antibodies will have access to this site. Recent studies have shown that serum antibody titres can be a weak correlate of immune reactivity upon infection51,99,100 (reviewed in Ref. 101), suggesting that serum antibody levels overestimate the strength of immune selection pressure. An improved understanding of the development and the maintenance of tissue-resident immune memory cells that are specific for influenza virus is required to assess the involvement of mucosal immunity in recall responses and its potential role in driving virus evolution.
Between-host and epidemic dynamics
New antigenic variants are likely to be lost upon transmission to subsequent hosts unless they are able to replicate to high levels within an individual who is infected. The virus population bottlenecks that occur during virus egress from the donor host, ingress into the recipient host and penetration of innate immune barriers are each likely to be substantial, resulting in major losses of newly generated viral diversity64,102,103,104,105 (Fig. 2b). The extent of these bottlenecks varies with route of transmission, with respiratory droplet transmission imposing a stronger bottleneck than contact transmission102,103.
Similar to within-host and between-individual transmission processes, larger-scale epidemic dynamics also impose strong bottlenecks on influenza virus evolution. The seasonal nature and short duration of influenza virus epidemics combined with the large viral diversity in an ongoing epidemic mean that newly emerged variants, even when they are more fit than the dominant variants, have little time to compete with existing variants, to replicate to high levels and to seed subsequent epidemics.
The timing of seasonal influenza virus epidemics varies globally, with the majority of influenza virus infections in temperate regions occurring in the winter months and during the rainy season in tropical regions (though epidemic timing is variable in some tropical countries)106,107,108 (Fig. 4a). Experimental work in animal models has shown that respiratory droplet transmission of seasonal influenza viruses occurs most efficiently in lower temperature and lower relative humidity environments compared with higher temperature or higher humidity environments109. This could explain the seasonality of influenza virus epidemics in the temperate regions of the Northern and Southern Hemispheres. However, in many parts of the tropics, influenza virus epidemics coincide with local rainy seasons, during which temperatures and relative humidity are both high. This implies that in the tropics, influenza virus transmission that occurs via direct contact or fomites could have a more important role than respiratory droplet transmission. If true, the larger population bottlenecks that are associated with contact transmission could facilitate more rapid evolution in tropical environments than the possibly respiratory droplet dominated transmission in temperate environments. The onset of epidemics has also been linked to fluctuations in absolute humidity110,111.
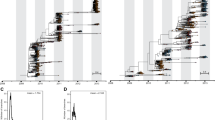
a | The mean influenza virus epidemic activity by month based on virologically confirmed samples of influenza A virus with haemagglutinin (HA) subtype 3 and neuraminidase (NA) subtype 2 glycoproteins (A/H3N2), A/H1N1, B/Victoria, and B/Yamagata viruses from 2000 to 2015 (Ref. 164) is shown. The 2009 A/H1N1 pandemic caused outbreaks with atypical seasonality around the world. To avoid biasing the estimates of seasonality, A/H1N1 viruses collected in 2009 were excluded. For countries for which data was not available for all years from 2000 to 2015, data for years available was used. The number of virologically confirmed cases reported in FluNet164 were calculated for each month for each country and converted into mean case frequencies per month by dividing each monthly total by the total number of cases recorded for that country. Colours represent the relative level of influenza virus activity for that country during that month: dark red represents high influenza prevalence; dark blue indicates low prevalence. b | Estimates of virus migration rate between regions based on phylogenetic inferences of influenza HA genetic sequence data are depicted by arrows. Line thickness indicates the mean number of virus migration events per year. Arrowheads indicate the strength of directionality of virus migration. The circle area for each region indicates that region's contribution to the global ancestry of viruses. Circles are coloured to differentiate regions of the world but do not otherwise affect the interpretation of the figure. Africa could not be included in these inferences owing to the substantially lower levels of surveillance in Africa relative to the rest of the world. Lat, latitude. Part b adapted with permission from Ref. 24, Macmillan Publishers Limited.
Winter epidemics in temperate regions and rainy season epidemics in the tropics could also be linked to changes in human behaviour, for example, by increases in time spent indoors. Host mixing in confined spaces can have an important role in viral transmission, as evidenced by summertime outbreaks of influenza virus in elderly care homes and prisons112,113,114 and by the effect of school closures on limiting viral transmission115. Additionally, seasonal fluctuations in human immunity, particularly during the winter months in which pro-inflammatory responses are upregulated116, could also have a role in the seasonality of influenza epidemics.
Global dynamics
Owing to the acute nature of influenza virus epidemics, the long-term circulation of influenza viruses in the human population is driven by the global movement of viruses. The extent to which viruses move internationally versus persisting locally between different epidemics has been of interest since at least the 1800s117,118. Various hypotheses have been proposed to explain the global movement of viruses, including back and forth migration between the temperate regions of the Northern and Southern Hemispheres due to differences in the timing of winter, continual seeding of epidemics in temperate regions from the tropics and seeding of viruses from China. The hypothesis based on the seeding of viruses from China originates from the perceived historical importance of China as a source of pandemic viruses119 and the sheer magnitude of the human population in China.
Phylogenetic analyses of virus genome data sets from New York, United States, and South Island, New Zealand, and Australia representing the Northern and Southern Hemispheres, respectively, provided evidence against the local persistence of A/H3N2 viruses between epidemics120. Subsequent studies on the global dynamics of A/H3N2 viruses provided further evidence of local extinction of viruses between epidemics and identified East and Southeast (E–SE) Asia as the global source of A/H3N2 viruses24,121,122,123,124,125,126. The advent of influenza virus surveillance data from India (Box 2) led to the discovery that the global source region for A/H3N2 viruses also includes India in addition to E–SE Asia24 and confirmed previous hypotheses121. In E–SE Asia and India, A/H3N2 viruses become locally extinct between epidemics, but variation in local climates enables viruses to continuously circulate by passing from epidemic to epidemic in the region.
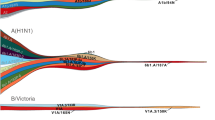
Recently, the global circulation pattern of seasonal influenza was shown to vary with the rate of antigenic evolution of different virus types and subtypes24. Unlike A/H3N2 viruses that spread globally from E–SE Asia and India each year, A/H1N1 and influenza B viruses sporadically persist locally between epidemics in multiple regions of the world, giving rise to multiple co-circulating genetic lineages. These co-circulating lineages occasionally resulted in divergent antigenic variants, suggesting a need for region-specific influenza B virus vaccine components21.
The differences in the global dynamics of seasonal influenza viruses are likely a result of differences in the rates of virus evolution, potentially leading to the observed variations in age distributions of infection24; that is, viruses that evolve rapidly to escape immunity can infect people of all ages more often than viruses that evolve at a slower rate, which primarily infect children and only sporadically infect adults. For example, antigenic regions of the HA glycoprotein of A/H3N2 viruses evolve at a faster rate than A/H1N1 virus and influenza B virus antigenic regions127, and these differences in rates of evolution lead to larger and more frequent A/H3N2 virus epidemics (which could, in turn, drive evolution128), resulting in people of a wide variety of ages becoming infected. A/H1N1 and influenza B viruses tend to cause smaller and less frequent epidemics and to infect younger individuals. Combined with the more frequent global travel of adults than young children, these differences in the distribution of age of infection provide a mechanism for A/H3N2, A/H1N1 and influenza B viruses to follow similar patterns of movement around the world but at the same time provide a mechanism for A/H3N2 viruses to spread more rapidly than A/H1N1 and influenza B viruses (Fig. 4b).
Options for the control of influenza
Vaccines are the primary tool for preventing and controlling seasonal influenza virus infections. The effect of vaccination on virus evolution is likely to be small compared with the effect of influenza virus infection, as <10% of the world population is routinely vaccinated against influenza virus129. However, the impact of virus evolution on vaccine efficacy is substantial130,131. Accordingly, the formulation of influenza virus vaccines is re-assessed twice annually (in February in the Northern Hemisphere and in September in the Southern Hemisphere) to try to ensure that the viruses in the vaccine match circulating viruses that are most likely to cause future epidemics38,39,132. These decisions are made approximately nine months before the vaccines are intended to confer protection in order to accommodate the time that is necessary for producing, distributing and administering the vaccine. The long vaccine production process is largely caused by the use of chicken egg-based vaccine production technologies, and this duration creates a window of opportunity for new virus variants to emerge, often resulting in decreased vaccine efficacy133. Egg-based vaccine production also has the potential to cause unintended antigenic change to the vaccine virus when the human-adapted virus is evolved to replicate efficiently in eggs134. New vaccine production methods could reduce these inefficiencies (Table 1), potentially leading to more efficient vaccine production processes and more effective vaccines. Influenza virus vaccines can now be produced using cell-based culture manufacturing and recombinant technologies without culturing the virus in chicken eggs (reviewed in Refs 135,136), thus allowing the use of viruses that are genetically closer to wild-type viruses in vaccines and reducing the risk of viral evolution during laboratory passage136,137,138. Recombinant technologies have the added benefit of substantially reducing production times compared with egg-based and cell-based technologies, as production can begin once a virus genome has been sequenced rather than starting with a live virus136. If the majority of influenza virus vaccines were produced using recombinant technologies, the vaccine strain selection decisions could be made closer to the time of intended vaccine use, thus reducing the window of opportunity for the emergence of new antigenic variants in the human population.
A complementary approach to improve influenza virus vaccine production is to attempt to predict future patterns of influenza virus evolution. The ability to predict how influenza viruses will evolve would substantially improve options for vaccine development and efficacy. To date, the accurate prediction of future antigenic change to influenza viruses has been hindered by a limited understanding of how changes in viral genotype affect viral phenotype and of the interplay of factors governing the rate of antigenic evolution.
Recent efforts have made progress in predicting influenza virus genetic evolution by analysing genetic sequence data and phylogenetic branching patterns to predict the success of genetic lineages139,140 and may become useful for predicting which virus lineages are likely to survive or to become extinct. Attempts to integrate antigenic data into these predictions have resulted in improvements in prediction accuracy141. These methods, combined with tools that track the real-time evolution of influenza viruses, like NextFlu142, are helping to integrate different types of surveillance data, particularly antigenic data, and have the potential to help determine the vaccine strains that are included in influenza virus vaccines.
There is also a need to improve how vaccines are used to maintain immunity. The annual re-vaccination designed to maintain protection against influenza viruses can have a negative effect on vaccine efficacy, owing to the repeated recall of previously generated immune memory with a limited stimulation of novel immune responses143,144,145,146. This decline in vaccine efficacy associated with vaccination history is thought to be the result of a decline in boosts of antibody titres with each repeated exposure to similar viruses and antigenic differences between the vaccine virus and circulating viruses147,148. There are four broad strategies for combatting this problem: generating broadly neutralizing responses to conserved epitopes to reduce the need for revaccination (that is, creating a universal vaccine)47,149; better stimulating mucosal immunity to improve the breadth of cross-protection150,151; vaccinating with viruses that are sufficiently antigenically dissimilar to previously circulating viruses to leverage backboosting for protection against circulating strains27,152; and improving the adjuvant composition of vaccines to enhance the generation of novel immune responses153,154.
In addition, influenza virus vaccine efficacy could likely be improved by better vaccine strain selection strategies132. Vaccine strain selection is largely based on the antigenic characterization of circulating viruses by use of antisera in binding assays that are produced during primary influenza virus infections in ferrets. These assays can overlook the complexity of the immunity that results from multiple infections and vaccinations18 and might not accurately reflect protection in individuals who have been exposed multiple times, particularly in elderly people. Emerging technologies for immune phenotyping and next-generation sequencing of antibody repertoires to improve our understanding of protection could facilitate a more strategic selection of vaccine strains to elicit better protection in individuals who have been previously exposed to influenza viruses through infections and repeated vaccinations155,156,157.
Conclusions
In this Review, we have discussed how processes that operate at the level of the virus, host and human population act in concert to determine the rate of influenza virus evolution and, in turn, shape the global dynamics of seasonal influenza viruses. Despite the high number of influenza virus infections each year, the opportunities for evolutionary selection pressures to act on are likely to be rare. The factors that limit these opportunities are inherent to the biology of the virus (owing to the limited number of mutations that can be accommodated by the virus without substantially decreasing fitness), immunological (owing to the way complex exposure histories limit the generation of novel immunity) and epidemiological (owing to the acute nature of infections and epidemics).
Studies of these evolutionary processes will benefit from the increasing resolution of next-generation sequencing tools, which can be used to identify mutations during within-host and between-host processes, particularly with regard to the de novo generation of new mutants and the selective advantages that mutants confer to escape stochastic extinction. Similarly, there is a growing need for better computational and mathematical modelling frameworks for integrating different data sources about processes occurring within and between hosts. Moreover, standardizing practices across different laboratories and the sharing of primary virological data beyond genetic sequence data, particularly antigenic data, is crucial for heightening our understanding of the evolution of seasonal influenza viruses.
Immune repertoire sequencing for profiling immune responses during infection and vaccination could lead to the identification of markers of the host immune response that are shared among individuals and thus provide opportunities for the systematic characterization of within-host and population-level selective pressures that affect virus evolution158. These common immune markers could be used for predicting the efficacy of vaccines during the design stage159. However, the use of such large-scale sequencing approaches in influenza virus vaccine design requires an understanding of how changes in viral genotype affect specific immune phenotypes, which is currently lacking.
Improving our understanding of influenza virus evolution and our ability to predict patterns of viral evolution will lead to better influenza virus control. However, as influenza virus is a global pathogen, new measures for controlling influenza virus will need to be implemented worldwide. Currently, global seasonal influenza virus vaccine production is ∼500 million doses per year129 for a human population of >7 billion people. Reducing the global burden of influenza virus will require substantial increases in vaccine production and increases in access to influenza virus vaccines, antiviral drugs and non-pharmaceutical interventions. This is particularly important for areas where the public health infrastructure is poor.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
07 November 2017
In Figure 4 of the original online version of the article, the influenza virus epidemic activity by month was incorrectly labelled. This has now been corrected in the online and print versions. We apologize to the authors and to readers for any confusion caused.
References
Stöhr, K. Influenza—WHO cares. Lancet Infect. Dis. 2, 517 (2002).
Zambon, M. C. Epidemiology and pathogenesis of influenza. J. Antimicrob. Chemother. 44, 3–9 (1999).
Russell, C. A. et al. Improving pandemic influenza risk assessment. eLife 3, e03883 (2014).
Tong, S. et al. New World bats harbor diverse influenza A viruses. PLoS Pathog. 9, e1003657 (2013).
Francis, T. A. New type of virus from epidemic influenza. Science 92, 405–408 (1940).
Biere, B., Bauer, B. & Schweiger, B. Differentiation of influenza B virus lineages Yamagata & Victoria by real-time PCR. J. Clin. Microbiol. 48, 1425–1427 (2010).
Kanegae, Y. et al. Evolutionary pattern of the hemagglutinin gene of influenza B viruses isolated in Japan: cocirculating lineages in the same epidemic season. J. Virol. 64, 2860–2865 (1990).
Chen, W. et al. A novel influenza A virus mitochondrial protein that induces cell death. Nat. Med. 7, 1306–1312 (2001).
Jagger, B. W. et al. An overlapping protein-coding region in influenza A virus segment 3 modulates the host response. Science 337, 199–204 (2012).
Cohen, M. et al. Influenza A penetrates host mucus by cleaving sialic acids with neuraminidase. Virol. J. 10, 321 (2013).
Westgeest, K. B. et al. Genomewide analysis of reassortment and evolution of human influenza A(H3N2) viruses circulating between 1968 and 2011. J. Virol. 88, 2844–2857 (2014).
Smith, D. J. et al. Mapping the antigenic and genetic evolution of influenza virus. Science 305, 371–376 (2004). This is a seminal study that documented the continuous genetic but punctuated antigenic evolution of A/H3N2 viruses and introduced antigenic cartography — a computational tool for quantifying differences in virus antigenic phenotype.
Westgeest, K. B. et al. Genetic evolution of the neuraminidase of influenza A (H3N2) viruses from 1968 to 2009 and its correspondence to haemagglutinin evolution. J. Gen. Virol. 93, 1996–2007 (2012).
Sandbulte, M. R. et al. Discordant antigenic drift of neuraminidase and hemagglutinin in H1N1 and H3N2 influenza viruses. Proc. Natl Acad. Sci. USA 108, 20748–20753 (2011).
Salk, J. E. & Suriano, P. C. Importance of antigenic composition of influenza virus vaccine in protecting against the natural disease. Am. J. Publ. Health Nat. Health 39, 345–355 (1949).
Kilbourne, E. D. et al. The total influenza vaccine failure of 1947 revisited: major intrasubtypic antigenic change can explain failure of vaccine in a post-World War II epidemic. Proc. Natl Acad. Sci. USA 99, 10748–10752 (2002).
Tricco, A. C. et al. Comparing influenza vaccine efficacy against mismatched and matched strains: a systematic review and meta-analysis. BMC Med. 11, 153 (2013).
Linderman, S. L. et al. Potential antigenic explanation for atypical H1N1 infections among middle-aged adults during the 2013–2014 influenza season. Proc Natl Acad. Sci. USA 111, 15798–15803 (2014).
Cobey, S. & Hensley, S. E. Immune history and influenza virus susceptibility. Curr. Opin. Virol. 22, 105–111 (2017).
DiLillo, D. J., Palese, P., Wilson, P. C. & Ravetch, J. V. Broadly neutralizing anti-influenza antibodies require Fc receptor engagement for in vivo protection. J. Clin. Invest. 126, 605–610 (2016).
Terajima, M. et al. Complement-dependent lysis of influenza A virus-infected cells by broadly cross-reactive human monoclonal antibodies. J. Virol. 85, 13463–13467 (2011).
Jegaskanda, S., Weinfurter, J. T., Friedrich, T. C. & Kent, S. J. Antibody-dependent cellular cytotoxicity is associated with control of pandemic H1N1 influenza virus infection of macaques. J. Virol. 87, 5512–5522 (2013).
Chen, R. & Holmes, E. C. The evolutionary dynamics of human influenza B virus. J. Mol. Evol. 66, 655–663 (2008).
Bedford, T. et al. Global circulation patterns of seasonal influenza viruses vary with antigenic drift. Nature 523, 217–220 (2015). This is the first detailed comparison of the global circulation dynamics of all four seasonal influenza viruses and is the most complete characterization of the global dynamics of seasonal influenza viruses to date.
Vijaykrishna, D. et al. The contrasting phylodynamics of human influenza B viruses. eLife 4, e05055 (2015).
Kucharski, A. J. et al. Estimating the life course of influenza A(H3N2) antibody responses from cross-sectional data. PLoS Biol. 13, e1002082 (2015).
Fonville, J. M. et al. Antibody landscapes after influenza virus infection or vaccination. Science 346, 996–1000 (2014).
Cheung, P. P. H. et al. Generation and characterization of influenza A viruses with altered polymerase fidelity. Nat. Commun. 5, 4794 (2014).
Virelizier, J.-L. Host defenses against influenza virus: the role of anti-hemagglutinin antibody. J. Immunol. 115, 434–439 (1975).
Bizebard, T. et al. Structure of influenza virus haemagglutinin complexed with a neutralizing antibody. Nature 376, 92–94 (1995).
Margine, I. et al. H3N2 influenza virus infection induces broadly reactive hemagglutinin stalk antibodies in humans and mice. J. Virol. 87, 4728–4737 (2013).
Moody, M. A. et al. H3N2 influenza infection elicits more cross-reactive and less clonally expanded anti-hemagglutinin antibodies than influenza vaccination. PLoS ONE 6, e25797 (2011).
Nachbagauer, R. et al. Age dependence and isotype specificity of influenza virus hemagglutinin stalk-reactive antibodies in humans. mBio 7, e01996-15 (2016). This is a detailed analysis of how broadly influenza neutralizing antibodies accumulate with age and of their possible role in the decreased rate of influenza infection among elderly individuals.
Wiley, D. C., Wilson, I. A. & Skehel, J. J. Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 289, 373–378 (1981).
Skehel, J. J. et al. A carbohydrate side chain on hemagglutinins of Hong Kong influenza viruses inhibits recognition by a monoclonal antibody. Proc. Natl Acad. Sci. USA 81, 1779–1783 (1984).
Gerhard, W., Yewdell, J., Frankel, M. E. & Webster, R. Antigenic structure of influenza virus haemagglutinin defined by hybridoma antibodies. Nature 290, 713–717 (1981).
Koel, B. F. et al. Substitutions near the receptor binding site determine major antigenic change during influenza virus evolution. Science 342, 976–979 (2013). This is a key paper showing that the majority of substantial antigenic changes as determined by experimental assays from 1968 to – have been associated with amino acid substitutions in just seven positions in the HA protein.
Barr, I. G. et al. WHO recommendations for the viruses used in the 2013–2014 Northern Hemisphere influenza vaccine: epidemiology, antigenic and genetic characteristics of influenza A(H1N1)pdm09, A(H3N2) and B influenza viruses collected from October 2012 to January 2013. Vaccine 32, 4713–4725 (2014).
Klimov, A. I. et al. WHO recommendations for the viruses to be used in the 2012 Southern Hemisphere influenza vaccine: epidemiology, antigenic and genetic characteristics of influenza A(H1N1)pdm09, A(H3N2) and B influenza viruses collected from February to September 2011. Vaccine 30, 6461–6471 (2012).
Koel, B. F. et al. Antigenic variation of clade 2.1 H5N1 virus is determined by a few amino acid substitutions immediately adjacent to the receptor binding site. mBio 5, e01070-14 (2014).
Abente, E. J. et al. The molecular determinants of antibody recognition and antigenic drift in the H3 hemagglutinin of swine influenza A virus. J. Virol. 90, 8266–8280 (2016).
Lewis, N. S. et al. The global antigenic diversity of swine influenza A viruses. eLife 5, e12217 (2016).
Lewis, N. S. et al. Antigenic and genetic evolution of equine influenza A (H3N8) virus from 1968 to 2007. J. Virol. 85, 12742–12749 (2011).
Doud, M. B., Hensley, S. E. & Bloom, J. D. Complete mapping of viral escape from neutralizing antibodies. PLoS Pathog. 13, e1006271 (2017). This study demonstrates the excellent use of tools for deep mutational scanning to investigate the virus genetic consequences of antibody selection.
Kirchenbaum, G. A., Carter, D. M. & Ross, T. M. Sequential infection in ferrets with antigenically distinct seasonal H1N1 influenza viruses boosts hemagglutinin stalk-specific antibodies. J. Virol. 90, 1116–1128 (2016).
Nachbagauer, R. et al. Induction of broadly reactive anti-hemagglutinin stalk antibodies by an H5N1 vaccine in humans. J. Virol. 88, 13260–13268 (2014).
Okuno, Y., Isegawa, Y., Sasao, F. & Ueda, S. A common neutralizing epitope conserved between the hemagglutinins of influenza A virus H1 and H2 strains. J. Virol. 67, 2552–2558 (1993).
Chai, N. et al. Two escape mechanisms of influenza A Virus to a broadly neutralizing stalk-binding antibody. PLoS Pathog. 12, e1005702 (2016). This study provides an important functional demonstration of the ability of influenza viruses to escape broadly neutralizing antibodies through a small number of amino acid substitutions.
Schulman, J. L., Khakpour, M. & Kilbourne, E. D. Protective effects of specific immunity to viral neuraminidase on influenza virus infection of mice. J. Virol. 2, 778–786 (1968).
Murphy, B. R., Kasel, J. A. & Chanock, R. M. Association of serum anti-neuraminidase antibody with resistance to influenza in man. N. Engl. J. Med. 286, 1329–1332 (1972).
Eichelberger, M. C. & Wan, H. Influenza neuraminidase as a vaccine antigen. Curr. Top. Microbiol. Immunol. 386, 275–299 (2015).
Sultana, I. et al. Stability of neuraminidase in inactivated influenza vaccines. Vaccine 32, 2225–2230 (2014).
Koelle, K., Cobey, S., Grenfell, B. & Pascual, M. Epochal evolution shapes the phylodynamics of interpandemic influenza A (H3N2) in humans. Science 314, 1898–1903 (2006).
Koelle, K. & Rasmussen, D. A. The effects of a deleterious mutation load on patterns of influenza A/H3N2's antigenic evolution in humans. eLife 4, e07361 (2015).
Zinder, D., Bedford, T., Gupta, S. & Pascual, M. The roles of competition and mutation in shaping antigenic and genetic diversity in influenza. PLoS Pathog. 9, e1003104 (2013).
Recker, M., Pybus, O. G., Nee, S. & Gupta, S. The generation of influenza outbreaks by a network of host immune responses against a limited set of antigenic types. Proc. Natl Acad. Sci. USA 104, 7711–7716 (2007).
Meyer, A. G. & Wilke, C. O. Geometric constraints dominate the antigenic evolution of influenza H3N2 hemagglutinin. PLoS Pathog. 11, e1004940 (2015).
Bedford, T., Rambaut, A. & Pascual, M. Canalization of the evolutionary trajectory of the human influenza virus. BMC Biol. 10, 38 (2012).
Gog, J. R. The impact of evolutionary constraints on influenza dynamics. Vaccine 26, C15–C24 (2008).
Andreasen, V. & Sasaki, A. Shaping the phylogenetic tree of influenza by cross-immunity. Theor. Popul. Biol. 70, 164–173 (2006).
Hensley, S. E. et al. Hemagglutinin receptor binding avidity drives influenza A virus antigenic drift. Science 326, 734–736 (2009).
Gong, L. I. & Bloom, J. D. Epistatically interacting substitutions are enriched during adaptive protein evolution. PLoS Genet. 10, e1004328 (2014). The study is a key example of how the specific genetic context in which mutations occur can have substantial effect on the resulting fitness of viruses.
Bloom, J. D., Gong, L. I. & Baltimore, D. Permissive secondary mutations enable the evolution of influenza oseltamivir resistance. Science 328, 1272–1275 (2010).
Leonard, A. S. et al. Deep sequencing of influenza A virus from a human challenge study reveals a selective bottleneck and only limited intrahost genetic diversification. J. Virol. 90, 11247–11258 (2016).
Debbink, K. et al. Vaccination has minimal impact on the intrahost diversity of H3N2 influenza viruses. PLoS Pathog. 13, e1006194 (2017). This study provides an interesting analysis of human virus samples relating within-host virus data to vaccination status and finds a minimal role for vaccine-induced immunity as a source of evolutionary selection pressure.
McCrone, J. T. et al. The evolutionary dynamics of influenza A virus within and between human hosts. bioRxiv http://dx.doi.org/10.1101/176362 (2017).
Xue, K. S. et al. Parallel evolution of influenza across multiple spatiotemporal scales. eLife 6, e26875 (2017).
Parvin, J. D., Moscona, A., Pan, W. T., Leider, J. M. & Palese, P. Measurement of the mutation rates of animal viruses: influenza A virus and poliovirus type 1. J. Virol. 59, 377–383 (1986).
Nobusawa, E. & Sato, K. Comparison of the mutation rates of human influenza A and B viruses. J. Virol. 80, 3675–3678 (2006).
Bloom, J. D. An experimentally determined evolutionary model dramatically improves phylogenetic fit. Mol. Biol. Evol. 31, 1956–1978 (2014).
Pauly, M. D., Procario, M. C. & Lauring, A. S. A novel twelve class fluctuation test reveals higher than expected mutation rates for influenza A viruses. eLife 6, e26437 (2017).
Sidorenko, Y. & Reichl, U. Structured model of influenza virus replication in MDCK cells. Biotechnol. Bioeng. 88, 1–14 (2004).
Wu, N.-H. et al. The differentiated airway epithelium infected by influenza viruses maintains the barrier function despite a dramatic loss of ciliated cells. Sci. Rep. 6, 39668 (2016).
Guillot, L. et al. Involvement of toll-like receptor 3 in the immune response of lung epithelial cells to double-stranded RNA and influenza A virus. J. Biol. Chem. 280, 5571–5580 (2005).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Marois, I., Cloutier, A., Garneau, E. & Richter, M. V. Initial infectious dose dictates the innate, adaptive, and memory responses to influenza in the respiratory tract. J. Leukoc. Biol. 92, 107–121 (2012).
Le Goffic, R. et al. Influenza A virus protein PB1-F2 exacerbates IFN-β expression of human respiratory epithelial cells. J. Immunol. 185, 4812–4823 (2010).
Le Goffic, R. et al. Transcriptomic analysis of host immune and cell death responses associated with the influenza A virus PB1-F2 protein. PLoS Pathog. 7, e1002202 (2011).
Everitt, A. R. et al. IFITM3 restricts the morbidity and mortality associated with influenza. Nature 484, 519–523 (2012).
Zimmermann, P., Manz, B., Haller, O., Schwemmle, M. & Kochs, G. The viral nucleoprotein determines Mx sensitivity of influenza A viruses. J. Virol. 85, 8133–8140 (2011).
Gnirss, K. et al. Tetherin sensitivity of influenza A viruses is strain specific: role of hemagglutinin and neuraminidase. J. Virol. 89, 9178–9188 (2015).
Andrews, S. F. et al. Immune history profoundly affects broadly protective B cell responses to influenza. Sci. Transl Med. 7, 316ra192 (2015).
Slütter, B. et al. Dynamics of influenza-induced lung-resident memory T cells underlie waning heterosubtypic immunity. Sci. Immunol. 2, eaag2031 (2017).
Baccam, P., Beauchemin, C., Macken, C. A., Hayden, F. G. & Perelson, A. S. Kinetics of influenza A virus infection in humans. J. Virol. 80, 7590–7599 (2006).
Kitphati, R. et al. Kinetics and longevity of antibody response to influenza A H5N1 virus infection in humans. Clin. Vaccine Immunol. 16, 978–981 (2009).
Ochsenbein, A. F. et al. Protective long-term antibody memory by antigen-driven and T help-dependent differentiation of long-lived memory B cells to short-lived plasma cells independent of secondary lymphoid organs. Proc. Natl Acad. Sci. USA 97, 13263–13268 (2000).
Neuzil, K. M. et al. Immunogenicity and reactogenicity of 1 versus 2 doses of trivalent inactivated influenza vaccine in vaccine-naive 5–8-year-old children. J. Infect. Dis. 194, 1032–1039 (2006).
Renegar, K. B., Small, P. A., Boykins, L. G. & Wright, P. F. Role of IgA versus IgG in the control of influenza viral infection in the murine respiratory tract. J. Immunol. 173, 1978–1986 (2004).
Stokes, C. R., Soothill, J. F. & Turner, M. W. Immune exclusion is a function of IgA. Nature 255, 745–746 (1975).
Nachbagauer, R. & Krammer, F. Universal influenza virus vaccines and therapeutic antibodies. Clin Microbiol. Infect. 23, 222–228 (2017).
Wrammert, J. et al. Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. J. Exp. Med. 208, 181–193 (2011).
Davenport, F. M., Hennessy, A. V. & Francis, T. Epidemiologic and immunologic significance of age distribution of antibody to antigenic variants of influenza virus. J. Exp. Med. 98, 641–656 (1953).
Davenport, F. M., Hennessy, A. V., Stuart-Harris, C. H. & Francis, T. Epidemiology of influenza; comparative serological observations in England and the United States. Lancet 269, 469–474 (1955).
Miller, M. S. et al. Neutralizing antibodies against previously encountered influenza virus strains increase over time: a longitudinal analysis. Sci Transl Med 5, 198ra107 (2013).
Lessler, J. et al. Evidence for antigenic seniority in influenza A (H3N2) antibody responses in southern China. PLoS Pathog. 8, e1002802 (2012).
Davenport, F. M. & Hennessy, A. V. A serologic recapitulation of past experiences with influenza A; antibody response to monovalent vaccine. J. Exp. Med. 104, 85–97 (1956).
Hobson, D., Curry, R. L., Beare, A. S. & Ward-Gardner, A. The role of serum haemagglutination-inhibiting antibody in protection against challenge infection with influenza A2 and B viruses. J. Hyg. 70, 767–777 (1972).
Swayne, D. E. et al. Antibody titer has positive predictive value for vaccine protection against challenge with natural antigenic-drift variants of H5N1 high-pathogenicity avian influenza viruses from Indonesia. J. Virol. 89, 3746–3762 (2015).
Fox, A. et al. Hemagglutination inhibiting antibodies and protection against seasonal and pandemic influenza infection. J. Infect. 70, 187–196 (2015).
Ohmit, S. E., Petrie, J. G., Cross, R. T., Johnson, E. & Monto, A. S. Influenza hemagglutination-inhibition antibody titer as a correlate of vaccine-induced protection. J. Infect. Dis. 204, 1879–1885 (2011).
Hensley, S. E. Challenges of selecting seasonal influenza vaccine strains for humans with diverse pre-exposure histories. Curr. Opin. Virol. 8, 85–89 (2014).
Frise, R. et al. Contact transmission of influenza virus between ferrets imposes a looser bottleneck than respiratory droplet transmission allowing propagation of antiviral resistance. Sci. Rep. 6, 29793 (2016).
Varble, A. et al. Influenza A virus transmission bottlenecks are defined by infection route and recipient host. Cell Host Microbe 16, 691–700 (2014). This is an elegant experimental paper showing how route of transmission is an important factor in virus population bottleneck size.
Moncla, L. H. et al. Selective bottlenecks shape evolutionary pathways taken during mammalian adaptation of a 1918-like avian influenza virus. Cell Host Microbe 19, 169–180 (2016).
Poon, L. L. M. et al. Quantifying influenza virus diversity and transmission in humans. Nat. Genet. 48, 195–200 (2016).
Tamerius, J. D. et al. Environmental predictors of seasonal influenza epidemics across temperate and tropical climates. PLoS Pathog. 9, e1003194 (2013).
Viboud, C., Alonso, W. J. & Simonsen, L. Influenza in tropical regions. PLoS Med. 3, e89 (2006).
Hirve, S. et al. Influenza seasonality in the tropics and subtropics — when to vaccinate? PLoS ONE 11, e0153003 (2016).
Lowen, A. C., Mubareka, S., Steel, J. & Palese, P. Influenza virus transmission is dependent on relative humidity and temperature. PLoS Pathog. 3, e151 (2007).
Deyle, E. R., Maher, M. C., Hernandez, R. D., Basu, S. & Sugihara, G. Global environmental drivers of influenza. Proc. Natl Acad. Sci. USA 113, 13081–13086 (2016).
Shaman, J. & Kohn, M. Absolute humidity modulates influenza survival, transmission, and seasonality. Proc. Natl Acad. Sci. USA 106, 3243–3248 (2009).
Young, L. C. et al. Summer outbreak of respiratory disease in an Australian prison due to an influenza A/Fujian/411/2002(H3N2)-like virus. Epidemiol. Infect. 133, 107–112 (2005).
Finnie, T. J. R., Copley, V. R., Hall, I. M. & Leach, S. An analysis of influenza outbreaks in institutions and enclosed societies. Epidemiol. Infect. 142, 107–113 (2014).
Gaillat, J., Dennetière, G., Raffin-Bru, E., Valette, M. & Blanc, M. C. Summer influenza outbreak in a home for the elderly: application of preventive measures. J. Hosp. Infect. 70, 272–277 (2008).
Cauchemez, S., Valleron, A.-J., Boëlle, P.-Y., Flahault, A. & Ferguson, N. M. Estimating the impact of school closure on influenza transmission from Sentinel data. Nature 452, 750–754 (2008).
Dopico, X. C. et al. Widespread seasonal gene expression reveals annual differences in human immunity and physiology. Nat. Commun. 6, 7000 (2015).
Hirsch, A. & Creighton, C. Handbook of geographical and historical pathology. (London: The New Sydenham Society, 1883).
Hope-Simpson, R. E. The role of season in the epidemiology of influenza. J. Hyg. 86, 35–47 (1981).
Shortridge, K. F., Peiris, J. S. M. & Guan, Y. The next influenza pandemic: lessons from Hong Kong. J. Appl. Microbiol. 94 (Suppl.), 70S–79S (2003).
Nelson, M. I., Simonsen, L., Viboud, C., Miller, M. A. & Holmes, E. C. Phylogenetic analysis reveals the global migration of seasonal influenza A viruses. PLoS Pathog. 3, e131 (2007).
Russell, C. A. et al. The global circulation of seasonal influenza A (H3N2) viruses. Science 320, 340–346 (2008).
Rambaut, A. et al. The genomic and epidemiological dynamics of human influenza A virus. Nature 453, 615–619 (2008).
Chan, J., Holmes, A. & Rabadan, R. Network analysis of global influenza spread. PLoS Comput. Biol. 6, e1001005 (2010).
Lemey, P. et al. Unifying viral genetics and human transportation data to predict the global transmission dynamics of human influenza H3N2. PLoS Pathog. 10, e1003932 (2014).
Bielejec, F., Lemey, P., Baele, G., Rambaut, A. & Suchard, M. A. Inferring heterogeneous evolutionary processes through time: from sequence substitution to phylogeography. Syst. Biol. 63, 493–504 (2014).
Bedford, T., Cobey, S., Beerli, P. & Pascual, M. Global migration dynamics underlie evolution and persistence of human influenza A (H3N2). PLoS Pathog. 6, e1000918 (2010).
Bedford, T. et al. Integrating influenza antigenic dynamics with molecular evolution. eLife 3, e01914 (2014).
Ferguson, N. M., Galvani, A. P. & Bush, R. M. Ecological and immunological determinants of influenza evolution. Nature 422, 428–433 (2003).
Partridge, J. & Kieny, M. P. Global production capacity of seasonal influenza vaccine in 2011. Vaccine 31, 728–731 (2013).
Flannery, B. et al. Enhanced genetic characterization of influenza A(H3N2) Viruses and vaccine effectiveness by genetic group, 2014–2015. J. Infect. Dis. 214, 1010–1019 (2016).
Skowronski, D. M. et al. A perfect storm: impact of genomic variation and serial vaccination on low influenza vaccine effectiveness during the 2014–2015 season. Clin. Infect. Dis. 63, 21–32 (2016).
WHO Writing Group et al. Improving influenza vaccine virus selection: report of a WHO informal consultation held at WHO headquarters, Geneva, Switzerland, 14–16 June 2010. Influenza Other Respir. Viruses 6, 142–152 (2012).
de Jong, J. C., Beyer, W. E. P., Palache, A. M., Rimmelzwaan, G. F. & Osterhaus, A. D. M. E. Mismatch between the 1997/1998 influenza vaccine and the major epidemic A(H3N2) virus strain as the cause of an inadequate vaccine-induced antibody response to this strain in the elderly. J. Med. Virol. 61, 94–99 (2000).
Skowronski, D. M. et al. Low 2012–2013 influenza vaccine effectiveness associated with mutation in the egg-adapted H3N2 vaccine strain not antigenic drift in circulating viruses. PLoS ONE 9, e92153 (2014).
Wong, S.-S. & Webby, R. J. Traditional and new influenza vaccines. Clin. Microbiol. Rev. 26, 476–492 (2013). This is an excellent review of the processes, challenges and future directions for influenza virus vaccine production.
Krammer, F. & Palese, P. Advances in the development of influenza virus vaccines. Nat. Rev. Drug Discov. 14, 167–182 (2015).
Brand, C. & Palese, P. Sequential passage of influenza virus in embryonated eggs or tissue culture: emergence of mutants. Virology 107, 424–433 (1980).
McWhite, C. D., Meyer, A. G. & Wilke, C. O. Sequence amplification via cell passaging creates spurious signals of positive adaptation in influenza virus H3N2 hemagglutinin. Virus Evol. 2, vew026 (2016).
Łuksza, M. & Lässig, M. A predictive fitness model for influenza. Nature 507, 57–61 (2014).
Neher, R. A., Russell, C. A. & Shraiman, B. I. Predicting evolution from the shape of genealogical trees. eLife 3, e03568 (2014).
Neher, R. A., Bedford, T., Daniels, R. S., Russell, C. A. & Shraiman, B. I. Prediction, dynamics, and visualization of antigenic phenotypes of seasonal influenza viruses. Proc. Natl Acad. Sci. USA 113, E1701–E1709 (2016).
Neher, R. A. & Bedford, T. nextflu: real-time tracking of seasonal influenza virus evolution in humans. Bioinformatics 31, 3546–3548 (2015).
Nolan, T. et al. Safety and immunogenicity of a prototype adjuvanted inactivated split-virus influenza A (H5N1) vaccine in infants and children. Vaccine 26, 6383–6391 (2008).
Van Damme, P. et al. Long-term persistence of humoral and cellular immune responses induced by an AS03A-adjuvanted H1N1 2009 influenza vaccine: an open-label, randomized study in adults aged 18–60 years and older. Hum. Vaccin. Immunother. 9, 1512–1522 (2013).
Andrews, N. J. et al. Predictors of immune response and reactogenicity to AS03B-adjuvanted split virion and non-adjuvanted whole virion H1N1 pandemic influenza vaccines. Vaccine 29, 7913–7919 (2011).
Huijskens, E. et al. Immunogenicity, boostability, and sustainability of the immune response after vaccination against Influenza A virus (H1N1) 2009 in a healthy population. Clin. Vaccine Immunol. 18, 1401–1405 (2011).
Smith, D. J., Forrest, S., Ackley, D. H. & Perelson, A. S. Variable efficacy of repeated annual influenza vaccination. Proc. Natl Acad. Sci. USA 96, 14001–14006 (1999).
Skowronski, D. M. et al. Serial vaccination and the antigenic distance hypothesis: effects on influenza vaccine effectiveness during A(H3N2) epidemics in Canada, 2010–2011 to 2014–2015. J. Infect. Dis. 215, 1059–1099 (2017).
Corti, D. et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 333, 850–856 (2011).
Tumpey, T. M., Renshaw, M., Clements, J. D. & Katz, J. M. Mucosal delivery of inactivated influenza vaccine induces B-cell-dependent heterosubtypic cross-protection against lethal influenza A H5N1 virus infection. J. Virol. 75, 5141–5150 (2001).
Hoft, D. F. et al. Comparisons of the humoral and cellular immune responses induced by live attenuated influenza vaccine (LAIV) and inactivated influenza vaccine (IIV) in adults. Clin. Vaccine Immunol. 24, e00414-16 (2016).
Li, C. et al. Selection of antigenically advanced variants of seasonal influenza viruses. Nat. Microbiol. 1, 16058 (2016).
Leroux-Roels, I. et al. Antigen sparing and cross-reactive immunity with an adjuvanted rH5N1 prototype pandemic influenza vaccine: a randomised controlled trial. Lancet Lond. Engl. 370, 580–589 (2007).
Khurana, S. et al. Vaccines with MF59 adjuvant expand the antibody repertoire to target protective sites of pandemic avian H5N1 influenza virus. Sci. Transl. Med. 2, 15ra5 (2010).
Lee, J. et al. Molecular-level analysis of the serum antibody repertoire in young adults before and after seasonal influenza vaccination. Nat. Med. 22, 1456–1464 (2016).
Jiang, N. et al. Lineage structure of the human antibody repertoire in response to influenza vaccination. Sci. Transl. Med. 5, 171ra19 (2013).
Nakaya, H. I. et al. Systems biology of immunity to MF59-adjuvanted versus nonadjuvanted trivalent seasonal influenza vaccines in early childhood. Proc. Natl Acad. Sci. USA 113, 1853–1858 (2016).
Wang, C. et al. B-Cell repertoire responses to varicella-zoster vaccination in human identical twins. Proc. Natl Acad. Sci. USA 112, 500–505 (2015).
Boyd, S. D. & Jackson, K. J. L. Predicting vaccine responsiveness. Cell Host Microbe 17, 301–307 (2015).
Monto, A. S. & Maassab, H. F. Ether treatment of type B influenza virus antigen for the hemagglutination inhibition test. J. Clin. Microbiol. 13, 54–57 (1981).
Mosterín Höpping, A., Fonville, J. M., Russell, C. A., James, S. & Smith, D. J. Influenza B vaccine lineage selection — an optimized trivalent vaccine. Vaccine 34, 1617–1622 (2016).
Heikkinen, T., Ikonen, N. & Ziegler, T. Impact of influenza B lineage-level mismatch between trivalent seasonal influenza vaccines and circulating viruses, 1999–2012. Clin. Infect. Dis. 59, 1519–1524 (2014).
Saito, T. et al. Antigenic alteration of influenza B virus associated with loss of a glycosylation site due to host-cell adaptation. J. Med. Virol. 74, 336–343 (2004).
WHO. FluNet. WHO http://www.who.int/influenza/gisrs_laboratory/flunet/en/ (2017).
Rogers, M. B. et al. Intrahost dynamics of antiviral resistance in influenza A virus reflect complex patterns of segment linkage, reassortment, and natural selection. mBio 6, e02464-14 (2015).
Russell, C. A. et al. The potential for respiratory droplet-transmissible A/H5N1 influenza virus to evolve in a mammalian host. Science 336, 1541–1547 (2012).
Österlund, P. et al. Incoming influenza A virus evades early host recognition, while influenza B virus induces interferon expression directly upon entry. J. Virol. 86, 11183–11193 (2012).
Crotta, S. et al. Type I and type III interferons drive redundant amplification loops to induce a transcriptional signature in influenza-infected airway epithelia. PLoS Pathog. 9, e1003773 (2013).
Miao, H. et al. Quantifying the early immune response and adaptive immune response kinetics in mice infected with influenza A virus. J. Virol. 84, 6687–6698 (2010).
Tas, J. M. J. et al. Visualizing antibody affinity maturation in germinal centers. Science 351, 1048–1054 (2016).
Choi, Y. S. & Baumgarth, N. Dual role for B-1a cells in immunity to influenza virus infection. J. Exp. Med. 205, 3053–3064 (2008).
Deenick, E. K. et al. Naive and memory human B cells have distinct requirements for STAT3 activation to differentiate into antibody-secreting plasma cells. J. Exp. Med. 210, 2739–2753 (2013).
Tinoco, J. C. et al. Immunogenicity, reactogenicity, and safety of inactivated quadrivalent influenza vaccine candidate versus inactivated trivalent influenza vaccine in healthy adults aged ≥18 years: a phase III, randomized trial. Vaccine 32, 1480–1487 (2014).
Gerdil, C. The annual production cycle for influenza vaccine. Vaccine 21, 1776–1779 (2003).
Cox, R. J. et al. A phase I clinical trial of a PER. C6® cell grown influenza H7 virus vaccine. Vaccine 27, 1889–1897 (2009).
Kistner, O. et al. Development of a mammalian cell (Vero) derived candidate influenza virus vaccine. Vaccine 16, 960–968 (1998).
Dormitzer, P. R. Rapid production of synthetic influenza vaccines. Curr. Top. Microbiol. Immunol. 386, 237–273 (2015).
Cox, M. M. J. Recombinant protein vaccines produced in insect cells. Vaccine 30, 1759–1766 (2012).
Cox, M. M. J., Patriarca, P. A. & Treanor, J. FluBlok, a recombinant hemagglutinin influenza vaccine. Influenza Other Respir. Viruses 2, 211–219 (2008).
Alexandrova, G. I. et al. Study of live recombinant cold-adapted influenza bivalent vaccine of type A for use in children: an epidemiological control trial. Vaccine 4, 114–118 (1986).
Rudenko, L., Isakova-Sivak, I. & Donina, S. H7N3 live attenuated influenza vaccine has a potential to protect against new H7N9 avian influenza virus. Vaccine 31, 4702–4705 (2013).
Ramezanpour, B., Pronker, E. S., Kreijtz, J. H. C. M., Osterhaus, A. D. M. E. & Claassen, E. Market implementation of the MVA platform for pre-pandemic and pandemic influenza vaccines: a quantitative key opinion leader analysis. Vaccine 33, 4349–4358 (2015).
Altenburg, A. F. et al. Modified vaccinia virus ankara (MVA) as production platform for vaccines against influenza and other viral respiratory diseases. Viruses 6, 2735–2761 (2014).
Fries, L. F., Smith, G. E. & Glenn, G. M. A. Recombinant viruslike particle influenza A (H7N9) vaccine. N. Engl. J. Med. 369, 2564–2566 (2013).
Fiers, W. et al. M2e-based universal influenza A vaccine. Vaccine 27, 6280–6283 (2009).
Mallajosyula, V. V. A. et al. Influenza hemagglutinin stem-fragment immunogen elicits broadly neutralizing antibodies and confers heterologous protection. Proc. Natl Acad. Sci. USA 111, E2514–E2523 (2014).
Acknowledgements
The authors thank A. Han for his help with Fig. 1b–e. This work was supported by a Wellcome Trust Ph.D. Studentship to V.N.P. and by a Royal Society University Research Fellowship and a Wellcome Trust Collaborative Award to C.A.R.
Author information
Authors and Affiliations
Contributions
V.N.P. and C.A.R. contributed to researching data for article. V.N.P. and C.A.R. substantially contributed to the discussion of content. V.N.P. and C.A.R. wrote the article. V.N.P. and C.A.R. reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Epidemics
-
Infectious disease outbreaks involving a large number of people in a defined geographic location over a defined period of time.
- Fomites
-
Surfaces or objects that can be contaminated by pathogens.
- Pandemics
-
Global infectious disease epidemics.
- Genome reassortment
-
A form of genomic rearrangement where two or more influenza viruses infect the same cell and exchange genomic segments, resulting in a genetically novel virus.
- Antigenic clusters
-
A set of influenza virus variants with similar antigenic profiles.
- Haemagglutination inhibition (HAI) assays
-
Experimental assays used to antigenically characterize viruses based on the ability of host serum to inhibit the virus-induced agglutination of red blood cells.
- Within-host selection
-
Evolutionary selection that occurs at the level of an individual host, generally pertaining to virus fitness or virus interaction with the host immune response.
- Immunodominant
-
A property of an antigen, causing it to be the primary focus of the immune response.
- Plaque assays
-
Experimental assays that measure virus growth rates.
- Microneutralization assays
-
Experimental assays that measure the ability of host serum to neutralize specific antigenic strains.
- Deep mutational scanning
-
An experimental protocol for assessing the mutability and effect of amino acid substitutions at specific positions or across entire proteins.
- Epitopes
-
The parts of an antigen that are recognized by the host adaptive immune response.
- Antigenicity
-
The quality determining the appearance of an antigen to the immune system.
- Avidity
-
The strength of binding between an antigen and a receptor based on multiple chemical bonds.
- Dynamical models
-
A mathematical abstraction of the time-dependent behaviour of an object or system.
- Mucociliary clearance
-
The removal of pathogens by the movement of mucus in the upper respiratory tract by ciliated cells.
- Infectious dose
-
The number of pathogen particles that initiate an infection.
- Antigenic distance
-
A measure of antigenic similarity derived from quantitative representations of haemagglutination inhibition assay data.
- Immune waning
-
The process by which antibody titres or general immune reactivity declines with time in the absence of stimulation.
- Immunological backboosting
-
The recall of previously acquired immune memory upon infection or vaccination with a partially cross-reactive antigen.
- Antigenic seniority
-
The phenomenon of having higher antibody titres to influenza virus variants encountered earlier in life than to more recent viruses.
- Bottlenecks
-
Contractions in population diversity associated with reductions in population size.
- Adjuvant
-
A pharmacological agent that affects the breadth and/or strength of the immune response.
- Antisera
-
The antibody-containing portions of the blood, which are specific for a given pathogen.
- Immune repertoire sequencing
-
Targeted genetic sequencing of the B cell or T cell receptor genes.
Rights and permissions
About this article
Cite this article
Petrova, V., Russell, C. The evolution of seasonal influenza viruses. Nat Rev Microbiol 16, 47–60 (2018). https://doi.org/10.1038/nrmicro.2017.118
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro.2017.118
This article is cited by
-
Bioinformatics analysis identifies a key gene HLA_DPA1 in severe influenza-associated immune infiltration
BMC Genomics (2024)
-
Impaired influenza A virus replication by the host restriction factor SAMHD1 which inhibited by PA-mediated dephosphorylation of the host transcription factor IRF3
Virology Journal (2024)
-
Predictive evolutionary modelling for influenza virus by site-based dynamics of mutations
Nature Communications (2024)
-
Multivalent next generation influenza virus vaccines protect against seasonal and pre-pandemic viruses
Scientific Reports (2024)
-
Determinants of epidemic size and the impacts of lulls in seasonal influenza virus circulation
Nature Communications (2024)